Abstract
Crucial to variety improvement programs is the reliable and accurate prediction of genotype’s performance across environments. However, due to the impactful presence of genotype by environment (G×E) interaction that dictates how changes in expression and function of genes influence target traits in different environments, prediction performance of genomic selection (GS) using single-environment models often falls short. Furthermore, despite the successes of genome-wide association studies (GWAS), the genetic insights derived from genome-to-phenome mapping have not yet been incorporated in predictive analytics, making GS models that use Gaussian kernel primarily an estimator of genomic similarity, instead of the underlying genetics characteristics of the populations. Here, we developed a GS framework that, in addition to capturing the overall genomic relationship, can capitalize on the signal of genetic associations of the phenotypic variation as well as the genetic characteristics of the populations. The capacity of predicting the performance of populations across environments was demonstrated by an overall gain in predictability up to 31% for the winter wheat DH population. Compared to Gaussian kernels, we showed that our multi-environment weighted kernels could better leverage the significance of genetic associations and yielded a marked improvement of 4–33% in prediction accuracy for half-sib families. Furthermore, the flexibility incorporated in our Bayesian implementation provides the generalizable capacity required for predicting multiple highly genetic heterogeneous populations across environments, allowing reliable GS for genetic improvement programs that have no access to genetically uniform material.
Similar content being viewed by others
Introduction
Variety improvement programs are tasked with capturing heritable genomic response to selection across multiple growing environments and field seasons. While climatic uncertainty is outpacing variety development, the condition of global food, fuel, and fiber insecurity has become more vulnerable (Feynman and Ruzmainkin 2007). In the face of diverse abiotic stresses, when considering genomic selection for variety improvement (GS, Meuwissen et al. 2001), reliable prediction of genotype performance across environmental variabilities has become increasingly critical.
However, selection using single-environment (SE) models becomes unreliable in the presence of genotype-by-environment (G × E) interaction (Burgueño et al. 2012; Crossa et al. 2017), due to the heterogeneity of genetic variance across environments, or imperfect genetic correlation of the same traits across sites/seasons (Crossa et al. 2004). Recently, GS models capable of assessing single population performance across multiple environments (ME) have been proposed (López-Cruz et al. 2015; Crossa et al. 2016; Lado et al. 2016; Montesinos-López et al. 2016; Spindel and McCouch 2016; Cuevas et al. 2018). For example, Cuevas et al. (2018) examined the prediction accuracy of six different GS models with G × E interactions using two maize and two wheat datasets. Though some degree of advantage over the conventional SE counterparts can be identified, such gain can only be observed when the phenotypic correlation between environments was high (i.e., above 0.6), and when the traits of interest had moderate to high heritability (Burgueño et al. 2012; López-Cruz et al. 2015; Cuevas et al. 2016; Monteverde et al. 2018). The negative impact of the G × E to the performance of GS models is evident, even more so when inbred lines or homogeneous growing conditions are unavailable (Resende et al. 2012). For example, in tree genetic improvement programs that primarily use open-pollinated families, the half-sib pedigree structure has eventually prevented partitioning G × E interaction from genetic variance owing to the lack of genetically uniform replications, thus impeding the performance of GS models (Beaulieu et al. 2014; Chen et al. 2018; Gamel El-Dien et al. 2018; Alves et al. 2020; Thistlethwaite et al. 2020). Even to this date, studies dealing with genetically heterogeneous lines and half-sibs progeny without replications usually approach this problem by removing or accounting for the environmental effects, instead of directly estimating G × E in the model. For example, Albrecht et al. (2014) studied both Pedigree-BLUP and Genomic-BLUP for heterogeneous populations in ME prediction by employing cross-validation to assess cross-environment prediction performance. To address this challenge in GS, here we developed a statistical method capable of incorporating G × E effect while evaluating the predictability of genetically different populations across environments.
Genome-wide approaches have played an instrumental role in the discovery of new biological insights underpinning complex trait variation (Frazer et al. 2009; Huang et al. 2010; Wang et al. 2019). Despite the tens of thousands of variant-trait associations cataloged (Buniello et al. 2019), few studies have incorporated knowledge learned from genome-wide association studies (GWAS) in a full GS framework. In Lloyd-Jones et al. (2019), it was demonstrated that the inclusion of GWAS summary statistics could return favorable results for height and BMI for over 500,000 individuals from UK Biobank. Through different approaches, Bian and Holland (2017) reached a similar conclusion by directly employing GWAS-estimated effects from the maize NAM RILs (recombinant inbred lines). Furthermore, MacLeod et al. (2016) also proposed that including additional functional knowledge, such as non-synonymous coding change, promoter regions, and known causal variants, could add to predictability.
Predicted by population genetics models, studies attempting to understand the impact of rare or less common variants on complex traits have shown an inverse relationship between the variant’s effect size and its frequency in the population (Park et al. 2011; Bomba et al. 2017). Empirical results from recent GWAS studies are mostly in agreement- that is, common variants have small effects, and rare variants have large effects (Bloom et al. 2019; Fournier et al. 2019; Wainschtein et al. 2019). Low-frequency and rare variants with small to modest effects that are thought to contribute to the missing heritability of many complex traits (Manolio et al. 2009; Eichler et al. 2010) may often have been overlooked because of the process in array production (Ziegler et al. 2008. Bouwman et al. 2017; Zhang et al 2018). As a consequence of not able to capture these rare but favorable alleles, selection based on the genomic estimated breeding values (GEBVs) could lead to loss of genetic diversity which further reduces the long-term genetic gain and prediction accuracy (Jannink 2010; Eynard et al. 2015; Liu et al. 2015; Doublet et al. 2019; Meuwissen et al. 2020; Vanavermaete et al. 2020). In this study, we proposed a flexible GS framework that incorporates marker information beyond just genotypic values, while extending the capability of conventional ME models. Our study uses examples in winter wheat and Interior spruce populations to demonstrate the advantage of including trait- and population-specific genetic characteristics, such as single nucleotide polymorphism (SNP) allele frequency and strength of association with the target phenotypes. Compared to the existing Gaussian Kernel (GK) that assigns a uniform weight to every SNP, our proposed Weighted Kernel (WK) captured more realistic functional genetic relationship of individuals within and cross environments by differentiating the contribution of SNPs. This capacity to address the trait- and population-specific environmental effects is not limited to the use of clonal genetic material or inbred lines, making modeling G × E in GS feasible for trials that utilize highly heterogeneous genetic resource to examine genetic adaptability across the range of a species or environmental variabilities (Risk et al. 2021).
High-throughput technologies have revolutionized biological and medical research and will continue to explore other omics spaces responsible for trait variation and adaptive responses to environmental variability (Halstead et al. 2021; Hasin et al. 2017; Li et al. 2019; Kim et al. 2016; Westhues et al. 2017). Integrating various omics information has become increasingly crucial for complex trait prediction and disease diagnostics (Tieri et al. 2011; Gomez-Cabrero et al. 2014; Higdon et al. 2015; Huang et al. 2017). The ability of kernel-based approaches to leverage the complementing favorable properties of predictors (Schrag et al. 2018), and the association significance to trait variation across growing conditions, would have the potential to provide consistent predictability across traits and environments.
Materials and methods
Duster x Billings hard red winter wheat Doubled Haploid (DH) population
Developed cooperatively by the Oklahoma Agriculture Experiment Station (OAES) and the USDA-ARS, a total of 242 DH lines derived from the intercross of Duster and Billings winter wheat varieties were used in the study. Traits analyzed include grain yield (GY) in kilograms per hectare (kg/ha), sodium dodecyl sulfate sedimentation value (Lorenzo and Kronstad 1987) adjusted for flour protein content (SDS), kernel weight measured by the single kernel characterization system (Perten Instruments, Segeltorp, Sweden) (SKCSKW), and wheat protein on a 12% moisture basis (WHTPRO). Each of these traits was evaluated in three harvest years with varied rainfall (i.e., 19.8, 41.3, and 45.2 cm for 2014, 2015, and 2016, respectively) in Stillwater, OK, USA (36.12 N, 97.09 W), representing three different environments. The average of two field replicates of each DH line per year was used in the analysis.
Genotypes were derived using genotype-by-sequencing (GBS) technology, and 16,265 SNP markers were selected after filtering markers with >50% missing ratio. Missing genotypes of markers were imputed by the marker mean (Nazzicari et al. 2016). Although the genetic profile is the same across three years, the effects of environment on phenotype can vary in different years. Hence, we estimated the single- (\(h_{SE}^2\)) and multi-environment narrow-sense heritability (\(h_{ME}^2\)) from GBLUP using single-year phenotypes and the average of 3-year phenotypes for each trait, respectively. The estimation was implemented in the R package BGLR (Pérez and de los Campos 2014).
Interior spruce population
The Interior spruce breeding population includes a total of 1126 38-year-old trees growing over three sites in British Columbia Canada, i.e., Prince George Tree Improvement Station (PGTIS), Aleza Lake, and Quesnel. Each site has 25 families with various sample sizes (Gamal El-Dien et al. 2015). To reduce the impact of unbalanced sample size on modeling, we randomized each family with respect to its minimum sample size (range from 6 to 16), resulting in 340 trees per site. Phenotypes used for prediction are height in m (HT) and diameter at breast height in cm (DBH) as growth traits; and two wood quality attributes, resistance to drilling (WDres) and wood density in kg/m3 using X-ray densitometry (WDX-ray). The genotypic information regarding GBS SNP can also be found in Gamal El-Dien et al. (2015). The single- (\(h_{SE}^2\)) and multi-environment narrow-sense heritability (\(h_{ME}^2\)) of each trait were estimated from GBLUP by Gamal El-Dien et al. (2015) and were reported in Table 1.
Statistical models
Single-environment (SE) model
The kernel matrix in GS models is normally used to represent the genetic correlation between individuals that can be derived from either pedigree information or molecular marker data. To account for the relatedness in genetic background, the SE model implemented here was an extension of model 1 in Cuevas et al. (2017), by adding a random background genetic effect, bj. The SE model is used for comparison with multi-environment (ME) model we proposed in the next section and the SE model is expressed as follows:
where yj is the response vector with length nj, nj is the total number of phenotypic observations in the jth environment, j = 1, …, m, m is the number of environments; 1nj is a vector of ones with length nj, μj is the overall phenotypic mean of individuals in the jth environment; gj is the random genetic effect of individuals in the jth environment, and we assume \({{{\mathbf{g}}}}_j\sim N( {{{{\mathbf{0}}}},\,\sigma _{g_j}^2{{{\mathbf{K}}}}_j})\) where \(\sigma _{g_j}^2\)is the genetic variance of individuals in the jth environment, Kj (size nj × nj) is the kernel matrix used to describe genetic similarity between individuals in the jth environment; bj is the random background genetic effect of the jth environment that is not explained by genetic markers in the gj, and we assume \({{{\mathbf{b}}}}_j\sim N( {{{{\mathbf{0}}}},\,\sigma _{b_j}^2{{{\mathbf{B}}}}_j})\) where \(\sigma _{b_j}^2\) is the background genetic variance of individuals in the jth environment, Bj (size nj × nj) is a matrix representing the background genetic relationship of two individuals in the jth environment; ej is the random error term of the jth environment, and we assume \({{{\mathbf{e}}}}_j\sim N( {{{{\mathbf{0}}}},\,\sigma _{e_j}^2{\rm I}_{n_j}})\) where \(\sigma _{e_j}^2\) is the residual variance of the jth environment and Inj is the identity matrix with size nj; gj, bj and ej are assumed to be independent.
Multi-environment (ME) model
To fully capture G × E interaction, we proposed a generalization of model 3 in Cuevas et al. (2017). The generalized ME model is capable of predicting different individuals across different environments and the model is expressed as follows:
where y = (y1, y2, …, ym)T; \(\mu = \left( {{{{\mathbf{1}}}}_{n_1}\mu _1,\,{{{\mathbf{1}}}}_{n_2}\mu _2, \ldots ,\,{{{\mathbf{1}}}}_{n_m}\mu _m} \right)^T\); g = (g1, g2, …, gm)T and g ~ N(0, ∑g); b = (b1, b2, …, bm)T and b ~ N(0, ∑b); e = (e1, e2, …, em)T and e ~ N(0, ∑e); g, b, and e are assumed to be independent; ym, μm, gm, bm and em are defined the same as in SE model.
In general, the genetic covariance matrix is
where \(\sigma _{g_m}^2\) and Km are defined the same as in SE model; σg1m is the genetic covariance of individuals in the 1st environment and mth environment; K1m (size n1 × nm) is the kernel matrix representing genetic relationship between individuals explained by genetic markers from the 1st and mth environment.
The background genetic covariance matrix is
where \(\sigma _{b_m}^2\) and Bm are defined the same as in SE model; \(\sigma _{b_{1m}}\) is the background genetic covariance of individuals in the 1st and the mth environment; B1m (size n1 × nm) is a matrix constructed to present the background relationship between two individuals that is not explained by genetic markers in the 1st and mth environment. With this model, background genetic relationship can be appropriately incorporated into both SE and ME models. For instance, when two individuals from the same family in the Interior spruce population, a half-sib relatedness of 0.25 was assigned to indicate their shared genetic background. While when the informative background relationship is unavailable, an identity matrix can be used for both background genetic variance and covariance matrices (Crossa et al. 2017), which was the case for Oklahoma wheat DH population demonstrated in this study.
The covariance matrix of the residuals is
where \(\sigma _{e_m}^2\) and Inm are defined the same as in SE model.
Gaussian kernel (GK) and Weighted kernel (WK)
Here in this study, we compared the model prediction performance with two different kernels, Gaussian kernel (GK) used in de Los Campos et al. (2010) and our proposed weighted kernel (WK). The GK in (3) transforms genetic distance into genetic correlation between individuals.
where KG(.,.) is a positive definite function evaluated by marker genotypes; xi, xk are vectors of marker genotypes for the ith and kth individuals respectively, I, k = 1, …, nj, xi = (xi1, …, xil, …, xip)T and xk = (xk1, …, xkl, …, xkp)T, l = 1, …, p, p is the total number of markers; the allelic states of xil are coded as 0, 1, 2 for AA, Aa, and aa respectively; h is a positive bandwidth parameter that controls the rate of decay of the genetic correlation between two individuals. To determine the optimal value of the parameter, either a grid search method from cross-validation procedure or an empirical Bayesian approach (Pérez-Elizalde et al. 2015) can be applied. In this study, h = 1 was used for simplicity. \(d_{ik}^2\) is squared Euclidean distance between two individuals i and k explained by marker genotypes, and s is the largest value of all \(d_{ik}^2\).
Motivated by Wu et al. (2011) and Yan et al. (2014), we proposed to model additional information such as the frequency and the effects of the variants by a WK method in Eq. (4).
where \(d_{ik}^{^\ast 2} = \mathop {\sum}\nolimits_{l = 1}^p {w_l\left( {x_{il} - x_{kl}} \right)^2}\), s* is the sample maximum of \(d_{ik}^{ \ast 2}\), and the weight wl assigned to the lth marker is based on its minor allele frequency (MAF) and p-values from GWAS model with G×E interaction. The detailed formula of wl is the following
where c1 is a constant; MAFl is the minor allele frequency of the lth marker; α and β are the parameters of Beta distribution density function; pvalue1l and pvalue2l are p-values of main genotypic effect and G × E interaction effect respectively from GWAS for the lth marker, these p-values are adjusted by false discovery rate at 0.05 to account for the multiple hypothesis testing problem (Benjamini and Hochberg 1995; Storey and Tibshirani 2003).
To account for the potential effects of low-frequency variants while incorporating the test statistics from GWAS, we proposed the following formula (6) to determine the value of c1.
where pvalue1 = (pvalue1l, … ,pvalue1l, … ,pvalue1p)T, pvalue2 = (pvalue2l, … ,pvalue2l, … ,pvalue2p)T, and MAF = (MAF1, … ,MAFl, … ,MAFP)T.
As for the setting of α and β in Eqs. (5) and (6), Wu et al. (2011) and Yan et al. (2014) suggested to set α = 1 and β = 25 as a general way to control the impact of rare genetic variants in their GWAS research. In this study, we proposed to fix α = 1 and explore the impact of β on the performance of prediction as such beta density decreases as MAF increases (see Fig. S1 for details). As expected, when both MAF and p value are very small, the value of c1was found to be determined approximately by β, i.e., \(c_1 \approx \frac{{10}}{\beta }\). As a result, β cannot go to infinity to shrink c1 toward zero. To further document the impact of β on prediction performance of model with WK, five values were inspected, i.e., β = 12, 25, 50, 100, and 200. Thus, \(c_1 \approx \frac{{10}}{\beta } \approx\) 0.83, 0.40, 0.20, 0.10, and 0.05.
In addition, we compared the prediction performance of the model using the proposed WK with the model that implements the WK by MAF or p-value alone, denoted as WKMAF and WKPvalue respectively. As comparison, we denoted the WK contributed by both MAF and P value as WKMAF_Pvalue, and its weight wl is from Eq. (5). The weight in WKMAF is calculated by Eq. (7).
where c1, α, and β are defined the same as above.
Similarly, the weight in WKPvalue is formulated as following
Model implementation
SE model
Analysis was conducted in R (R Core Team 2020). The SE model was implemented using R package BGLR with 12,000 iterations and the first 6000 as burn-in for both wheat and spruce data sets (Pérez and de los Campos 2014).
ME model
Duster × Billings hard red winter wheat DH population
In the cases where the same genetic line was evaluated in multiple environments, such as Oklahoma winter wheat DH breeding populations, the ME model was fitted using R package MTM (de los Campos and Grüneberg 2016) with 20,000 iterations and the first 10,000 samples as burn-in.
Interior spruce population
Since no clonal or inbred line material was available for Interior spruce, we expanded the ME model for such scenario by a Bayesian approach to estimate all parameters in the ME model. The detailed implementation of Bayesian approach for ME model can be seen in the Supplementary Material. The model was implemented in R with 100,000 iterations and the first 50,000 as burn-ins. The convergences of Markov Chains for all models were assessed by visualizing the trace plots and running convergence diagnosis using R package CODA (Plummer et al. 2006).
Prediction accuracy evaluation
Both Pearson’s correlation coefficient (PCOR) between observed and predicted phenotypes, and its mean squared error (MSE) were used to assess the model prediction accuracy for each environment. To assess the model prediction performance, we split the data into a training set (TRN) and a testing set (TST). We applied the estimation of model parameters from TRN to TST to get predicted phenotypes. For SE model, we randomly selected 70% of single-environment data as TRN (n70 × 1) and the remaining 30% as TST (n30 × 1). For ME models, we followed cross-validation 2 (CV2) procedure in López-Cruz et al. (2015) to assign individuals to TRN and TST. CV2 mimics the practical prediction scenario related to plant breeders where individual plants are only tested in some environments (Burgueño et al. 2012). We randomly selected 70% of multi-environment data as TRN (n70 × m) and the rest 30% as TST (n30 × m). n70 and n30 stand for 70% and 30% of data, respectively. The random partition was repeated 50 times for SE and ME models respectively to generate an average prediction performance of each model. For WK, we selected the value that produced the highest prediction accuracy in TRN. The calculations of MAF and p values were based on each TRN-TST partition.
Results
For each dataset, we present the following: 1) MAF distribution; 2) summary of phenotypes and the estimated heritability, and 3) prediction performance of the proposed models in the single- and multi-environment settings. PCORs were used to illustrate the prediction performance. Additionally, the results of MSE were found to be consistent with PCORs, i.e., a lower MSE tends to have a higher PCOR. The detailed evaluation of prediction performance can be found in Tables S1, S2.
Minor allele frequency distribution
Duster × Billings hard red winter wheat DH population
The distribution of common and rare SNP allele frequency for Duster × Billings DH population is shown in Fig. 1A, The wheat DH population has ~64% of the SNPs with MAF < 0.2, about 59% <0.1 and 19% between 0.4 and 0.5. Further demonstrated in Fig. S1, the density of Beta(1: β) will converge to zero with large value of Beta random variables. Additionally, regarding to the context of MAF as the value of Beta random variable, 0.2 was considered as the large value.
Interior spruce population
The distribution of MAF was found with a greater degree of rare alleles in all three sites of Interior spruce. There were about 86%, 88%, and 91% of SNPs with MAF < 0.2, for, PGTIS, Aleza Lake, and Quesnel, respectively (Fig. 1B–D).
Summary of phenotypes and heritability estimates
Duster × Billings hard red winter wheat DH population
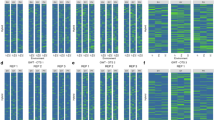
Boxplots of the four traits (GY, SDS, SKCSKW, and WHTPRO) across 3 years (2014–2016) for Oklahoma winter wheat are shown in Fig. 2A. Both trait distributions and phenotypic variation are quite different among the three years for all four traits. For observed phenotypic correlation between years, SDS exhibited the highest average phenotypic correlation (0.52–0.72, average = 0.59), while WHTPRO was the lowest (0.13–0.37, average = 0.22) (Table 1). Heritability for each year (\(h_{SE}^2\)), as well as cross-year estimates (\(h_{ME}^2\)), are also listed in Table 1. GY showed the highest and the most stable single-year heritability among the studied four traits (\(h_{SE}^2\) = 0.60–0.63). Conversely, the other three traits had much lower and varying heritability estimates. For multi-year heritability estimates (2014–2016), only GY and SDS have higher heritability than their single-year estimates (i.e., \(h_{SE}^2\) = 0.60–0.63, \(h_{ME}^2\) = 0.66 for GY;\(h_{SE}^2\) = 0.33–0.46, \(h_{ME}^2\) = 0.46 for SDS).
A Winter wheat, grain yield (GY), SDS sedimentation value (SDS), kernel weight (SKCSKW), and wheat protein (WHTPRO) in each environment (Environment 1/2/3 = Year 2014/2015/2016); B Interior spruce, height (HT), diameter at breast height (DBH), resistance to drilling (WDres), and wood density in kg/m3 using X-ray densitometry (WDX-ray) in each environment (Environment 1/2/3 = PGITS/Aleza Lake/Quesnel, PGTIS, Prince George Tree Improvement Station).
Interior spruce population
Growth phenotypes, HT and DBH, varied among the three Interior spruce sites, while traits related to wood density (e.g., WDres and WDX-ray), Aleza Lake and Quesnel showed similarity in distribution and ranges (Fig. 2B). However, unlike the winter wheat population, the observed pairwise phenotypic correlations were relatively low for all studied traits (Table 1). Overall, WDX-ray had the highest average phenotypic correlation at 13% over the three sites (5–19%), and the lowest was found in the correlation with DBH (1–8%, average at 4%). The single-site heritability ranged from moderate to high (i.e., \(h_{SE}^2\) ranged from 0.26 to 0.56). Generally, traits measured in Quesnel showed higher heritability than the other two sites. The overall heritability estimated across the three sites was reduced to 0.07–0.20 (\(h_{ME}^2\), Table 1), with the highest in HT and the lowest in DBH.
Single and multi-environment predictions
For the performance of WK, we present the results with β that produced the highest prediction accuracy of using WKMAF_Pvalue for both data, i.e., β = 12 for wheat (Fig. S2) and β = 200 for spruce (Fig. S3).
Duster × Billings hard red winter wheat DH population
The average prediction accuracies of SE and ME models are shown in Fig. 3 for the DH hard red winter wheat population. In general, significant improvement in prediction accuracy can be seen with modeling across multiple environments (Fig. 3). For example, the prediction accuracy of GY using single year Gaussian kernel (SE_GK, in Fig. 3) ranged from 0.38 (2016) to 0.55 (2014). With the same Gaussian kernel, ME_GK trained the model with data from all years and generated 6–10% improvement in GY prediction accuracy. The gain from ME models can be as significant as a four-time increase (0.1 in SE_GK and 0.38 for ME_GK, for the SKCSKW in Fig. 3); substantial increase in ME_GK prediction accuracy was also found for SDS with an average increase of 35% over the SE_GK (Fig. 3). SDS also showed the highest gain of the estimated genetic variance from ME_GK vs. SE_GK (Table S3).
Average Pearson’s correlation coefficients were collected over 50 replications of CV2 scheme; SE_GK, single-environment model with Gaussian kernel; ME_GK, multi-environment model with Gaussian kernel; ME_WKMAF, multi-environment model with weighted kernel (WK) by minor allele frequency (MAF); ME_WKPvalue, multi-environment model with WK by genome-wide association study p value; ME_WKMAF_Pvalue (with β = 12), multi-environment model with WK by both MAF and p value; GY, grain yield; SDS, SDS sedimentation value;, SKCSKW, kernel weight; WHTPRO, wheat protein.
Weighting with MAF and the association signal further improved the prediction performance by WK for ME models, with the exceptions of the reduced accuracy found in the ME_WKMAF model for GY and WHTPRO (Fig. 3). Among all three methods of WK, WKMAF_Pvalue performed similarly to WKPvalue, and both significantly outperformed WKMAF for all studied traits. Compared with the ME_GK models, the increase of prediction accuracy from ME_WKMAF_Pvalue models ranged from 1 to 3%, 4 to 5%, and 6 to 7% for SDS, SKCSKW, and WHTPRO, respectively (Fig. 3). For GY, the performance of ME_WKMAF_Pvalue and ME_GK was found similar in 2014 and 2015, but ME_WKMAF_Pvalue produced significantly higher prediction accuracy for 2016 (i.e., 31% higher than ME_GK, Fig. 3).
Interior spruce population
Different from what was observed in the wheat dataset, the advantage of ME_GK over SE_GK was not as evident for Interior spruce, which might be a result of the large amount of variance that cannot be accounted for in the multi-environment models (Table S4). The highest prediction accuracy for SE_GK was found in Quesnel for HT (Fig. 4); similar performance in HT was found for ME_GK as well. Additionally, wood quality traits showed consistent prediction accuracy for all Gaussian models, ranging from 0.19 to 0.31 for WDres and 0.24 to 0.28 for WDX-ray. In Table 1, DBH had the lowest multi-environment heritability estimates, which is further reflected by the average of 5% reduction in ME_GK prediction performance (Fig. 4).
Average Pearson’s correlation coefficients were collected over 50 replications of CV2 scheme; SE_GK, single-environment model with Gaussian kernel; ME_GK, multi-environment model with Gaussian kernel; ME_WKMAF, multi-environment model with weighted kernel (WK) by minor allele frequency (MAF); ME_WKPvalue, multi-environment model with WK by genome-wide association study p value; ME_WKMAF_Pvalue (with β = 200), multi-environment model with WK by both MAF and p value; HT, height; DBH, diameter at breast height; WDres, resistance to drilling; WDX-ray, wood density in kg/m3 using X-ray densitometry; PGTIS, Prince George Tree Improvement Station.
In general, modeling with MAF and specific-trait association improved predictability, even when predicting phenotypes for genetically heterogeneous material across environments. Among the WK implementations, WKMAF_Pvalue outperformed the other WK models almost in all traits, and the WKPvalue model showed slight advantage for WDres prediction in the PGTIS site (Fig. 4). HT was the most predictable phenotype, with a moderate prediction accuracy in Quesnel, increased from 0.48 of SE_GK and 0.47 of ME_GK to 0.56 ME_WKMAF_Pvalue. The greatest gain by using the WK models was, however, found in DBH in Aleza Lake; the benefit of using WK models for DBH was, however, diminished in PGTIS (Fig. 4). The benefit of including genomics signals was not significant for WDX-ray. Due to the relatively indifferent prediction performance for WDX-ray across all models, the benefit of incorporating MAF and association signal was not observed. We suspect that the SNP predictors generated for Interior spruce are in weak LD with the underlying genes and QTLs.
Discussion
GS performance can be influenced by many interrelated factors, including trait genetic architecture, heritability, and the relatedness among individuals between training and testing populations (Crossa et al. 2017). When ME prediction across sites or growing seasons was conducted with a more defined set of genetic diversity like populations derived from controlled crosses, the advantage of incorporating available genetic correlation between environments was evident. As shown in Fig. 3, our ME_GK models using conventional GK demonstrated a consistent improvement over the SE model, showing a 4–38% gain in predictability for Oklahoma winter wheat DH population. The greatest improvement for this population was observed in SDS for 2015, the trait that also showed the most consistent cross-year prediction in Hu et al. (2019). Our results demonstrated that, even in the presence of identifiable environmental variability (\(h_{ME}^2\) ranges from 0.33 to 0.66, Tables 1 and S3), the benefit of employing ME prediction can be anticipated in this case, because of the model capacity to leverage genotype’s environment-specific effect.
Shown in Fig. 4, the ME_GK model, on the other hand, exhibited a slightly unfavorable performance for Interior spruce, except for WDX-ray whose accuracies were found indifferent with the SE model. Compared to our results, the prediction analysis using the same half-sib families in Gamel El-Dien et al. (2015) presented a much-reduced GS accuracy with cross-site validations, even when the prediction accuracy was calculated by correlating the breeding values with the GEBVs. The non-additive effect of these traits was found significant in Gamel El-Dien et al. (2018), with WDX-ray being the only exception. In this study, the multi-site heritability estimates (\(h_{ME}^2\)) ranged from 0.07 to 0.20 (Table 1); this small amount of additive genetic variance would be one of the leading attributes that hinder the performance of the ME_GK model.
The conventional GK using genetic markers is only able to capture the overall genetic similarity between individuals. Although the bandwidth parameter in GK can adjust the distribution of genetic similarity (Pérez-Elizalde et al. 2015), such tuning is uniform to all genetic markers. For Interior spruce, the genetic marker data revealed a much lower relatedness of these trees within each site, suggesting the actual sibling relatedness within families rarely met the half-sib relatedness assumption. In the case where various degree of genetic relatedness between individuals exists within the same family across sites, the strength of incorporating genetic correlation in the ME models using only the Gaussian kernel might be confounded by the heterogeneous genetic background, resulting in an accuracy slightly lower to the SE models (Fig. 4).
The bandwidth parameter tuning in conventional kernel models could potentially create a better mapping between the overall genetic distance among individuals to the phenotypic variation (Pérez-Elizalde et al. 2015). However, it does not reflect the trait’s genomic functional space, leaving important biological insights, such as allele frequencies and the underlying genetic architecture, out of the genome-to-phenome mapping in the GK models. GWAS studies have been a powerful tool to assessing the association between genetic variants and trait variations. The genetic variants identified indicate their functional roles or a close linkage with important genetic determinants for the traits of interest (Wu et al. 2011; Yan et al. 2014; Lin et al. 2016). Several studies have suggested prioritizing GWAS variants when creating the genomic relationship matrix could improve SE predictability of unrelated individuals (de los Campos et al. 2013; Ober et al. 2015; Morgante et al. 2018).
Despite the increases in GWAS statistical power afforded in large international consortia (Willer et al. 2013; Wood et al. 2014; Liu et al. 2015; Astle et al. 2016; Bomda et al. 2017), GWAS still only accounts for a fraction of heritability for most complex traits, a well-known phenomenon called “missing heritability” (Manolio et al. 2009). Genetic variants outside of the reach of the GWAS statistical power are considered to also contribute to the missing heritability (Speed et al. 2012), including common variants with weak effects, low-frequency (MAF 1–5%), and rare variants (MAF < 1%) of small to modest effects, or their combination (Agarwala et al. 2013). When the true causative genetic variants remain unknown, GS has been proven more effective than classic marker-assisted selection. This is because GS employs all available markers as a “compete modeling” methodology for estimating trait performance (Jia 2017). Compared to phenotypic selection, GS could lead to the acceleration of annual inbreeding rate and the loss of genetic diversity as it encourages selecting individuals with high GEBV early in variety improvement programs and those closely related to the training populations (Bassi et al. 2016; Doekes et al. 2018; Forutan et al. 2018). In order to provide stable predictability across populations, GS might also contribute to rapid fixation of genomic regions where consistent marker effects across populations can be identified (Clark et al. 2011; Pszczola et al. 2012; Allier et al. 2019). When the breeding decision is made to optimize short-term genetic gain with conventional GS, rare but favorable alleles could be overlooked. That will essentially reduce selection accuracy and genetic gain in the long term. Demonstrated in simulation studies, up-weighting such alleles would provide 8–30.8% greater long-term gain than that of un-weighted prediction methods (Jannink 2010; Liu et al. 2015), further advocating the WK approaches proposed in this study for the long-term reliability of GS (Rutkoski et al. 2015; Zhang et al. 2018; Ramasubramanian and Beavis 2021).
Here, we presented a flexible GS framework capable of incorporating important genetic attributes to breeding populations and trait variability while addressing the shortcomings of conventional GS models. Shown in Figs. 3, 4, the advantage of incorporating trait- and population-specific genetic characteristics, like p-values of GWAS and MAF, was evident. The MAF component in our WK models aided in preserving rare but favorable variants, which are usually underpowered in GWAS, and in some cases, not even included in the analysis (Pongpanich et al. 2010; Marees et al. 2018). In addition, the WK considers the contribution of genetic markers to the trait-specific G × E. By further differentiating the effects of SNPs between growing environments, GS predictability can be improved for all traits studied for DH genotypes, as well as for the half-sib families of Interior spruce with considerable degree of environmental variability across sites. Finally, the Bayesian kernel methodology presented in the present study offers the flexibility required for predicting multiple populations across environments without using genetically clonal material. This kernel implementation can further encourage integration of other predictors, such as variables in environmental typing (Gianola 2021), to further improve GS performance of highly genetically heterogeneous populations across environments.
Data availability
Code for our proposed model is available here https://github.com/XiaoweiHu-Stat/Multivariate_WeightedKernel.
References
Agarwala V, Flannick J, Sunyaev S, Altshuler D (2013) Evaluating empirical bounds on complex disease genetic architecture. Nat Genet 45:1418–1427
Albrecht T, Auinger HJ, Wimmer V, Ogutu JO, Knaak C, Ouzunova M et al. (2014) Genome-based prediction of maize hybrid performance across genetic groups, testers, locations, and years. Theor Appl Genet 127:1375–1386
Allier A, Lehermeier C, Charcosset A, Moreau L, Teyssèdre S (2019) Improving short- and long-term genetic gain by accounting for within-family variance in optimal cross-selection. Front Genet 10:1006
Alves FC, Balmant KM, Resende Jr MFR, Kirst M, de Los Campos G (2020) Accelerating forest tree breeding by integrating genomic selection and greenhouse phenotyping. Plant Genome 13(3):e20048
Astle WJ, Elding H, Jiang T, Allen D, Ruklisa D, Mann AL et al. (2016) The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167:1415–1429
Bassi FM, Bentley AR, Charmet G, Ortiz R, Crossa J (2016) Breeding schemes for the implementation of genomic selection in wheat (Triticum spp.). Plant Sci Int J Exp Plant Biol 242:23–36
Beaulieu J, Doerksen T, Clément S, Mackay J, Bousquet J (2014) Accuracy of genomic selection models in a large population of open-pollinated families in white spruce. Heredity 113:343–352
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a pratical and powerful approach to multiple testing. J R Stat Soc B 57:289–300
Bian Y, Holland JB (2017) Enhancing genomic prediction with genome-wide association studies in multiparental maize populations. Heredity 118:585–593
Bloom JS, Boocock J, Treusch S, Sadhu MJ, Day L, Oates-Barker H et al. (2019) Rare variants contribute disproportionately to quantitative trait variation in yeast (CR Landry and N Barkai, Eds.). eLife 8:e49212
Bomba L, Walter K, Soranzo N (2017) The impact of rare and low-frequency genetic variants in common disease. Genome Biol 18:77
Bouwman AC, Hayes BJ, Calus MPL (2017) Estimated allele substitution effects underlying genomic evaluation models depend on the scaling of allele counts. Genet Sel Evol 49:79
Buniello A, MacArthur JAL, Cerezo M, Harris LW, Hayhurst J, Malangone C et al. (2019) The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res 47:D1005–D1012
Burgueño J, de los Campos G, Weigel K, Crossa J (2012) Genomic prediction of breeding values when modeling genotype × environment interaction using pedigree and dense molecular markers. Crop Sci 52:707–719
de los Campos G, Hickey JM, Pong-Wong R, Daetwyler HD, Calus MPL (2013) Whole-genome regression and prediction methods applied to plant and animal breeding. Genetics 193:327–345
Chen ZQ, Baison J, Pan J, Karlsson B, Andersson B, Westin J et al. (2018) Accuracy of genomic selection for growth and wood quality traits in two control-pollinated progeny trials using exome capture as the genotyping platform in Noway spruce. BMC Genom 19:946
Clark SA, Hickey JM, van der Werf JH (2011) Different models of genetic variation and their effect on genomic evaluation. Genet Sel Evol GSE 43:18
Crossa J, de los Campos G, Maccaferri M, Tuberosa R, Burgueño J, Pérez-Rodríguez P (2016) Extending the marker × environment interaction model for genomic-enabled prediction and genome-wide association analysis in Durum wheat. Crop Sci 56:2193–2209
Crossa J, Pérez-Rodríguez P, Cuevas J, Montesinos-López O, Jarquín D, de Los Campos G et al. (2017) Genomic selection in plant breeding: methods, models, and perspectives. Trends Plant Sci 22:961–975
Crossa J, Yang R-C, Cornelius PL (2004) Studying crossover genotype × environment interaction using linear-bilinear models and mixed models. J Agric Biol Environ Stat 9:362–380
Cuevas J, Crossa J, Montesinos-López OA, Burgueño J, Pérez-Rodríguez P, de Los Campos G (2017) Bayesian genomic prediction with genotype × environment interaction kernel models. G3 Bethesda Md 7:41–53
Cuevas J, Crossa J, Soberanis V, Pérez-Elizalde S, Pérez-Rodríguez P, Campos G de L, et al. (2016) Genomic prediction of genotype × environment interaction kernel regression models. Plant Genome 9:1–20. https://doi.org/10.3835/plantgenome2016.03.0024
Cuevas J, Granato I, Fritsche-Neto R, Montesinos-Lopez OA, Burgueño J, Sousa MBE et al. (2018) Genomic-enabled prediction kernel models with random intercepts for multi-environment trials. G3 Genes Genomes Genet 8:1347–1365
De los Campos G, Gianola D, Rosa GJM, Weigel KA, Crossa J (2010) Semi-parametric genomic-enabled prediction of genetic values using reproducing kernel Hilbert spaces methods. Genet Res 92:295–308
de los Campos G, Grüneberg A (2016) MTM (Multiple-Trait Model) package. https://quantgen.github.io/MTM/vignette.html
Doekes HP, Veerkamp RF, Bijma P, Hiemstra SJ, Windig JJ (2018) Trends in genome-wide and region-specific genetic diversity in the Dutch-Flemish Holstein-Friesian breeding program from 1986 to 2015. Genet Sel Evol 50:15
Doublet A-C, Croiseau P, Fritz S, Michenet A, Hozé C, Danchin-Burge C et al. (2019) The impact of genomic selection on genetic diversity and genetic gain in three French dairy cattle breeds. Genet Sel Evol 51:52
Eichler EE, Flint J, Gibson G, Kong A, Leal SM, Moore JH et al. (2010) Missing heritability and strategies for finding the underlying causes of complex disease. Nat Rev Genet 11:446–450
El-Dien OG, Ratcliffe B, Klápště J, Porth I, Chen C, El-Kassaby YA (2018) Multienvironment genomic variance decomposition analysis of open-pollinated Interior spruce (Picea glauca x engelmannii). Mol Breed 38:26
Eynard SE, Windig JJ, Leroy G, van Binsbergen R, Calus MP (2015) The effect of rare alleles on estimated genomic relationships from whole genome sequence data. BMC Genet 16:24
Feynman J, Ruzmaikin A (2007) Climate stability and the development of agricultural societies. Clim Change 84:295–311
Forutan M, Ansari Mahyari S, Baes C, Melzer N, Schenkel FS, Sargolzaei M (2018) Inbreeding and runs of homozygosity before and after genomic selection in North American Holstein cattle. BMC Genom 19:98
Fournier T, Abou Saada O, Hou J, Peter J, Caudal E, Schacherer J (2019) Extensive impact of low-frequency variants on the phenotypic landscape at population-scale (CR Landry and N Barkai, Eds.). eLife 8:e49258
Frazer KA, Murray SS, Schork NJ, Topol EJ (2009) Human genetic variation and its contribution to complex traits. Nat Rev Genet 10:241–251
Gamal El-Dien O, Ratcliffe B, Klápště J, Chen C, Porth I, El-Kassaby YA (2015) Prediction accuracies for growth and wood attributes of interior spruce in space using genotyping-by-sequencing. BMC Genom 16:370
Gianola D (2021) Opinionated views on genome-assisted inference and prediction during a pandemic. Front Plant Sci https://doi.org/10.3389/fpls.2021.717284
Gomez-Cabrero D, Abugessaisa I, Maier D, Teschendorff A, Merkenschlager M, Gisel A et al. (2014) Data integration in the era of omics: current and future challenges. BMC Syst Biol 8(Suppl 2):I1
Halstead MM, Islas-Trejo A, Goszczynski DE, Medrano JF, Zhou H, Ross PJ (2021) Large-scale multiplexing permits full-length transcriptome annotation of 32 bovine tissues from a single nanopore flow cell. Front Genet 12:664260
Hasin Y, Seldin M, Lusis A (2017) Multi-omics approaches to disease. Genome Biol 18:83
Higdon R, Earl RK, Stanberry L, Hudac CM, Montague E, Stewart E et al. (2015) The promise of multi-omics and clinical data integration to identify and target personalized healthcare approaches in autism spectrum disorders. Omics J Integr Biol 19:197–208
Hu X, Carver BF, Powers C, Yan L, Zhu L, Chen C (2019) Effectiveness of genomic selection by response to selection for winter wheat variety improvement. Plant Genome 12:1–15. https://doi.org/10.3835/plantgenome2018.11.0090
Huang S, Chaudhary K, Garmire LX (2017) More is better: recent progress in multi-omics data integration methods. Front Genet 8:84
Huang X, Wei X, Sang T, Zhao Q, Feng Q, Zhao Y et al. (2010) Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42:961–967
Jannink J-L (2010) Dynamics of long-term genomic selection. Genet Sel Evol 42:35
Jia Z (2017) Controlling the overfitting of heritability in genomic selection through cross validation. Sci Rep 7:13678
Kim M, Rai N, Zorraquino V, Tagkopoulos I (2016) Multi-omics integration accurately predicts cellular state in unexplored conditions for Escherichia coli. Nat Commun 7:13090
Lado B, Barrios PG, Quincke M, Silva P, Gutiérrez L (2016) Modeling Genotype × Environment interaction for genomic selection with unbalanced data from a wheat breeding program. Crop Sci 56:2165–2179
Li Z, Gao N, Martini JWR, Simianer H (2019) Integrating gene expression data into genomic prediction. Front Genet 25:126
Lin X, Lee S, Wu MC, Wang C, Chen H, Li Z et al. (2016) Test for rare variants by environment interactions in sequencing association studies. Biometrics 72:156–164
Liu H, Meuwissen TH, Sørensen AC, Berg P (2015) Upweighting rare favourable alleles increases long-term genetic gain in genomic selection programs. Genet Sel Evol 47:19
Lloyd-Jones LR, Zeng J, Sidorenko J, Yengo L, Moser G, Kemper KE et al. (2019) Improved polygenic prediction by bayesian multiple regression on summary statistics. Nat Commun 10:5086
López-Cruz M, Crossa J, Bonnett D, Dreisigacker S, Poland J, Jannink J-L et al. (2015) Increased prediction accuracy in wheat breeding trials using a marker × environment interaction genomic selection model. G3 Genes Genomes Genet 5:569–582
Lorenzo A, Kronstad WE (1987) Reliability of two laboratory techniques to predict bread wheat protein quality in nontraditional growing areas. Crop Sci 27:2
MacLeod IM, Bowman PJ, Vander Jagt CJ, Haile-Mariam M, Kemper KE, Chamberlain AJ et al. (2016) Exploiting biological priors and sequence variants enhances QTL discovery and genomic prediction of complex traits. BMC Genomics 17:144
Marees AT, de Kluiver H, Stringer S, Vorspan F, Curis E, Marie‐Claire C, Derks EM (2018) A tutorial on conducting genome‐wide association studies: quality control and statistical analysis. Int J Methods Psychiatr Res 27(2 Jun):e1608
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ et al. (2009) Finding the missing heritability of complex diseases. Nature 461:747–753
Meuwissen TH, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829
Meuwissen THE, Sonesson AK, Gebregiwergis G, Woolliams JA (2020) Management of genetic diversity in the era of genomics. Front Genet 11:880. https://doi.org/10.3389/fgene.2020.00880
Montesinos-López OA, Montesinos-López A, Crossa J, Toledo FH, Pérez-Hernández O, Eskridge KM et al. (2016) A genomic bayesian multi-trait and multi-environment model. G3 Genes Genomes Genet 6:2725–2744
Monteverde E, Rosas JE, Blanco P, Vida FP, de, Bonnecarrère V, Quero G et al. (2018) Multienvironment models increase prediction accuracy of complex traits in advanced breeding lines of rice. Crop Sci 58:1519–1530
Morgante F, Huang W, Maltecca C, Mackay TFC (2018) Effect of genetic architecture on the prediction accuracy of quantitative traits in samples of unrelated individuals. Heredity 120:500–514
Nazzicari N, Biscarini F, Cozzi P et al. (2016) Marker imputation efficiency for genotyping-by-sequencing data in rice (Oryza sativa) and alfalfa (Medicago sativa). Mol Breed 36:69
Ober U, Huang W, Magwire M, Schlather M, Simianer H, Mackay TFC (2015) Accounting for genetic architecture improves sequence based genomic prediction for a drosophila fitness trait. PLoS ONE 10:e0126880
Park J-H, Gail MH, Weinberg CR, Carroll RJ, Chung CC, Wang Z et al. (2011) Distribution of allele frequencies and effect sizes and their interrelationships for common genetic susceptibility variants. Proc Natl Acad Sci 108:18026–18031
Pérez P, de los Campos G (2014) Genome-wide regression and prediction with the BGLR statistical package. Genetics 198:483–495
Pérez-Elizalde S, Cuevas J, Pérez-Rodríguez P, Crossa J (2015) Selection of the bandwidth parameter in a Bayesian kernel regression model for genomic-enabled prediction. J Agric Biol Environ Stat 20:512–532
Plummer M, Best N, Cowles K, Vines K (2006) CODA: convergence diagnosis and output analysis for MCMC. R N. 6:7–11
Pongpanich M, Sullivan PF, Tzeng JY (2010) A quality control algorithm for filtering SNPs in genome-wide association studies. Bioinformatics 26(14):1731–1737
Pszczola M, Strabel T, Mulder HA, Calus MPL (2012) Reliability of direct genomic values for animals with different relationships within and to the reference population. J Dairy Sci 95:389–400
Ramasubramanian V, Beavis WD (2021) Strategies to assure optimal trade-offs among competing objectives for the genetic improvement of soybean. Front Genet 12:675500
Resende Jr MFR, Muñoz P, Acosta JJ, Peter GF, Davis JM, Grattapaglia D, Resende MDV, Kirst M (2012) Accelerating the domestication of trees using genomic selection: accuracy of prediction models across ages and environments. N Phytol 193(3):617–624
Risk C, McKenney DW, Pedlar J, Lu P (2021) A compilation of North American tree provenance trials and relevant historical climate data for seven species. Sci Data 8:29
Rutkoski J, Singh RP, Huerta-Espino J, Bhavani S, Poland J, Jannink JL et al. (2015) Genetic gain from phenotypic and genomic selection for quantitative resistance to stem rust of wheat. Plant Genome 8:eplantgenome2014.10.0074
R Core Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, https://www.R-project.org/
Schrag TA, Westhues M, Schipprack W, Seifert F, Thiemann A, Scholten S et al. (2018) Beyond genomic prediction: combining different types of omics data can improve prediction of hybrid performance in maize. Genetics 208:1373–1385
Speed D, Hemani G, Johnson MR, Balding DJ (2012) Improved heritability estimation from genome-wide SNPs. Am J Hum Genet 91:1011–1021
Spindel JE, McCouch SR (2016) When more is better: how data sharing would accelerate genomic selection of crop plants. N. Phytol 212:814–826
Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci 100(16):9440–5
Thistlethwaite FR, Gamal El-Dien O, Ratcliffe B, Klápště J, Porth I, Chen C et al. (2020) Linkage disequilibrium vs. pedigree: genomic selection prediction accuracy in conifer species. PLoS One 15:e0232201
Tieri P, de la Fuente A, Termanini A, Franceschi C (2011) Integrating Omics data for signaling pathways, interactome reconstruction, and functional analysis. Methods Mol Biol 719:415–433
Vanavermaete D, Fostier J, Maenhout S, De Baets B (2020) Preservation of genetic variation in a breeding population for long-term genetic gain. G3 10:2753–2762
Wainschtein P, Jain DP, Yengo L, Zheng Z, Anthropometry WGTopm, For PMCT-O, et al. (2019) Recovery of trait heritability from whole genome sequence data. ESPE Year book 16
Wang Q-J, Yuan Y, Liao Z, Jiang Y, Wang Q, Zhang L et al. (2019) Genome-wide association study of 13 traits in maize seedlings under low phosphorus stress. Plant Genome 12:1–13
Westhues J, Schrag TA, Heuer C, Thaller G, Utz HF, Schipprack W et al. (2017) Omics-based hybrid prediction in maize. Theor Appl Genet 130:1927–1939
Willer CJ, Schmidt EM, Sengupta S, Peloso GM, Gustafsson S, Kanoni S et al. (2013) Discovery and refinement of loci associated with lipid levels. Nat Genet 45:1274–1283
Wood AR, Esko T, Yang J, Vedantam S, Pers TH, Gustafsson S et al. (2014) Defining the role of common variation in the genomic and biological architecture of adult human height. Nat Genet 46:1173–1186
Wu MC, Lee S, Cai T, Li Y, Boehnke M, Lin X (2011) Rare-variant association testing for sequencing data with the sequence kernel association test. Am J Hum Genet 89:82–93
Yan Q, Tiwari HK, Yi N, Lin W-Y, Gao G, Lou X-Y et al. (2014) Kernel-machine testing coupled with a rank-truncation method for genetic pathway analysis. Genet Epidemiol 38:447–456
Zhang Q, Sahana G, Su G, Guldbrandtsen B, Lund MS, Calus MPL (2018) Impact of rare and low-frequency sequence variants on reliability of genomic prediction in dairy cattle. Genet Sel Evol 50:62
Ziegler A, König IR, Thompson JR (2008) Biostatistical aspects of genome-wide association studies. Biom J 50(1):8–28
Acknowledgements
Funding for this work was supported by grants from the Oklahoma Wheat Research Foundation (for XH, BFC, and CC), Oklahoma Center for the Advancement of Science and Technology (OCAST) award number PS15-011-2 and PS19-004 for CC. This research represents the research outcomes for the USDA HATCH project OKL03011 (CC). Genotyping effort of this manuscript was also supported by the National Science Foundation award number NSF-MRI 1626257 (CC). This work was completed utilizing the High-Performance Computing Center facilities of Oklahoma State University at Stillwater, and also in part by the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by National Science Foundation grant number ACI-1548562. Specifically, it used the Bridges system, which is supported by NSF award number ACI-1445606, at the Pittsburgh Supercomputing Center (PSC) under the resource allocation MCB-180177.
Author information
Authors and Affiliations
Contributions
XH, LZ, and CC designed and discussed the study. XH conducted full data analysis. XH and CC wrote the draft of the manuscript. BFC, YAE, LZ, and CC reviewed and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Chenwu Xu.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hu, X., Carver, B.F., El-Kassaby, Y.A. et al. Weighted kernels improve multi-environment genomic prediction. Heredity 130, 82–91 (2023). https://doi.org/10.1038/s41437-022-00582-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-022-00582-6
- Springer Nature Switzerland AG