Abstract
Predicting glioma grade plays a pivotal role in treatment and prognosis. However, several current methods for grading depend on the characteristics of the whole tumor. Predicting grade by analyzing tumor subregions has not been thoroughly investigated, which aims to improve the prediction performance. To predict glioma grade via analysis of tumor heterogeneity with features extracted from tumor subregions, it is mainly divided into four magnetic resonance imaging (MRI) sequences, including T2-weighted (T2), fluid-attenuated inversion recovery (FLAIR), pre-gadolinium T1-weighted (T1), and post-gadolinium T1-weighted methods. This study included the data of 97 patients with glioblastomas and 42 patients with low-grade gliomas before surgery. Three subregions, including enhanced tumor (ET), non-enhanced tumor, and peritumoral edema, were obtained based on segmentation labels generated by the GLISTRBoost algorithm. One hundred radiomic features were extracted from each subregion. Feature selection was performed using the cross-validated recursive feature elimination with a support vector machine (SVM) algorithm. SVM classifiers with grid search were established to predict glioma grade based on unparametric and multiparametric MRI. The area under the receiver operating characteristic curve (AUC) was used to evaluate the performance of the classifiers, and the performance of the subregions was compared with the results of the whole tumor. In uniparametric analysis, the features from the ET subregion yielded a higher AUC value of 0.8697, 0.8474, and 0.8474 than those of the whole tumor of FLAIR, T1, and T2. In multiparametric analysis, the ET subregion achieved the best performance (AUC = 0.8755), which was higher than the uniparametric results. Radiomic features from the tumor subregion can potentially be used as clinical markers to improve the predictive accuracy of glioma grades.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Gliomas are highly malignant intracranial tumors [1,2,3]. According to the 2016 World Health Organization (WHO) classification scheme for central nervous system tumors, gliomas can be classified as grades I–IV. Stage I and II are considered low-grade gliomas (LGG), and stage III and IV are focused on as high-grade gliomas (HGG) [4]. Glioblastomas (GBM) are the highest-grade malignant gliomas, which are the most malignant and frequently occurring type of primary astrocytomas [5]. In the treatment process, the prediction of glioma grade is one of the challenging tasks [6,7,8]. Glioma grade is an essential indicator of disease occurrence and patient survival [9], which can guide the formulation of surgery or radiation therapy treatment plans [10, 11]. Therefore, it is necessary to develop an accurate and effective method to predict with reasonable accuracy.
The current clinical diagnosis of glioma generally depends on the judgment of the physician assisting with magnetic resonance imaging technology and their experience [12,13,14]. Standard MRI sequences refer to T2-weighted (T2), fluid-attenuated inversion recovery (FLAIR), anterior gadolinium T1-weighted (T1), and posterior gadolinium T1-weighted (T1Gd) techniques [15,16,17]. Moreover, radiomics is an emerging field in medicine and oncology, which can extract information from imaging techniques [18, 19]. Various valuable features can be obtained from MRI sequences, which cannot be easily observed by the naked eye, and these features can efficiently assist in clinical diagnosis. Those features are mathematical descriptions of the visual properties of an image [20,21,22]. For instance, numerical values are included, such as actual pixel intensities, edge strengths, shape descriptors, and regional variation or texture of pixel value [23], which can provide accurate and reliable evidence to aid in clinical decision-making [24, 25]. For instance, a review study conducted by Lohmann et al. illustrated the relationship between imaging features and clinicopathological manifestations of gliomas, in which the value of the features in diagnosing gliomas was demonstrated. However, this study did not present specific protocols due to the limited space [26]. Moreover, Han et al. identified gliomas by constructing clinical and combined models. The results showed that the imaging features could help identify and distinguish glioma grade. However, the radiographic data they used were only T1-enhanced sequences, while other sequences may contain additional functional and biological information [15]. Despite the limitations of the studies mentioned above, it has demonstrated that imaging features can be used for the clinical diagnosis of gliomas [27,28,29,30]. Therefore, some effective protocols for glioma grading are recommended, which are used for clinical diagnosis and prognostic degree. For example, Lu et al. explored the problem of predicting glioma grade by features to identify better gliomas, to construct the machine learning-based segmentation model and prediction models. The final test results were consistent with clinical results, which demonstrated the clinical value of using tumor features to predict glioma grade. However, they did not combine the high-risk characteristics with the prognosis and establish an effective method to analyze glioma heterogeneity [31].
The above-mentioned studies demonstrated the importance of tumor characteristics in predicting glioma grade. However, there are still deficiencies in predicting the grade of gliomas, in that the high heterogeneity and multiple tumor subregions are not considered. Heterogeneity within tumors generally occurs at the molecular level, which reflects macroscopically by different phenotypes on magnetic resonance imaging, which are divided into three tumor subregions [32, 33]. Tumor subregions usually considered are core-enhanced tumors (ET), non-enhanced tumors (NET), and peritumor edema (ED) [34]. With the help of multiparametric MRI, tumor regions can be identified explicitly as active tumors and necrotic and edematous by MRI sequences.
Previous studies have analyzed the features of the MRI sequence of the whole tumor, while it is ignored that there may be more valuable features in MRI sequences of tumor subregions. For example, Chiu et al. demonstrated that the features of tumor subregions showed a higher correlation with the pathological features of glioma than the whole tumor [35]. Zhou et al. demonstrated that quantitative features from the core-enhanced tumor subregion had the discriminative capability in predicting glioma grading [36]. Yang et al. investigated that the subregions showed much more heterogeneous information. They integrated three radiomic features from different tumor subregions to establish a nomogram that could predict glioma grade, whose performance was superior to the one constructed from the radiomic features of the tumor [37]. However, they only studied the radiomic score of tumor subregions and did not investigate the features of MRI sequences. Furthermore, Yin et al. revealed that the subregional features might contain more information than features extracted from the whole tumor, which could predict glioma grading better. However, the comparison between the whole tumor and tumor subregions was only based on a single parametric MRI (T1), which was not comprehensive enough [38]. It was shown that the tumor subregion-based features had better potential than features extracted from the whole tumor. However, it remains unclear whether radiological features from each single tumor subregion based on multiparametric MRI provide more favorable information than from the whole tumor in predicting glioma grade.
Therefore, this paper aims to investigate the prediction of glioma grade by the radiomic features from tumor subregions based on multiparametric MRI. Features from subregions have performed better than those from the whole tumor, improving the accuracy of the diagnosis. The results showed that the ET subregion outperformed other tumor subregions and the whole tumor in predicting glioma grade.
2 Materials and Methods
2.1 Methodology Framework
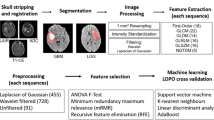
The methodology framework is shown in Fig. 1, which was carried out in the following stages. Firstly, an MRI dataset including four sequences was obtained. Secondly, the tumor subregions and whole tumors were obtained according to segmentation labels generated by the GLISTRBoost algorithm. Thirdly, radiomic features were extracted from the subregions and the whole tumor. Fourthly, the optimal features were selected for model construction. The support vector machine method was used to predict glioma grade. Finally, the performance of the model was accessed through the receiver operating characteristic (ROC) analysis.
2.2 Study Population and Image Data Description
The dataset was obtained from the existing public database on The Cancer Image Archive (TCIA, https://www.cancerimagingarchive.net/): International Multimodal Brain Tumor Segmentation Challenge 2015 (BRATS 2015). Four structural modalities (i.e., T1, T1Gd, T2, FLAIR) acquired at the baseline pre-operative time point were included. The data were obtained from the actual clinical practice of radiographically scanning patients diagnosed with gliomas, which comprised the scan collections of GBM (n = 97) and low-grade glioma (LGG, n = 62). The dataset described computer-aided and manually corrected tumor segmentation labels, which were revised and manually corrected by an expert board-certified neuroradiologist. Those labels can serve as gold standards, according to TCIA. Cases with incomplete tumor segmentation labels were removed. Therefore, in this study, 139 patients were obtained [34, 39, 40], including 97 GBM and 42 LGG cases in Neuroimaging Informatics Technology Initiative (NIfTI) format. Two representative cases are studied, including one GBM and one LGG, as shown in Fig. 2. This public data set has no patient identifier and does not need the approval of the institutional review committee.
Cases for glioma with different grades. The first row shows a FLAIR-based MRI for a patient with GBM from the a axial direction, b sagittal direction, and c coronal direction. The second row shows a FLAIR-based MRI for a patient with LGG in the d axial direction, e sagittal direction, and f coronal direction
2.3 Pre-processing and Subregion Segmentation
MRI data and tumor segmentations were downloaded as NIfTI format files from TCIA. The NIfTI files contained struct data, and segmentation labels were stored as three-dimensional image matrices in the struct data. These labels were generated by GLISTRBoost based on a hybrid generation–discrimination model [41]. GLISTRBoost is defined as a combined generative–discriminative tumor segmentation method. In addition, board-certified neuroradiologists reviewed these segmentation markers and manually corrected misclassifications. Segmentations were matched with the image by matrix multiplication to obtain tumor subregions, including core-enhanced tumor (ET), non-enhanced tumor (NET), and peritumor edema (ED). All segmentations were combined to represent a single whole tumor region. Figure 3 illustrates the results of tumor segmentation for two selected cases (GBM and LGG) using the above-mentioned segmentation.
Tumor segmentation results for cases with different grades. Red, yellow, and green represent the NET, ET, and ED subregions. The first row shows the segmented results of GBM in the a axial direction, b sagittal direction, and c coronal direction and the second row shows the segmented results of LGG in the d axial direction, e sagittal direction, and f coronal direction
2.4 Feature Extraction
Radiomic features were calculated through an open-source toolkit, i.e., PyRadiomics (version 3.0.1) [42]. A total of 100 features were extracted from the tumor subregions (ET, NET, and ED subregions) and the whole tumor of four MRI sequences (T1, T2, T1-weighted, and FLAIR). The extracted features referred to 14 shape features, 18 first-order features, and 62 second-order features. Second-order features consisted of 22 gray-level co-occurrence matrix (GLCM) features, 16 gray-level run length matrix (GLRLM) features, 16 gray-level size-zone matrix (GLSZM) features, and 14 gray-level dependence matrix (GLDM) features. Those features are listed in Supplementary Table S1. To improve the comparability of features and eliminate dimensional effects, the feature data were standardized using the z-score formula before feature selection.
2.5 Feature Selection
Different kinds of radiomic features from each region (three subregions and the whole tumor) of each MRI sequence were combined and used as an input dataset for the machine learning model. Therefore, 16 datasets of features were constructed. Each dataset comprised 70% of the data for training and the remaining 30% for data validation. Tenfold cross-validated recursive feature elimination with an SVM classifier was used to select the good predictive features [43]. In recursive feature elimination, first, all features were rated, and then a different number of features was selected sequentially in cross-validation. It was used to determine the number of features with the highest scores and the most practical combination of features for each training session. The evaluation metric based on the area under the receiver operating characteristic curve (AUC) was used to determine the optimal features during the cross-validation, which were used for the final model construction and prediction.
2.6 Model Construction and Evaluation
Wilcoxon rank sum test was adopted to analyze differences in individual features among glioma grades. The superiority of the Wilcoxon rank sum test depends on its ability to handle non-normal data and small sample sizes. Compared to other statistical tests, such as the two-sample t test, the Wilcoxon rank sum test does not require assumptions about the underlying distribution of the data, which makes it a more robust and reliable test for the data set. Another advantage of the Wilcoxon rank sum test is that it can be easily understood and interpreted. It provides a straightforward and intuitive measure of the difference between the two groups, making it an ideal choice for exploratory analysis or for testing hypotheses about the differences between two groups. The P value of a single feature from subregions and the whole tumor was calculated, and P < 0.05 indicated a statistically significant difference. In uniparametric analysis, univariate SVM classifiers were constructed to investigate the diagnostic performance of individual features in differentiating the GBM and LGG states. Each feature was used as an individual dataset to build a classifier.
Multivariate SVM classifiers were subsequently established to predict glioma grade. The grid search method was applied to select the most suitable combination of hyperparameters in various parameters, including alternative regularization parameters and kernel types. During grid search, tenfold cross-validation with AUC as the evaluation index was performed for the training set. Then, the test set was used to validate the models under the optimal hyperparameters. Each classifier was randomly trained five times, and the average results were calculated. Besides, multiparametric analysis was implemented by combining the features of four MRI sequences as the input dataset. The performance was compared between the tumor subregions and the whole tumor. The results of the multiparametric analysis were also compared with those of the uniparametric analysis. The ROC curve was plotted, and AUC was calculated to indicate the diagnostic performance. Performance metrics in the prediction model were adopted, such as a 95% confidence interval, accuracy, specificity, and sensitivity.
Wilcoxon rank sum test was performed in MATLAB (version 2019b). Feature extraction, feature selection, and model construction were implemented in Python (version 3.7.11). Performance evaluation of the model was done using R (version 4.2.1).
3 Results and Discussion
3.1 Difference Analysis of Individual Features
Eight features with the lowest P values of four MRI sequences are listed in Supplementary Tables S2–S5. It was found that the eight features from the ET subregion had a significantly lower P value than the whole tumor features. Four of the eight features from the ET subregion were shape features.
The box plots of the representative shape features (mesh volume and voxel volume) from the ET subregion and whole tumor are provided in Fig. 4. For the shape features, mesh volume (P = 3.1873E−13) and voxel volume (P = 3.2976E−13) from the ET subregion provided significant differences between GBM and LGG grade in four MRI sequences. It was obtained that there was no significant difference in mesh volume (P = 0.3389) and voxel volume (P = 0.3412) from the tumor.
3.2 Performance Analysis of Individual Features
The diagnostic performance (in terms of AUC) of the eight top-ranked features is provided from the T1-based ET subregion and that of the whole tumor is listed in Table 1. The AUC values ranged from 0.8223 to 0.8902 for the eight features extracted from the ET subregion. The feature of gray-level non-uniformity (GLRLM) from the ET subregion was the best, with an AUC of 0.8902. The AUC value was the highest among the four sequences. For the whole tumor, variance and gray-level variance obtained the highest AUC of 0.7968. The AUC values of the eight features in the ET subregion were higher than those previously. The results showed that the capability of individual features from the ET subregion were relatively higher than the capability of the whole tumor in distinguishing GBM from LGG. Similar results were found for the FLAIR and T2-based ET subregion and the entire tumor (Tables 2, 3). In FLAIR, T1Gd, and T2, the shape feature of mesh volume from the ET subregion obtained the highest AUC of 0.8870 (Tables 2, 3, 4).
Various features were explored in univariate analysis, including shape features, first-order features, and other gray-level matrix features. For diagnostic performance, individual features from the ET subregion were better than those from whole regions. Gray-level non-uniformity of GLRLM (GLN) from the ET subregion in T1 showed the most remarkable ability for differentiating GBM grade. GLN measures the similarity between gray-level intensity values within an image where a lower value correlates with more striking similarities in intensity values. A high correlation was revealed between GLN and various immune markers, and GLN was then used to determine glioma grade and other texture features [44]. Results demonstrated the usefulness of the proposed texture features as a non-invasive biomarker for predicting glioma treatment response. Furthermore, the shape feature mesh volume from the ET subregion performed better than all features extracted from the whole tumor in FLAIR, T2, and T1Gd. In addition, the ET subregion-based three-shape features, such as voxel volume, surface area, and least axis length, performed better than those of the whole tumor. The relationship between the volume shape characteristics of the ET subregion and glioma grade was reported from previous research works. Our findings are somewhat similar to their findings [45]. Those shape features from the ET subregion showed significant differences between GBM and LGG grades in the Wilcoxon rank sum test. However, shape features from the whole tumor did not show significant differences. Therefore, shape features from the ET subregion with a small P value can be used as glioma grade indicators. It is suggested that the distribution of samples from GBM and LGG differed significantly.
3.3 SVM-Based Multivariate Analysis for Predicting Glioma Grade
Table 5 lists the diagnostic performance of multivariate classifiers among tumor subregions and the whole tumor. In uniparametric analysis, the FLAIR-based ET subregion produced the highest AUC of 0.8697. The AUC value was higher than that of the NET and ED subregions. The ET subregion also performed best among the three subregions in the other three MRI sequences. Compared with the whole tumor, the AUC of the ET subregion was higher in FLAIR, T1 (AUC = 0.8474), and T2 (AUC = 0.8474). In T1Gd, the whole tumor produced the highest AUC of 0.8718.
Previous studies predicted glioma grade based on the radiomic characteristics of the whole tumor [34, 46]. Recent studies have analyzed the performance of tumor and edema areas [47]. This paper compared the performance of subregions and the whole tumor in predicting glioma grade. The prediction model with the best performance was established in the multiparametric classifier of the ET subregion. A reasonable explanation is that the ET subregion is considered the region of most intense tumor activity in high-grade gliomas. Since GBM destroys the blood–brain barrier and has rich vascular components, the ET subregion can occupy many contrast agents and show a high enhancement level in MRI-enhanced scanning. Therefore, the characteristic tumor information of the ET subregion is more helpful in distinguishing GBM from LGG than the whole tumor [34].
In the multiparametric analysis, the ET subregion yielded the highest AUC value at 0.8755 in classification among all the subregions and the whole tumor. The performance of the ET subregion was better than the that of other subregions and the whole tumor. In addition, multiparametric classifiers outperformed uniparametric classifiers in the same subregions. ROC and AUC of classifiers of the ET subregion and the whole tumor are presented in Fig. 5. Moreover, the accuracy, sensitivity, and specificity derived from the tumor subregions and the whole tumor are listed in Supplementary Table S6. In FLAIR, T1, and T2, the ET subregion yielded higher accuracy than the other subregions and the whole tumor.
Lambin et al. indicated that using multiple data (e.g., multiparametric MRI data) in radiomics could improve the accuracy and specificity of radiomics [48], which employed multiparametric prediction analysis, i.e., combining the features of four MRI sequences to select and construct SVM classifiers. The whole tumor and subregions showed better prediction performance for multiparametric classification than all uniparametric results. Because the four imaging sequences were acquired differently, the extracted tumor features were different, except for the shape features. In FLAIR, parenchymal lesions and lesions containing bound water had distinct signals. A high signal is evident in the ED subregion of the lesion in T2. In T1Gd, the enhanced tumor areas with rich blood supply appear bright. Image information acquired from multiparametric MRI can effectively complement each other. The key features of the four MRIs can be identified and filtered comprehensively during feature extraction and selection. Moreover, Ning et al. demonstrated the feasibility of integrating multimodal MRI radiomics and in-depth features, which was used to develop a promising non-invasive model for diagnosing glioma grade [49].
Furthermore, the wrapped recursive feature elimination method with cross-validation (REFCV) was used during the feature selection process, which can determine the number of features with the highest scores and the most practical combination of features for each training session [50]. Wrapping feature selection can directly take the performance of the classifier, which is used as the evaluation standard of the feature subset. The feature subset most beneficial to the performance of the classifier is selected. AUC was used as an evaluation metric in both feature selection and classification and SVM was the classifier. Therefore, the feature selection results of the model were more suitable for the model we had chosen. It can improve the accuracy of the experimental results and the standardization of diagnostic prediction.
Public data and open-source software were used in our work. For example, Wu et al. concluded that the public database data could greatly improve clinical data standardization, reproducibility, and generalizability for radiological features. Besides, mature image processing technology and data mining tools were adopted, to perform complex tasks such as automatic tumor segmentation, image processing, and radiomics feature extraction. It can help us improve the efficiency of disease diagnosis and data processing [51].
A preliminary analysis was conducted, and the performance of features from the subregions and the whole tumor was compared in identifying the glioma grade. Several limitations are discussed in this study. Firstly, it was a retrospective study with a relatively small sample size. More samples were collected to validate and refine the present work for future studies. Besides, data were collected from one institution, so the sample may not be representative of the general population, which magnifies the effect of individual difference. Therefore, multiple datasets should be acquired from different institutions to perform external verification of the current model, which improves the generalization ability. Furthermore, we analyzed features only from the four modalities of MRI. In the future, features from other imaging (e.g., diffusion-weighted imaging) can be included. Finally, it would be worthwhile to compare the performance of machine learning models with the diagnoses of physicians.
4 Conclusion
This study predicted glioma grade based on the radiomic features from the tumor subregions, and the performance was compared between the subregions and the whole tumor. The results showed that the ET subregion performed better than the other tumor subregions and the whole tumor in predicting glioma grade. Radiomic features from the ET subregion have the potential to serve as clinical markers, which enhances the prediction of glioma grade. In addition, multiparametric prediction was more accurate than uniparametric prediction. The ET subregion-based multiparametric model best predicted glioma grade (AUC = 0.8755). Therefore, combining multiple uniparametric MRI can provide more comprehensive radiomics information and accurate prediction. Clinicians can depend on the model to accurately predict glioma grades so that treatment plans can be developed promptly. It is beneficial to improve the cure rate of patients with glioma. Further work is required before these methods can be utilized to facilitate the non-invasive evaluation of glioma grade in clinical practice.
Availability of Data and Materials
Data and material data used in the study were obtained from the public database: The Cancer Image Archive (TCIA) (https://www.cancerimagingarchive.net/).
Abbreviations
- MRI:
-
Magnetic resonance imaging
- T1:
-
Gadolinium T1-weighted
- T2:
-
Gadolinium T2-weighted
- FLAIR:
-
Fluid-attenuated inversion recovery
- T1Gd:
-
Posterior gadolinium T1-weighted
- ET:
-
Core-enhanced tumor
- ED:
-
Peritumor edema
- NET:
-
Non-enhanced tumor
- SVM:
-
Support vector machine
- ROC:
-
Receiver operating characteristic
- GBM:
-
Glioblastoma multiforme
- LGG:
-
Low-grade glioma
- AUC:
-
Area under curve
References
Touat, M., Idbaih, A., Sanson, M., Ligon, K.L.: Glioblastoma targeted therapy: updated approaches from recent biological insights. Ann. Oncol. 28, 1457–1472 (2017)
Pallud, J., McKhann, G.M.: Diffuse low-grade glioma-related epilepsy. Neurosurg. Clin. N. Am. 30, 43–54 (2019)
Lah, T.T., Novak, M., Breznik, B.: Brain malignancies: glioblastoma and brain metastases. Semin. Cancer Biol. 60, 262–273 (2020)
Komori, T.: Grading of adult diffuse gliomas according to the 2021 WHO classification of tumors of the central nervous system. Lab. Invest. 102, 126–133 (2022)
Alves, A.L.V., et al.: Role of glioblastoma stem cells in cancer therapeutic resistance: a perspective on antineoplastic agents from natural sources and chemical derivatives. Stem Cell Res. Ther. 12, 206 (2021)
Zhang, L., et al.: Noninvasively evaluating the grading of glioma by multiparametric magnetic resonance imaging. Acad. Radiol. 28, e137–e146 (2021)
Daisy, P.S., Anitha, T.S.: Can artificial intelligence overtake human intelligence on the bumpy road towards glioma therapy? Med. Oncol. 38, 53 (2021)
Jaroch, K., Modrakowska, P., Bojko, B.: Glioblastoma metabolomics-in vitro studies. Metabolites 11, 315 (2021)
Chikada, A., Takenouchi, S., Arakawa, Y., Nin, K.: A descriptive analysis of end-of-life discussions for high-grade glioma patients. Neurooncol. Pract. 8, 345–354 (2021)
Rahman, R.: Toward the next generation of high-grade glioma clinical trials in the era of precision medicine. Cancer J. 27, 410–415 (2021)
Kara, B., et al.: Clinical features, treatment and outcome of childhood glial tumors. Turk. Neurosurg. 32, 135–142 (2022)
Malik, N., et al.: MRI radiomics to differentiate between low grade glioma and glioblastoma peritumoral region. J. Neurooncol. 155, 181–191 (2021)
Sidibe, I., Tensaouti, F., Roques, M., Cohen-Jonathan-Moyal, E., Laprie, A.: Pseudoprogression in glioblastoma: role of metabolic and functional MRI-systematic review. Biomedicines 10, 285 (2022)
Wu, Z., et al.: Advances in magnetic resonance imaging contrast agents for glioblastoma-targeting theranostics. Regen. Biomater. 8, rbab062 (2021)
Han, Y., et al.: Differentiation between glioblastoma multiforme and metastasis from the lungs and other sites using combined clinical/routine MRI radiomics. Front. Cell. Dev. Biol. 9, 710461 (2021)
Kong, Z., et al.: Imaging biomarkers guided anti-angiogenic therapy for malignant gliomas. Neuroimage Clin. 20, 51–60 (2018)
Maurer, C.J., et al.: Do gliosarcomas have distinct imaging features on routine MRI? Neuroradiol. J. 34, 501–508 (2021)
Ehret, F., Kaul, D., Clusmann, H., Delev, D., Kernbach, J.M.: Machine learning-based radiomics in neuro-oncology. Acta Neurochir. Suppl. 134, 139–151 (2022)
Taha, B., Boley, D., Sun, J., Chen, C.: Potential and limitations of radiomics in neuro-oncology. J. Clin. Neurosci. 90, 206–211 (2021)
Bullmore, E., Sporns, O.: Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198 (2009)
Hu, L.S., Hawkins-Daarud, A., Wang, L., Li, J., Swanson, K.R.: Imaging of intratumoral heterogeneity in high-grade glioma. Cancer Lett. 477, 97–106 (2020)
Nilsson, M., Englund, E., Szczepankiewicz, F., van Westen, D., Sundgren, P.C.: Imaging brain tumour microstructure. Neuroimage 182, 232–250 (2018)
Erickson, B.J., Korfiatis, P., Akkus, Z., Kline, T.L.: Machine learning for medical imaging(1). Radiographics 37, 505–515 (2017)
Tagliafico, A.S., et al.: Overview of radiomics in breast cancer diagnosis and prognostication. Breast 49, 74–80 (2020)
Zhang, S., et al.: Role of MRI-based radiomics in locally advanced rectal cancer (Review). Oncol. Rep. 47, 34 (2022)
Lohmann, P., et al.: Radiomics in neuro-oncology: basics, workflow, and applications. Methods 188, 112–121 (2021)
Gaudino, S., et al.: MR imaging of brain pilocytic astrocytoma: beyond the stereotype of benign astrocytoma. Childs Nerv. Syst. 33, 35–54 (2017)
Li, D., et al.: Visualization of diagnostic and therapeutic targets in glioma with molecular imaging. Front. Immunol. 11, 592389 (2020)
Rees, J.: Advances in magnetic resonance imaging of brain tumours. Curr. Opin. Neurol. 16, 643–650 (2003)
Zander, E., et al.: A functional artificial neural network for noninvasive pretreatment evaluation of glioblastoma patients. Neurooncol. Adv. 4, vdab167 (2022)
Lu, D., et al.: The prognosis prediction of GBM based on high-risk subregion and multi-parametric MR imaging. Int. Symp. Image Comput. Digit. Med. (ISICDM) 27, 214–217 (2019)
Fathi Kazerooni, A., Bakas, S., Saligheh Rad, H., Davatzikos, C.: Imaging signatures of glioblastoma molecular characteristics: a radiogenomics review. J. Magn. Reson. Imaging 52, 54–69 (2020)
O’Connor, J.P., et al.: Imaging intratumor heterogeneity: role in therapy response, resistance, and clinical outcome. Clin. Cancer Res. 21, 249–257 (2015)
Bakas, S., et al.: Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci. Data 4, 170117 (2017)
Chiu, F.-Y., Le, N.Q.K., Chen, C.-Y.: A multiparametric MRI-based radiomics analysis to efficiently classify tumor subregions of glioblastoma: a pilot study in machine learning. J. Clin. Med. 10 (2021)
Zhou, M., et al.: Identifying spatial imaging biomarkers of glioblastoma multiforme for survival group prediction. J. Magn. Reson. Imaging 46, 115–123 (2017)
Yang, Y., et al.: An improvement of survival stratification in glioblastoma patients via combining subregional radiomics signatures. Front. Neurosci. 15, 683452 (2021)
Yin, L., Liu, Y., Zhang, X., Lu, H., Liu, Y.: The effect of heterogenous subregions in glioblastomas on survival stratification: a radiomics analysis using the multimodality MRI. Technol. Cancer Res. Treat. 20, 15330338211033060 (2021)
Bakas, S., et al.: Segmentation labels and radiomic features for the pre-operative scans of the TCGA-LGG collection [Data Set]. The Cancer Imaging Archive. Sci. Data (2017)
Clark, K., et al.: The Cancer Imaging Archive (TCIA): maintaining and operating a public information repository. J. Digit. Imaging 26, 1045–1057 (2013)
Bakas, S., et al.: GLISTRboost: combining multimodal MRI segmentation, registration, and biophysical tumor growth modeling with gradient boosting machines for glioma segmentation. Brainlesion 9556, 144–155 (2016)
van Griethuysen, J.J.M., et al.: Computational radiomics system to decode the radiographic phenotype. Cancer Res. 77, e104–e107 (2017)
Pedregosa, F., et al.: Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011)
Chaddad, A., et al.: Deep radiomic signature with immune cell markers predicts the survival of glioma patients. Neurocomputing 469, 366–375 (2022)
Cao, H., et al.: A quantitative model based on clinically relevant MRI features differentiates lower grade gliomas and glioblastoma. Eur. Radiol 30, 3073–3082 (2020)
Matsui, Y., et al.: Prediction of lower-grade glioma molecular subtypes using deep learning. J. Neurooncol. 146, 321–327 (2020)
Zhang, H., et al.: Preoperative contrast-enhanced MRI in differentiating glioblastoma from low-grade gliomas in the cancer imaging archive database: a proof-of-concept study. Front. Oncol. 11, 761359 (2021)
Lambin, P., et al.: Radiomics: the bridge between medical imaging and personalized medicine. Nat. Rev. Clin. Oncol. 14, 749–762 (2017)
Ning, Z., et al.: Multi-modal magnetic resonance imaging-based grading analysis for gliomas by integrating radiomics and deep features. Ann. Transl. Med. 9, 298 (2021)
Karabadji, N.E.I., Seridi, H., Khelf, I., Azizi, N., Boulkroune, R.: Improved decision tree construction based on attribute selection and data sampling for fault diagnosis in rotating machines. Eng. Appl. Artif. Intell. 35, 71–83 (2014)
Fathi Kazerooni, A., et al.: Applications of radiomics and radiogenomics in high-grade gliomas in the era of precision medicine. Cancers 13, 5921 (2021)
Acknowledgements
All authors acknowledge the Tumor Imaging Archive for providing the experimental data used in this study.
Funding
Fully self-funded. The author(s) received no financial support for the research, authorship, and/or publication of this article.
Author information
Authors and Affiliations
Contributions
This study was done through the authors' own efforts, from the formulation of the research idea to the design of the research method, the design and execution of the program, management of data, statistical analysis, and final interpretation. SGD conceived the study, performed all the modeling, conducted the statistical analysis, and helped draft the manuscript. YQZ participated in experiments and drafted the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
There is no conflict of interest of competing interests and, therefore, not applicable.
Ethics approval and consent to participate
These data have eliminated personal privacy. Therefore, this study is not relevant to personal privacy.
Consent for publication
All authors have read and approved the final manuscript. (Both parties complete this review, I hereby consent for the International Journal of Computational Intelligence Systems to the public of this research).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Deng, S., Zhu, Y. Prediction of Glioma Grade by Tumor Heterogeneity Radiomic Analysis Based on Multiparametric MRI. Int J Comput Intell Syst 16, 51 (2023). https://doi.org/10.1007/s44196-023-00230-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44196-023-00230-9