Abstract
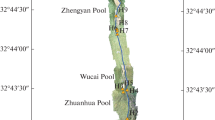
The Xifeng Hot Spring is one of the eight largest hot springs in China, which is rich in radon gas and sulphur in karst scenery. Little is known about the microbiota structure in the spring. The water was collected from three sites containing the outlet of spring water discharge site (OWD), spring pool for tourist (SPT) and sewage effluent pool (SEP) in the Xifeng Hot Spring and further analyzed by culture-independent technique and culture-dependent method. A total of 57 phyla were identified from the water samples. The dominate phyla at OWD was Bacteroidetes (46.93%), while it was Proteobacteria in both sites of SEP and SPT with relative richness of 61.9% and 94.9%, respectively. Two bacteria, Deinococcus and Hymenobacter, that confirmed to be radiation-resistant, seven sulphur bacteria and three thermophilic bacteria were detected from Xifeng Hot Spring. Furthermore, it was found that genus Flavobacterium was susceptible to environmental change with abundance of 11 ~ 2825 times higher in OWD than the other two groups. Compared bacteria from the OWD group with that from 14 hot springs in six countries, total 94 unique genera bacteria were found out from the Xifeng Hot Spring including four thiometabolism-related bacteria (Propionispira, Desulforegula, Desulfobacter and Desulfococcus) and the thermophilic bacterium (Symbiobacterium). Using microbial culturing and isolation technology, sixteen strains were isolated from the water samples of three sites. The diversity of microbiota was abundant and variable along with the niche changed in conditions and surroundings. It indicated that numbers of valuable bacteria resources could be explored from the special surroundings of Xifeng Hot Spring.





Similar content being viewed by others
Data Availability
All of nucleotide sequences of partial 16S rRNA gene from the isolated strains were deposited in NCBI with accession numbers of OQ932838-OQ932853. The sequencing data of water samples are available at NCBI with the BioProject number of PRJNA964588.
References
Mashzhan A, Javier-Lopez R, Kistaubayeva A, Savitskaya I, Birkeland NK (2021) Metagenomics and culture-based diversity analysis of the bacterial community in the Zharkent geothermal spring in Kazakhstan. Curr Microbiol 78(8):2926–2934
Peng Z, Yang L, Ruizhi A, Nanqian Q, Zhen D, Sang B (2022) Spatio-temporal niche of dominant protozoa species in the midstream and downstream of Lhasa river, Tibet. China. Scientia Silvae Sinicae 58(01):78–88
Jingyi Z, Lan L, Yuzhen M, Zhaoyu H, Xiaojuan Z, Zetao L, Min X, Wenjun L (2020) Biodiversity and enzyme activity screening of aquatic prokaryotes from hot springs of Conghua in Guangdong Province. Biotic Resources 42(05):557–567
Ma R (2021) Species diversity analysis and bioactive components of Actinobacteria from Hehua hot spring in Baoshan, Yunnan province. Dali University (Master's thesis)
Sabat AJ, van Zanten E, Akkerboom V, Wisselink G, van Slochteren K, de Boer RF, Hendrix R, Friedrich AW, Rossen JWA, Kooistra-Smid A (2017) Targeted next-generation sequencing of the 16S–23S rRNA region for culture-independent bacterial identification - increased discrimination of closely related species. Sci Rep 7(1):3434
Myer PR, Kim M, Freetly HC, Smith TPL (2016) Evaluation of 16S rRNA amplicon sequencing using two next-generation sequencing technologies for phylogenetic analysis of the rumen bacterial community in steers. J Microbiol Methods 127:132–140
Ye SH, Siddle KJ, Park DJ, Sabeti PC (2019) Benchmarking metagenomics tools for taxonomic classification. Cell 178(4):779–794
Pham VH, Kim J (2012) Cultivation of unculturable soil bacteria. Trends Biotechnol 30(9):475–484
Xian WD, Salam N, Li MM, Zhou EM, Yin YR, Liu ZT, Ming YZ, Zhang XT, Wu G, Liu L, Xiao M, Jiang HC, Li WJ (2020) Network-directed efficient isolation of previously uncultivated Chloroflexi and related bacteria in hot spring microbial mats. NPJ Biofilms Microbiomes 6(1):20
Song X, Duan Q, Meng F, Cao Z (2014) Geological genesis analysis of the Xifeng Hot Spring in Guizhou. Bull Geol Sci Technol 33(05):216–220
Zhang Z, Luo T, Chen Z, Chen G, Yang C, Xiao J (2022) Characteristics of δ~(34) and Sso_4~(2-)、δ_(13)C_(DIC) isotopes in radon-rich geothermal water and their significance— Xifeng Hot Spring as an example. Eco-Environ Knowl 0254–6108:643–652
Rui M, Cheng C, Hai-Feng L, Xue-Xi Z, Ke-Xin S, Meng Y, Wen X, Xiao-Yan Y, Kai-Ling W (2020) Species diversity and enzymatic activity of microorganisms in Chinese hot springs: a review. Microbiology China 47(09):2959–2973
Noll M, Matthies D, Frenzel P, Derakshani M, Liesack W (2005) Succession of bacterial community structure and diversity in a paddy soil oxygen gradient. Environ Microbiol 7(3):382–395
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodriguez AM, Chase J, Cope EK, Da Silva R, Diener C, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibbons SM, Gibson DL, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley GA, Janssen S, Jarmusch AK, Jiang L, Kaehler BD, Kang KB, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu YX, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton JT, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson MS 2nd, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, van der Hooft JJJ, Vargas F, Vazquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang Y, Zhu Q, Knight R, Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852–857
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJ, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581–583
Prodan A, Tremaroli V, Brolin H, Zwinderman AH, Nieuwdorp M, Levin E (2020) Comparing bioinformatic pipelines for microbial 16S rRNA amplicon sequencing. Plos One 15(1):e0227434
Margassery LM, Kennedy J, O’Gara F, Dobson AD, Morrissey JP (2012) Diversity and antibacterial activity of bacteria isolated from the coastal marine sponges Amphilectus fucorum and Eurypon major. Lett Appl Microbiol 55(1):2–8
Bertani G (1951) Studies on lysogenesis. I. The mode of phage liberation by lysogenic Escherichia coli. J Bacteriol 62(3):293–300
Lau MC, Aitchison JC, Pointing SB (2009) Bacterial community composition in thermophilic microbial mats from five hot springs in central Tibet. Extremophiles 13(1):139–149
Rawat N, Joshi GK (2019) Bacterial community structure analysis of a hot spring soil by next generation sequencing of ribosomal RNA. Genomics 111(5):1053–1058
Habib N, Khan IU, Salam N, Xiao M, Ahmed I, Zhi XY, Li WJ (2018) Tepidimonas sediminis sp. nov. and Tepidimonas alkaliphilus sp. nov., two novel moderately thermophilic species isolated from a hot spring. Antonie Van Leeuwenhoek 111(7):1023–1031
Ueda K, Ohno M, Yamamoto K, Nara H, Mori Y, Shimada M, Hayashi M, Oida H, Terashima Y, Nagata M, Beppu T (2001) Distribution and diversity of symbiotic thermophiles, Symbiobacterium thermophilum and related bacteria, in natural environments. Appl Environ Microbiol 67(9):3779–3784
Panosyan H, Birkeland NK (2014) Microbial diversity in an Armenian geothermal spring assessed by molecular and culture-based methods. J Basic Microbiol 54(11):1240–1250
Tindall BJ, Rossello-Mora R, Busse HJ, Ludwig W, Kampfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60(Pt 1):249–266
Lee EM, Jeon CO, Choi I, Chang KS, Kim CJ (2005) Silanimonas lenta gen. nov., sp. nov., a slightly thermophilic and alkaliphilic gammaproteobacterium isolated from a hot spring. Int J Syst Evol Microbiol 55(Pt 1):385–389
Blackwood BP, Hunter CJ (2016) Cronobacter spp. Microbiol Spectr 4(2). https://doi.org/10.1128/microbiolspec.EI10-0002-2015
Samarasinghe SN, Wanigatunge RP, Magana-Arachchi DN (2021) Bacterial diversity in a Sri Lankan geothermal spring assessed by culture-dependent and culture-independent approaches. Curr Microbiol 78(9):3439–3452
Rozanov AS, Bryanskaya AV, Ivanisenko TV, Malup TK, Peltek SE (2017) Biodiversity of the microbial mat of the Garga hot spring. BMC Evol Biol 17(Suppl 2):254
Brochier-Armanet C, Boussau B, Gribaldo S, Forterre P (2008) Mesophilic Crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nat Rev Microbiol 6(3):245–252
Li F, Altermatt F, Yang J, An S, Li A, Zhang X (2020) Human activities’ fingerprint on multitrophic biodiversity and ecosystem functions across a major river catchment in China. Glob Chang Biol 26(12):6867–6879
Yuan M, Zhang W, Dai S, Wu J, Wang Y, Tao T, Chen M, Lin M (2009) Deinococcus gobiensis sp nov, an extremely radiation-resistant bacterium. Int J Syst Evol Microbiol 59(Pt 6):1513–7
Billi D, Friedmann EI, Hofer KG, Caiola MG, Ocampo-Friedmann R (2000) Ionizing-radiation resistance in the desiccation-tolerant cyanobacterium Chroococcidiopsis. Appl Environ Microbiol 66(4):1489–1492
Nayak T, Sengupta I, Dhal PK (2021) A new era of radiation resistance bacteria in bioremediation and production of bioactive compounds with therapeutic potential and other aspects: An in-perspective review. J Environ Radioact 237:106696
Deng W, Yang Y, Gao P, Chen H, Wen W, Sun Q (2016) Radiation-resistant Micrococcus luteus SC1204 and its proteomics change upon gamma irradiation. Curr Microbiol 72(6):767–775
Lee SE, Ten LN, Park Y, Maeng S, Zhang J, Kim MK, Cha IT, Lee KE, Lee BH, Jung H, Kim MK (2021) Hymenobacter busanensis sp. nov., radiation-resistant species isolated from soil in South Korea. Arch Microbiol 203(2):755–762
Ueki A, Watanabe M, Ohtaki Y, Kaku N, Ueki K (2014) Description of Propionispira arcuata sp. nov., isolated from a methanogenic reactor of cattle waste, reclassification of Zymophilus raffinosivorans and Zymophilus paucivorans as Propionispira raffinosivorans comb. nov. and Propionispira paucivorans comb. nov. and emended description of the genus Propionispira. Int J Syst Evol Microbiol 64(Pt 10):3571–3577
Rees GN, Patel BK (2001) Desulforegula conservatrix gen. nov., sp. nov., a long-chain fatty acid-oxidizing, sulfate-reducing bacterium isolated from sediments of a freshwater lake. Int J Syst Evol Microbiol 51(Pt 5):1911–1916
Zhou L, Ou P, Zhao B, Zhang W, Yu K, Xie K, Zhuang WQ (2021) Assimilatory and dissimilatory sulfate reduction in the bacterial diversity of biofoulant from a full-scale biofilm-membrane bioreactor for textile wastewater treatment. Sci Total Environ 772:145464
Zhang Z, Luo T, Cheng Z, Chen G, Yang C, Xiao J (2022) Characteristics of δ34Sso42- and δ13CDIC isotopes in radon-rich geothermal water and their significance - Xifeng hot spring as an example. Environmental Chemistry 41(02):643–652
Shimizu T, Masuda S (2020) Persulphide-responsive transcriptional regulation and metabolism in bacteria. J Biochem 167(2):125–132
Chieffi D, Fanelli F, Fusco V (2020) Arcobacter butzleri: Up-to-date taxonomy, ecology, and pathogenicity of an emerging pathogen. Compr Rev Food Sci Food Saf 19(4):2071–2109
Yang Z, Sadakane T, Hosokawa H, Kuroda M, Inoue D, Ike M (2021) Factors affecting antimonate bioreduction by Dechloromonas sp. AR-2 and Propionivibrio sp AR-3. 3 Biotech 11(4):163
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Lucy Seldin
Importance: This paper identified the microbial community in the water samples of Xifeng Hot Spring. From those, two radiation-resistant bacteria, seven sulphur metabolism bacteria, three known thermophilic bacteria were classified, and 94 unique genus bacteria were screened from Xifeng Hot Spring compared with other fourteen springs. It is valuable for the exploration of these microbial resources in extreme environments in the near future.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, L., Long, H., Huang, S. et al. Bacterial diversity in water from Xifeng Hot Spring in China. Braz J Microbiol 54, 1943–1954 (2023). https://doi.org/10.1007/s42770-023-01070-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01070-7