Abstract
Purpose
The growing prevalence of soil salinity presents a significant threat to agriculture production on a global scale. Previous studies on salt stress, shown that silicon (Si) has an alleviating effect on plants exposed to stress. However, the results of the alleviating effect of Si on epigenetic level is not yet understood. In this study, we tried to understand how methylation mechanisms affect the alleviating effect of Si by testing on Arabidopsis epigenetic mutants (met1-7, drm2-2 and ros1-4).
Methods
The Col-0 and mutant plants were exposed to silicon and NaCl simultaneously and separately during two weeks. After that in order to see the physiological effects of Si on methylation mutants, which is known to be effective in antioxidant pathways of Col-0 plants, osmolyte accumulation and membrane damage were analyzed and to see the effects at the molecular level, the expression profiles of the CSD2, CAT3 and APX1 genes and global methylation changes were analyzed.
Results
As a general result of the osmolyte accumulation, ion leak, global methylation and gene expression analyzes performed in this study, it was determined that salt stress also had negative effects on Arabidopsis epigenetic mutants. It was concluded that the mitigating effect of Si on NaCl stress was most clearly determined as a result of global DNA methylation analyses.
Conclusions
It was found that treating Arabidopsis methylation mutants with Si during salt stress could improve the plants’ ability to withstand salt. The results of this study provide information about the alleviating effect of Si based on methylation of separate and co-exposure to Si and NaCl, and also provide an epigenetic perspective to explain the mechanisms of Si improving plant durability under stress conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Since plants are sessile, they regulate their relationships with their environment against the many different environmental stressors they encounter (Abdulkhair and Alghuthaymi 2016; Peck and Mittler 2020). Currently, agricultural areas are becoming unfavorable for plants due to climatic fluctuations or/and anthropogenic interferences that disrupt the hydrological properties of the soil. Salinity and drought are the most common type of stress plants encounter in nature. The high concentration of salts in the soil makes soil water inaccessible to plants due to its low water potential (Sánchez-Romera and Aroca 2020).
Under salinity stress, plants exert adaptive molecular mechanisms to mitigate various morphological and physiological changes (Rahnama et al. 2010; James et al. 2011; Abobatta 2020). Plants respond to salinity at the osmotic and ionic stress tolerance levels. In case of insufficient ionic homeostasis, the increased Na+ content in the cell causes reactive oxygen species (ROS) to accumulate. Accumulated ROS negatively affect cell homeostasis by causing enzyme inactivation and DNA damage. In order to cope with this situation, the osmotic balance is controlled by antioxidant defense systems consisting of both enzymatic and non-enzymatic components. Enzymatic antioxidants form the main line of defense against free radicals in the cell, and in general, the accumulation of compounds is positively correlated against salt stress. At the beginning of these enzymatic antioxidants are Superoxide dismutase (SOD), Catalase (CAT) and Ascorbate peroxidase (APX) enzymes (Shi et al. 2006; Tuteja 2007; Munns and Tester 2008; Isayenkov and Maathuis 2019). SOD has the ability to convert oxygen and hydrogen peroxide. SODs are divided into 3 classes based on their metal cofactors: Cu/Zn-SOD (CSD) (found in cytosol, chloroplast, peroxisome), Mn-SOD (MSD) (found in mitochondria or peroxisome), and Fe-SOD (FSD) (found mostly in mitochondria and chloroplast). The enzyme CAT removes hydrogen peroxide which is reduced by two electrons from oxygen. APX is primarily found in the chloroplast and cytosol and work as the key enzyme of the ascorbate cycle, converting ascorbic acid to dehydroascorbate, eliminating peroxidase (Asada 1992, Willekens et al. 1997; Al-aghabary et al. 2005; Prashanth et al. 2008; Bose et al. 2014; Ighodaro and Akinloye 2018).
Silicon (Si) has vital roles in plants’ life as a macro element. This element is the most abundant element in various compound forms on earth after oxygen, and constitutes 25–30% of the earth’s crust. Plantstake Si from the soil as silicic acid [Si(OH)4], an uncharged monomeric molecule, under less than 9 pH (Shi et al. 2013; Luyckx et al. 2017; Tripathi et al. 2017; Saitoh et al. 2021). Although the beneficial effects of Si are minimal under optimal conditions, the rise in stress factors (abiotic and/or biotic) increases the importance of Si for both agricultural and horticultural activities. It is stated that the harvest quality of various crops is increased by fertilization using silicon. However, the mechanisms of Si-mediated stress responses are still not fully elucidated (Savvas et al. 2009; Ashraf et al. 2010; Bakhat et al. 2018; Siam et al. 2018).
Many stages of development and responses to environmental cues are regulated directly or indirectly by epigenetic mechanisms (Sudan et al. 2018). DNA methylation is the most studied type of epigenetic modification in the cell. It occurs via the transfer of the methyl group from S-adenosyl methionine (SAMe) to cytosines by methyltransferase enzymes after replication. Demethylation of DNA can be passive and/or active in plants. Passive DNA demethylation occurs when methyltransferases are inactive during the cell cycle following DNA replication, leaving the newly synthesized strand unmethylated. Active DNA demethylation occurs when DNA glycosylases, which are normally associated with DNA repair, recognize 5-methylcytosine and remove the methyl group from DNA. DNA demethylation can activate expressed alleles of some imprinted genes (Gehring et al. 2009; Zhu 2009). DNA methylation in plants occurs in two types; symmetrical CG, CHG, and asymmetrical CHH (where H represents A, T, or C) (Kong et al. 2018). With the completion of A. thaliana genome sequencing in 2000, it was revealed that there are three main enzymes responsible for the methylation of cytosines in plants. These three methylation enzymes are: METHYLTRANSFERASE 1 (MET1), plant-specific CHROMOMETHYLASE 3 (CMT3) and DOMAINS REARRANGED METHYLASE 2 (DRM2). Plant-specific Pol IV and Pol V, developed from Polymerase II, are involved in RNA-mediated DNA methylation (RdDM) (Chinnusamy and Zhu 2009; Law and Jacobsen 2010). Also, four DNA demethylase genes have been identified in Arabidopsis, namely DEMETER (AtDME), SILENCING REPRESSOR (AtROS1), DEMETER LIKE 2 (AtDML2) and DEMETER LIKE 3 (AtDML3) (Chan et al. 2005). ROS1 acts as a transcriptional gene silencing repressor by catalyzing the removal of 5-mC and actively demethylating target DNA without the need for replication via a base excision repair pathway (Bharti et al. 2015).
The molecular mechanism of silicon has not yet been elucidated. Although the impact of changes in methylation level on gene regulation is well known, there are extremely limited studies on methylation level changes related to Si metabolism. In this study, the effect of methylation on silicon metabolism was examined by studying two methylation and one demethylation mutants of the Arabidopsis plant. To examine the salt stress mitigating effects of Si, combined salt (NaCl) and silicon [Si(OH)4] treatment was applied to Arabidopsis thaliana methylation mutants (met1-7, drm2-2, and ros1-4). Then, osmolyte accumulation and ion leakage analyzes were performed within the scope of testing physiological changes in mutant plants. The expression of CSD2, CAT3 and APX1 genes, which encode enzymes involved in antioxidant pathways, was examined. Additionally, methylation changes of the mutants used in this study after Si and/or NaCl treatment were analyzed at the global DNA methylation level.
2 Materials and Methods
2.1 Plant Material and Definition of Mutants
The seeds used in this study are mutants of the genes MET1, DRM2 and ROS1 all of which are Columbia-0 ecotypes. Previously described A. thaliana met1-7, drm2-2 and ros1-4 mutants were used as the experimental group while Col-0 (Columbia-0) plants were used as the control group [Col-0 seeds obtained by Dr. Ralf Stracke [The Center for Biotechnology (CeBiTec), Bielefeld University]. The characteristics of the homozygous seeds, NASC codes, Salk numbers are given in Table 1.
2.2 Seed Germination, Growth Conditions and Stress Treatment
All seeds were surface sterilized and placed on to Murashige and Skoog medium (MS) (Murashige and Skoog 1962) for in vitro culturing. Col-0, met1-7, drm2-2 and ros1-4 seeds were incubated for 7 days to germinate under fluorescent light in a plant growth chamber [16 h light / 8 h dark conditions, 105 µmol m− 2s− 1 (Sanyo, MLR-352 H)] at 25 °C.
Arabidopsis plants were transferred to the medium prepared including 100 mM NaCl (Merck, 1.06404), 1 mM Si (Sigma Aldrich, 306,363) and 100 mM NaCl + 1 mM Si for stress application and incubated for 14 days. After stress treatment, plants were used directly for physiological analysis, the rest was stored at -80 °C to use in molecular analysis. For Si application, silicic acid, which is the form in which plants can uptake the easiest from the soil, was used (Luyckx et al. 2017; Tripathi et al. 2017).
2.3 Physiological Analysis
The osmolyte accumulation was measured using the semi-micro osmometer (Knauer K-7400) as stated by Dourado et al. (2019), and ion leakage in the cell membrane wasmeasured using a conductivity meter (HORIBA B-137) as described by Shi et al. (2006).
2.4 DNA Isolation and Global DNA Methylation (5-mC) Level Analysis
DNA isolation from ∼ 100 mg (is equal to 2 plants for Col-0 and 4 or 5 plants for mutants) of plants harvested after treatment was performed with a GeneJET Plant Genomic DNA Purification Kit (Thermo Scientific, K0792) according to the manufacturer’s protocol. Then, the DNA samples were checked for their integrity, purity and quantity. Their integrities were analyzed by agarose gel electrophoresis and their purities as well as quantities were checked by NanoDrop 2000 (Thermo Scientific). The DNA samples were stored at -20 °C to be used in the methylation experiment. Global methylation changes (%) in genomic DNA were detected with the MethylFlash Global DNA Methylation (5-mC) ELISA Easy Kit (EpiGentek, p-1030) using an indirect ELISA method which is based on antigen-antibody reactions, according to the manufacturer recommended instructions using 100 ng of gDNA for each sample.
2.5 RNA Isolation and qPCR Analysis
After treatment, 100 mg (is equal to 2 plants for Col-0 and 4 or 5 plants for mutants) plant samples were collected from all experimental groups for isolation of RNA with Hibrizol (Hibrigen, Türkiye). Their integrity was checked with agarose gel electrophoresis (1%). Their purity was checked using a NanoDrop 2000 (Thermo Scientific) and their quantity was measured. After quantitative and qualitative controls, cDNA synthesis was performed with the High-Capacity cDNA Reverse Transcription Kit (Thermo Scientific, 4,368,814).
The gene expression profiles of CSD2, CAT3 and APX1 genes were evaluated in met1-7, drm2-2 and ros1-4 epigenetic mutants and Col-0. The primer sequences of the selected genes were designed using the Primer3 program (http://primer3.ut.ee/). Their sequences and product lengths are given in Table 2. qPCR experiments were performed with the LightCycler Nano instrument (Roche, Basel, Switzerland) using 2X SYBR Green qPCR Mix (Hibrigen, Türkiye). The Actin 8 gene was chosen as the endogenous control for normalization of qPCR results. The cycling conditions were set as hold (50 °C, 60 s), denaturation (94 °C, 30 s.), annealing (59 °C for Actin 8, CAT3 and APX1; 68 °C for CSD2 30 s.), extension (72 °C, 30 s.), and hold (40 °C, 30 s.), respectively. The Ct values were evaluated relative to the Actin 8 gene using the 2−ΔΔCt method (Livak and Schmittgen 2001).
2.6 Statistical Analysis
Physiological experiments with Col-0, met1-7, ros1-4 and drm2-2 plants, global DNA methylation level and gene expression analysis were performed in three biological and two technical repetitions. The data obtained in the experiments were statistically analyzed using GraphPad Prism® 9.0.0 software using a one-way ANOVA with the post-hoc Tukey’s test. P values less than 0.001 are given three asterisks, and P values less than 0.0001 are given four asterisks (ns P > 0,05, * P ≤ 0,05, ** P ≤ 0,01, *** P ≤ 0,001, **** P ≤ 0,0001).
3 Results
3.1 Physiological Effects of Stress Application
The analyzes to determine osmolyte accumulation and ion leakage were performed to evaluate the physiological effects of Si, NaCl and the combined Si and NaCl treatments on epigenetic mutants. For this purpose, osmolyte accumulation and ion leakage were evaluated between the experimental groups.
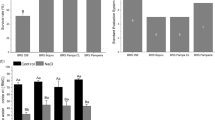
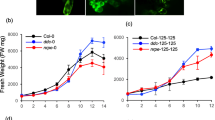
It is observed that NaCl application in Col-0 plants causes a significant increase compared to control and combined applications of Si and NaCl, while Si treatment does not cause a significant change at the end of (Fig. 1). A significant increase was observed in the met1-7 mutant only between the control and NaCl treatment. In the drm2-2 mutant, Si application caused a significant decrease compared to the control group, while NaCl application caused a significant increase of osmolytes. It was determined that combined Si and NaCl application significantly reduced the amount of osmolytes compared to NaCl application. This showed us that Si has a mitigating effect on NaCl stress in the drm2-2 mutant. In ros1-4 mutants, as compared to the control, the application of Si and NaCl resulted in a significant increase in osmolytes. When the osmolyte accumulation data were examined in general, it was observed that NaCl application caused an increase in the amount of osmolytes in all epigenetic mutants, similar to the Col-0 plant.
According to the ion leakage analysis results shown in Fig. 2, it was observed that Si applied to Col-0 and drm2-2 mutants did not cause ion leakage, but in the met1-7 mutant, Si had a slightly increasing effect on ion leakage. While an increase in ion leakage was observed after NaCl application in Col-0 and ros1-4 mutants, a mitigating effect of Si was observed when Si and NaCl was applied simultenously. Salt stress significantly increased ion leakage in all experimental groups except the drm2-2 mutant.
3.2 Global DNA Methylation
Global DNA methylation analysis showed that Si application caused significant degree of hypermethylation in all the experimental groups except the ros1-4 mutants (Fig. 3). On the other hand, while NaCl application caused hypomethylation in all groups (more severe in epigenetic mutants), the combined applications of Si and NaCl eliminated the hypomethylation effect of NaCl.
3.3 Changes in Gene Expression Profiles
It was observed that when Si was applied to the Col-0 plant alone, CSD2 gene expression which is one of the initial steps of the antioxidant process activated following stress in plants, did not change, but increased when Si and NaCl applied simultaneously. In the met1-7 mutant, unlike the Col-0 plant, it was seen that all the applied treatments upregulated the gene expression and the maximum level was reached in the combined Si and NaCl application. It might be suggested that by activating the antioxidant system in the met1-7 mutant, silicon exerts its mitigating effect on NaCl stress. A similar situation was seen in the drm2-2 mutant as in Col-0. While there was no significant increase in gene expression when Si was applied alone, it was observed that NaCl application caused a significant increase, however, the maximum level was reached in combined Si and NaCl application. In the ros1-4 mutant, similar to drm2-2, Si application did not cause a significant change in CSD2 gene expression, while NaCl and combined Si and NaCl application appeared to upregulate gene expression. As a result, the increase in the amount of gene expression of CSD2 in all experimental groups confirms that Si has a mitigating effect on salt stress (Fig. 4).
For the CAT3 gene, which plays a role in another step of the antioxidant mechanism, it was observed that combined Si and NaCl application caused a significant increase in Col-0 plants compared to Si and NaCl applications. In the met1-7 mutant, it was observed that NaCl application significantly increased CAT3 gene expression compared to both Si and combined Si and NaCl applications. It was observed that Si and NaCl applications induced a significant increase in CAT3 expression in drm2-2 mutant compared to the control group. Si and NaCl combined tretament significantly down-regulated CAT3 gene expression compared to both Si and NaCl applied separately. In the ros1-4 mutant, similar to drm2-2, NaCl application caused a significant increase compared to the control group. It was observed that the level of CAT3 gene expression caused by Si application was significantly lower than the level of gene expression in NaCl and the combined Si and NaCl applications (Fig. 5).
Considering that Si application along with NaCl stress showed the alleviating effect of Si via upregulating CAT3 gene expression in the Col-0 plant.
For the APX1 gene, which is involved in another step of the antioxidant mechanism, similar to CAT3, Si application in Col-0 plants caused a significant decrease in gene expression level compared to control and NaCl application. It was also observed that the combined Si and NaCl application caused a significant decrease compared to the control group. In the ros1-4 mutant, it was seen that NaCl application caused a significant increase compared to the control group and combined Si and NaCl application. APX1 gene level showed a similar pattern in all experimental groups (Fig. 6).
4 Discussion
Silicon, the second most abundant element in soil, has mostly been researched in terms of its role in plant absorption and its potential protective mechanisms against various stressors (Epstein 1994; Thorne et al. 2020; Celayir et al. 2023). It was demonstrated that silicon exerts a mitigating effect on salt stress, however the underlying molecular mechanisms remain partially understood (Liang et al. 1996; Goto et al. 2003; Tahir et al. 2006, 2012; Parveen and Ashraf 2010; Bae et al. 2012; Yin et al. 2013). In plants, salt stress causes ionic stress by disrupting the ionic balance of Na+, Cl−, K+ and Ca2+ together with osmotic stress. An excessive amount of Na+ ions entering the cells is transported to other leaves, tissues and organs where it changes the metabolisms in various ways. This interference causes physiological and molecular effects in plants. Studies have shown that silicon reduces the effects of salt stress by increasing K+ uptake against excessive Na+ intake to the plant (Liang et al. 1996; Zuccarini 2008; Yang and Guo 2018; Liu et al. 2019). However, although the effect of salt stress on epigenetic mechanisms is mostly studied, the effects of Si on epigenetic mutants exposed to salt stress have not been studied. In this study, we tried to understand the physiological and molecular effects of Si on methylation mutants under salt stress. We selected three epigenetic mutants with ecotypes Col-0 as plant materials; these mutants were met1-7, drm2-2 and ros1-4 with loss of function in MET1, DRM2 and ROS1 enzymes by T-DNA insert, respectively.
Osmolytes majorly contribute to maintaining cellular osmotic adjustment through cell turgidity, protect internal cell components and reduce ionic toxicity. In most of the plants, osmolyte accumulation is a common response after exposure to salinity stress. They increase their osmolytes like proline and glycine betaine in order to ameliorate the devastating effects of stress (Singh et al. 2022). An increase in the amount of osmolytes may indicate that stress response mechanisms are actively working (Kosar et al. 2019). In previous studies, it was stated that Si application increased the amount of osmolyte and the osmotic potential (Yin et al. 2013; Farouk et al. 2020). Abbas et al. (2015) examined the alleviating effect of Si on salt stress in okra. Spraying Si on leaves in salinity conditions significantly increased osmolytes and caused an increase in osmotic adaptation capacity and antioxidant activity. In line with our research, Celayir et al. (2023) observed in their study on the effects of Si on UV stress that the application of Si led to a higher concentration of osmolytes compared to the plants that were not treated with Si. However, when UV-B and Si were applied together, the amount of osmolyte decreased, whereas in our results the amount of osmolyte increased. Although the application of Si silicon alone in the drm2-2 mutant causes a decrease in the amount of osmolyte, it confirms that Si applied together with NaCl in Col-0 and drm2-2 reveals its protective effect under stress. The fact that changes in the amount of osmolyte occur at different rates in the mutantsshows that the activation of biomolecules effective in the absorption of silicon and salt might be affected by regional methylation differences.
After stress exposure, ion homeostasis as well as cell membrane permeability of the cell change. This ion flux is often interrelated with accumulation of ROS in the cell. In previous studies using various plants, it has been reported that Si application causes improvement in ion leakage caused by salt stress (Liang et al. 1996; Agarie et al. 1998; Reezi et al. 2009; Kim et al. 2014). Zhu et al. (2004) applied 1 mM Si, 50 mM NaCl and combination treatment to varieties of cucumber and obtained similar data with our results in the ion leakage analysis they performed on the 5th and 10th days of culture. Abdelaal et al. (2020) stated that when Si was added to the medium, ion leakage decreased in sweet pepper plants, while ion leakage increased at different salt stress conditions. Compared to our findings, similar results were observed in the Col-0 plant and the demethylation mutant ros1-4. On the other hand, it was found that Si application under salt stress in methylation mutants met1-7 and drm2-2 further increases ion leakage. This can be interpreted as the loss of methylation prevents Si from affecting membrane permeability, regardless of which regions are methylated.
DNA methylation in plants plays a crucial role in the regulation of many epigenetic phenomena. These include transcriptional silencing of transposons and transgenes, defense against stress conditions, silencing of genes that control seed formation and leaf morphology at flowering time (Zangi et al. 2020). Tobiasz-Salach et al. (2023) examined the physiological, biochemical, and epigenetic effects of Si under salinity stress. In order to identify alterations in DNA methylation in maize plants treated with Si under salt stress, they employed the methylation-sensitive amplified polymorphism (MSAP). According to their findings, plants respond to salt stress by altering their DNA methylation level, and the effects of NaCl together Si application were dose-dependent. They came to the conclusion that these modifications might point to pathways for salt-stressed plant adaptation. In addition, they discovered that applying Si topically can lessen the detrimental effects of salt in the soil, which is in line with previous studies. In another study, the goal of Stadnik et al. (2022) was to assess how silicon foliar treatment affected the oats’ (Avena sativa L.) physiological and epigenetic response to salt stress. Similar to the study conducted by Tobiasz-Salach et al. (2023), the impact of Si administration during salt stress on the DNA methylation level in oat plants was examined using MSAP analysis. Similarly, their research showed that the oat plants’ ability to withstand salt was enhanced by the exogenous application of silicon. This study examined both the physiological and epigenetic responses of plants to the experimental stimuli used. The lowering level of methylation is influenced by the administration of Si under saline conditions. Moreover, Celayir et al. (2023) reported that 1 mM Si application to Arabidopsis Col-0 plant caused hypermethylation, while Yeşildirek et al. (2023) reported that 100 mM NaCl application to met1-7 mutant caused hypomethylation. Also, Yeşildirek et al. (2023) stated in their study that the met1-7 mutant might have supported the genes involved in the RdDM pathway to cope with the DNA methylation deficiency in the CG regions. The fact that the drm2-2 mutant which was in this study gave parallel results with the met1-7 mutant among the applied NaCl condition suggests that a similar situation may also be in question for the drm2-2 mutant. It was observed that the hypermethylation state caused by salt stress was reversed in all epigenetic mutants in the presence of salt stress and Si.
The enzymatic antioxidants that work the most as a defense system against ROS accumulation are SOD, CAT and APX1 enzymes (Maqsood et al. 2023). In previous studies on various plants, it has been shown that the application of Si together with salt causes an increase in antioxidant enzyme activities, leading to an improvement in ROS accumulation (Zhu et al. 2004; Alzahrani et al. 2018; Ahmad et al. 2019; Alamri et al. 2020; Hurtado et al. 2020; Hernández-Salinas et al. 2022). In parallel with our results, Ma et al. (2016) reported thatTaCAT andTaSOD genes expression levels in wheat were increased in high levels when Si was applied to drought-stressed plants than to Si alone was treated. In our study, while no significant change was observed in theAPX gene in the Col-0 plant under salt stress, it was observed that theAPX gene expression increased under salt stress in ros1-4 mutant. It is thought that the gene expression enhancing effect of the epigenetic mutants used in the study was observed due to methylation defects in the gene regions associated with CAT3 expression.
Global methylation analysis showed that combined Si and NaCl application eliminated the hypomethylation caused by NaCl. CSD2 gene expression increased more in Si-NaCl combined application than in NaCl stress alone, regardless of the difference in epigenetic regulation differences. While CAT3 gene expression did not differ in control under NaCl stress, it increased in all epigenetic mutants in the study. In Si-NaCl combined application, this increase was suppressed to a certain extent. The APX gene, on the other hand, increased in NaCl application in mutants, and this increase was suppressed when Si and NaCl were applied together.
Plant responses to stress factors are greatly influenced by DNA methylation, one of the main epigenetic regulation mechanisms. Changes in DNA methylation also enable plants to withstand adverse environmental conditions and develop ‘epigenetic memory’ for the next stress situation. With this study, it was observed that Si, which has previously proven its ameliorating effect against the physiological and morphological negative effects of salt stress, can provide the same effect in terms of global DNA methylation. This suggests that RNA-directed DNA methylation (RdDM) pathway-based regulation may play an important role in the Si-Stress-Plant interaction when methylation and demethylation enzymes do not work properly.
5 Conclusion
One of the primary epigenetic control systems, DNA methylation, has a significant impact on how plants respond to stressors. Findings reveal that applying Si to Arabidopsis methylation mutants during salt stress increases the resistance of plants to salty conditions, depending on the methylation level at the epigenetic level. This study sheds light on the mitigating properties of Si, specifically the methylation that occurs in response to separate and simultaneous exposure to Si and NaCl. This gives us proof that the Si-Stress-Plant relationship may be significantly regulated by the RNA-directed DNA methylation (RdDM) pathway. In conclusion, it was found that treating Arabidopsis epigenetic mutants with Si during salt stress could improve the plants’ ability to withstand salt on epigenetic level.Moreover, our research provides an epigenetic perspective, providing an additional contribution to the mechanisms by which Si promotes plant resilience in stressful environments.
Data Availability
The data for the current study is included in this published article.
Change history
25 June 2024
A Correction to this paper has been published: https://doi.org/10.1007/s42729-024-01905-8
References
Abbas T, Balal RM, Shahid MA, Pervez MA, Ayyub CM, Aqueel MA, Javaid MM (2015) Silicon-induced alleviation of NaCl toxicity in okra (Abelmoschus esculentus) is associated with enhanced photosynthesis, osmoprotectants and antioxidant metabolism. Acta Physiol Plant 37:1–15. https://doi.org/10.1007/s11738-014-1768-5
Abdelaal KA, Mazrou YS, Hafez YM (2020) Silicon foliar application mitigates salt stress in sweet pepper plants by enhancing water status, photosynthesis, antioxidant enzyme activity and fruit yield. Plants 9:6:733. https://doi.org/10.3390/plants9060733
Abdulkhair WM, Alghuthaymi MA (2016) Plant pathogens. https://doi.org/10.5772/65325. Plant growth 49
Abobatta WF (2020) Plant responses and tolerance to extreme salinity: learning from halophyte tolerance to extreme salinity. Salt Drought Stress Tolerance Plants: Signal Networks Adapt Mech 177–210. https://doi.org/10.1007/978-3-030-40277-8_7
Agarie S, Hanaoka N, Ueno O, Miyazaki A, Kubota F, Agata W, Kaufman PB (1998) Effects of silicon on tolerance to water deficit and heat stress in rice plants (Oryza sativa L.), monitored by electrolyte leakage. Plant Prod Sci 12:96–103. https://doi.org/10.1626/pps.1.96
Ahmad P, Ahanger MA, Alam P, Alyemeni MN, Wijaya L, Ali S, Ashraf M (2019) Silicon (Si) supplementation alleviates NaCl toxicity in mung bean [Vigna radiata (L.) Wilczek] through the modifications of physio-biochemical attributes and key antioxidant enzymes. J Plant Growth Regul 38:70–82. https://doi.org/10.1007/s00344-018-9810-2
Al-aghabary K, Zhu Z, Shi Q (2005) Influence of silicon supply on chlorophyll content, chlorophyll fluorescence, and antioxidative enzyme activities in tomato plants under salt stress. J Plant Nutr 2712:2101–2115. https://doi.org/10.1081/PLN-200034641
Alamri S, Hu Y, Mukherjee S, Aftab T, Fahad S et al (2020) Silicon-induced postponement of leaf senescence is accompanied by modulation of antioxidative defense and ion homeostasis in mustard (Brassica juncea) seedlings exposed to salinity and drought stress. Plant Physiol Biochem 157:47–59. https://doi.org/10.1016/j.plaphy.2020.09.038
Alzahrani Y, Kuşvuran A, Alharby HF, Kuşvuran S, Rady MM (2018) The defensive role of silicon in wheat against stress conditions induced by drought, salinity or cadmium. Ecotoxicol Environ Saf 154:187–196. https://doi.org/10.1016/j.ecoenv.2018.02.057
Asada K (1992) Ascorbate peroxidase-a hydrogen scavenging enzyme in plants. Physiol Plant 85:235–241. https://doi.org/10.1111/j.1399-3054.1992.tb04728.x
Ashraf M, Afzal M, Ahmed R, Mujeeb F, Sarwar A, Ali L (2010) Alleviation of detrimental effects of NaCl by silicon nutrition in salt-sensitive and salt-tolerant genotypes of sugarcane (Saccharum officinarum L). Plant Soil 3261:381–391. https://doi.org/10.1007/s11104-009-0019-9
Bae EJ, Lee KS, Huh MR, Lim CS (2012) Silicon significantly alleviates the growth inhibitory effects of NaCl in salt-sensitive ‘Perfection’and ‘Midnight’Kentucky bluegrass (Poa pratensis L). Hortic Environ Biotechnol 53:477–483. https://doi.org/10.1007/s13580-012-0094-3
Bakhat HF, Bibi N, Zia Z, Abbas S, Hammad HM, Fahad S, Ashraf MR, Shah GM, Rabbani F, Saeed S (2018) Silicon mitigates biotic stresses in crop plants: a review. Crop Prot 104:21–34. https://doi.org/10.1016/j.cropro.2017.10.008
Bharti P, Mahajan M, Vishwakarma AK et al (2015) AtROS1 overexpression provides evidence for epigenetic regulation of genes encoding enzymes of flavonoid biosynthesis and antioxidant pathways during salt stress in transgenic tobacco. J Exp Bot 6619:5959–5969. https://doi.org/10.1093/jxb/erv304
Böhmdorfer G, Rowley MJ, Kuciński J et al (2014) RNA-directed DNA methylation requires stepwise binding of silencing factors to long non‐coding RNA. Plant J 792:181–191. https://doi.org/10.1111/tpj.12563
Bose J, Rodrigo-Moreno A, Shabala S (2014) ROS homeostasis in halophytes in the context of salinity stress tolerance. J Exp Bot 655:1241–1257. https://doi.org/10.1093/jxb/ert430
Celayir T, Yeni O, Yeşildirek YV, Arıkan B, Kara NT (2023) Molecular effects of Silicon on Arabidopsis thaliana Seedlings under UV-B stress. Photochem Photobiol 996:1393–1399. https://doi.org/10.1111/php.13788
Chan SL, Henderson IR, Jacobsen SE (2005) Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat Rev Genet 65:351–360. https://doi.org/10.1038/nrg1601
Chinnusamy V, Zhu JK (2009) RNA-directed DNA methylation and demethylation in plants. Sci China Life Sci 524:331–343. https://doi.org/10.1007/s11427-009-0052-1
Dourado PRM, de Souza ER, Lins CMT et al (2019) Osmotic adjustment in cowpea plants: interference of methods for estimating osmotic potential at full turgor. Plant Physiol Biochem 145:114–119. https://doi.org/10.1016/j.plaphy.2019.10.020
Epstein E (1994) The anomaly of silicon in plant biology. Proc Natl Acad Sci USA 911:11–17. https://doi.org/10.1073/pnas.91.1.11
Farouk S, Elhindi KM, Alotaibi MA (2020) Silicon supplementation mitigates salinity stress on Ocimum basilicum L. via improving water balance, ion homeostasis, and antioxidant defense system. Ecotoxicol Environ Saf 206:111396. https://doi.org/10.1016/j.ecoenv.2020.111396
Gehring M, Reik W, Henikoff S (2009) DNA demethylation by DNA repair. Trends Genet 25(2):82–90. https://doi.org/10.1016/j.tig.2008.12.001
Goto M, Ehara H, Karita S et al (2003) Protective effect of silicon on phenolic biosynthesis and ultraviolet spectral stress in rice crop. Plant Sci 1643:349–356. https://doi.org/10.1016/S0168-9452(02)00419-3
Hernández-Salinas M, Valdez-Aguilar LA, Alia-Tejacal I et al (2022) Silicon enhances the tolerance to moderate NaCl-salinity in tomato grown in a hydroponic recirculating system. J Plant Nutr 453:413–425. https://doi.org/10.1080/01904167.2021.1963772
Hurtado AC, Chiconato DA, de Mello, Prado et al (2020) Different methods of silicon application attenuate salt stress in sorghum and sunflower by modifying the antioxidative defense mechanism. Ecotoxicol Environ Saf 203:110964. https://doi.org/10.1016/j.ecoenv.2020.110964
Ighodaro OM, Akinloye OA (2018) First line defence antioxidants-superoxide dismutase (SOD), catalase (CAT) and glutathione peroxidase (GPX): their fundamental role in the entire antioxidant defence grid. Alexandria J Med 544:287–293. https://doi.org/10.1016/j.ajme.2017.09.001
Isayenkov SV, Maathuis FJ (2019) Plant salinity stress: many unanswered questions remain. Front Plant Sci 10:435515. https://doi.org/10.3389/fpls.2019.00080
James RA, Blake C, Byrt CS, Munns R (2011) Major genes for Na+ exclusion, Nax1 and Nax2 (wheat HKT1; 4 and HKT1; 5), decrease Na+ accumulation in bread wheat leaves under saline and waterlogged conditions. J Exp Bot 62.8:2939–2947. https://doi.org/10.1093/jxb/err003
Kim YH, Khan AL, Waqas M et al (2014) Silicon application to rice root zone influenced the phytohormonal and antioxidant responses under salinity stress. J Plant Grow Regul 33:137–149. https://doi.org/10.1007/s00344-013-9356-2
Kim JS, Lim JY, Shin H, Kim BG, Yoo SD, Kim WT, Huh JH (2019) ROS1-dependent DNA demethylation is required for ABA-inducible NIC3 expression. Plant Physiol 179 (4): 1810–1821. https://doi.org/10.1104/pp.18.01471
Kong W, Li B, Wang Q, Wang B et al (2018) Analysis of the DNA methylation patterns and transcriptional regulation of the NB-LRR-encoding gene family in Arabidopsis thaliana. Plant Mol Biol 966:563–575. https://doi.org/10.1007/s11103-018-0715-z
Kosar F, Akram NA, Sadiq M, Al-Qurainy F, Ashraf M (2019) Trehalose: a key organic osmolyte effectively involved in plant abiotic stress tolerance. J Plant Grow Regul 38:606–618. https://doi.org/10.1007/s00344-018-9876-x
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 113:204–220. https://doi.org/10.1038/nrg2719
Li L, Wu W, Zhao Y, Zheng B (2017) A reciprocal inhibition between ARID1 and MET1 in male and female gametes in Arabidopsis. J Integr Plant Biol 59:9:657–668. https://doi.org/10.1111/jipb.12573
Liang Y, Shen Q, Shen Z, Ma T (1996) Effects of silicon on salinity tolerance of two barley cultivars. J Plant Nutr 191:173–183. https://doi.org/10.1080/01904169609365115
Liu B, Soundararajan P, Manivannan A (2019) Mechanisms of Silicon-mediated amelioration of salt stress in plants. Plants 8.9:307 https://doi.org/10.3390/plants8090307
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2– ∆∆CT method. Methods 25:4:402–408. https://doi.org/10.1006/meth.2001.1262
López Sánchez A, Stassen JH, Furci L et al (2016) The role of DNA (de) methylation in immune responsiveness of Arabidopsis. Plant J 883:361–374. https://doi.org/10.1111/tpj.13252
Luyckx M, Hausman JF, Lutts S, Guerriero G (2017) Silicon and plants: current knowledge and technological perspectives. Front Plant Sci 8:411. https://doi.org/10.3389/fpls.2017.00411
Ma D, Sun D, Wang C, Qin H, Ding H, Li Y, Guo T (2016) Silicon application alleviates drought stress in wheat through transcriptional regulation of multiple antioxidant defense pathways. J Plant Growth Regul 35:1–10. https://doi.org/10.1007/s00344-015-9500-2
Maqsood MF, Shahbaz M, Zulfiqar U et al (2023) Enhancing wheat growth and yield through salicylic acid-mediated regulation of Gas Exchange, Antioxidant Defense, and osmoprotection under salt stress. Stresses 31:372–386. https://doi.org/10.3390/stresses3010027
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 153:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Parveen N, Ashraf M (2010) Role of silicon in mitigating the adverse effects of salt stress on growth and photosynthetic attributes of two maize (Zea mays L.) cultivars grown hydroponically. Pak J Bot 423:1675–1684
Peck S, Mittler R (2020) Plant signaling in biotic and abiotic stress. J Exp Bot 715:1649–1651. https://doi.org/10.1093/jxb/eraa051
Prashanth SR, Sadhasivam V, Parida A (2008) Overexpression of cytosolic copper/zinc superoxide dismutase from a mangrove plant Avicennia marinain indica rice var Pusa Basmati-1 confers abiotic stress tolerance. Transgenic Res 17:281–291. https://doi.org/10.1007/s11248-007-9099-6
Rahnama A, James RA, Poustini K, Munns R (2010) Stomatal conductance as a screen for osmotic stress tolerance in durum wheat growing in saline soil. Funct Plant Biol 373:255–263. https://doi.org/10.1071/FP09148
Reezi S, Kalantari MBS, Okhovvat SM, Jeong BR (2009) Silicon alleviates salt stress, decreases malondialdehyde content and affects petal color of salt-stressed cut rose (Rosa Xhybrida L.) ‘Hot lady’. Afr J Biotechnol 88:1502–1508
Sáenz-Mata J, Jiménez-Bremont JF (2012) HR4 gene is induced in the Arabidopsis-Trichoderma atroviride beneficial interaction. Int J Mol Sci 137:9110–9128. https://doi.org/10.3390/ijms13079110
Saitoh Y, Mitani-Ueno N, Saito K, Matsuki K, Huang S et al (2021) Structural basis for high selectivity of a rice silicon channel Lsi1. Nat Commun 12(1):6236. https://doi.org/10.1038/s41467-021-26535-x
Sánchez-Romera B, Aroca R (2020) Plant roots the hidden half for investigating salt and drought stress responses and tolerance. Salt and Drought stress tolerance in plants: signaling networks and adaptive mechanisms. 137–175. https://doi.org/10.1007/978-3-030-40277-8_6
Savvas D, Giotis D, Chatzieustratiou E, Bakea M, Patakioutas G (2009) Silicon supply in soilless cultivations of zucchini alleviates stress induced by salinity and powdery mildew infections. Environ Exp Bot 651:11–17. https://doi.org/10.1016/j.envexpbot.2008.07.004
Shi Q, Bao Z, Zhu Z, Ying Q, Qian Q (2006) Effects of different treatments of salicylic acid on heat tolerance, chlorophyll fluorescence, and antioxidant enzyme activity in seedlings of Cucumis sativus L. Plant Growth Regul 48:127–135. https://doi.org/10.1007/s10725-005-5482-6
Shi Y, Wang Y, Flowers TJ, Gong H (2013) Silicon decreases chloride transport in rice (Oryza sativa L.) in saline conditions. J Plant Physiol 1709:847–853. https://doi.org/10.1016/j.jplph.2013.01.018
Siam HS, Abd El-Moez MR, Holah SS, Abou Zeid ST (2018) Effect of silicon addition to different fertilizer on yield of rice (Oryza sativa L.) plants. I-Macro nutrients by different rice parts. Middle East J Appl Sci 81:177–190
Singh D, Singh CK, Singh D, Sarkar SK, Prasad SK, Sharma NL, Singh I (2022) Glycine betaine modulates chromium (VI)-induced morpho-physiological and biochemical responses to mitigate chromium toxicity in chickpea (Cicer arietinum L.) cultivars. Sci Rep 12. https://doi.org/10.1038/s41598-022-11869-3..1:8005
Stadnik B, Tobiasz-Salach R, Mazurek M (2022) Effect of Silicon on Oat Salinity Tolerance: analysis of the epigenetic and physiological response of plants. Agriculture 13(1):81. https://doi.org/10.3390/agriculture13010081
Sudan J, Raina M, Singh R (2018) Plant epigenetic mechanisms: role in abiotic stress and their generational heritability. 3 Biotech. https://doi.org/10.1007/s13205-018-1202-6. 8.3:172
Tahir MA, Rahmatullah T, Aziz M, Ashraf S, Kanwal S, Maqsood MA (2006) Beneficial effects of silicon in wheat (Triticum aestivum L.) under salinity stress. Pak J Bot 385:1715–1722. https://doi.org/10.9755/ejfa.v12i1.5170
Tahir MA, Aziz T, Farooq M, Sarwar G (2012) Silicon-induced changes in growth, ionic composition, water relations, chlorophyll contents and membrane permeability in two salt-stressed wheat genotypes. Arch Agron Soil Sci 583:247–256. https://doi.org/10.1080/03650340.2010.518959
Thorne SJ, Hartley SE, Maathuis FJ (2020) Is silicon a panacea for alleviating drought and salt stress in crops? Front Plant Sci 1221. https://doi.org/10.3389/fpls.2020.01221
Tobiasz-Salach R, Mazurek M, Jacek B (2023) Physiological, biochemical, and epigenetic reaction of Maize (Zea mays L.) to cultivation in conditions of varying Soil Salinity and Foliar Application of Silicon. Int J Mol Sci 24:2:1141. https://doi.org/10.3390/ijms24021141
Tripathi DK, Singh S, Singh VP, Prasad SM, Dubey NK, Chauhan DK (2017) Silicon nanoparticles more effectively alleviated UV-B stress than silicon in wheat (Triticum aestivum) seedlings. Plant Physiol Biochem 110:70–81. https://doi.org/10.1016/j.plaphy.2016.06.026
Tuteja N (2007) Mechanisms of high salinity tolerance in plants. Meth Enzymol 428:419–438. https://doi.org/10.1016/S0076-6879(07)28024-3
Willekens H, Chamnongpol S, Davey M, Schraudner M et al (1997) Catalase is a sink for H2O2 and is indispensable for stress defence in C3 plants. EMBO J 1616:4806–4816. https://doi.org/10.1093/emboj/16.16.4806
Yang Y, Guo Y (2018) Elucidating the molecular mechanisms mediating plant salt-stress responses. New Phytol 2172:523–539. https://doi.org/10.1111/nph.14920
Yeşildirek YV, Arıkan B, Kara NT (2023) Epigenetic changes of Arabidopsis MET1 Cytosine methyltransferase mutants under salt stress. Plant Biosyst 1573:507–515. https://doi.org/10.1080/11263504.2023.2165567
Yin L, Wang S, Li J, Tanaka K, Oka M (2013) Application of silicon improves salt tolerance through ameliorating osmotic and ionic stresses in the seedling of Sorghum bicolor. Acta Physiol Plant 3511:3099–3107. https://doi.org/10.1007/s11738-013-1343-5
Zangi M, Najjar MBB, Golalipour M, Aghdasi M (2020) MET1 DNA methyltransferase controls TERT Gene expression: a new insight to the role of telomerase in Development. Cell J 221:71–74. https://doi.org/10.22074/CELLJ.2020.6290
Zhu JK (2009) Active DNA demethylation mediated by DNA glycosylases. Annu Rev Genet 43:143–166. https://doi.org/10.1146/annurev-genet-102108-134205
Zhu Z, Wei G, Li J, Qian Q, Yu J (2004) Silicon alleviates salt stress and increases antioxidant enzymes activity in leaves of salt-stressed cucumber (Cucumis sativus L). Plant Sci 1673:527–533. https://doi.org/10.1016/j.plantsci.2004.04.020
Zuccarini P (2008) Effects of silicon on photosynthesis, water relations and nutrient uptake of Phaseolus vulgaris under NaCl stress. Biol Plant 52:157–160. https://doi.org/10.1007/s10535-008-0034-3
Acknowledgements
We gratefully acknowledge the financial support of TÜBİTAK-1002 with project number 121Z605, TUBITAK BIDEB, 2221 with Application Number 1059B211800734, and The Research Fund of the Istanbul University with Project ID 33713.
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: An extraneous note appeared above authors’affiliations in the article as originally published and has been removed.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yeşildirek, Y.V., Arıkan, B., Çelik, H. et al. Role of Silicon in Mediating Salt Stress Responses in Arabidopsis Methylation Mutants. J Soil Sci Plant Nutr (2024). https://doi.org/10.1007/s42729-024-01848-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42729-024-01848-0