Abstract
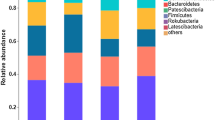
Long-term intensive greenhouse production commonly leads to continuous cropping obstacles and therefore declines the productivity of greenhouse system, soil microbiological mechanisms behind which remain poorly understood so far. Here, based on a continuous greenhouse cucumber cropping system, the differences in bacterial community structure of the soils with different cucumber cultivation history were assessed through high-throughput sequencing. The β diversity of bacterial community, but not α diversity, significantly changed as consecutive cucumber cultivation and also positively linked to cucumber yield. As for bacterial community members, at phylum level, prolonged cucumber cultivation increased average relative abundances of Chloroflexi and Gemmatimonadetes while decreasing average relative abundance of Nitrospirae; at genus level, continuous cucumber cultivation decreased average relative abundances of some beneficial microbes (i.e. Bacillus, Solirubrobacter, and Rubrobacter) and N-cycling related microbes (i.e. Nitrospira and Azoarcus) while increasing average relative abundance of some functional microbes (i.e. Agromyces, Thermomicrobium, Desulfotomaculum, Sphaerobacter, and Mycobacterium). Soil available phosphorus and organic carbon contents were the most important contributors to the shifts of bacterial community structure. Co-occurrence network analysis indicated long-term greenhouse cucumber cultivation weakened the interaction of species within bacterial community and led to less complex and connected organization among bacterial taxa. Overall, our results suggested that soil bacterial community structure and the interactions of bacterial taxa may play the important roles in the occurrence of continuous cropping obstacles in a long-term intensive greenhouse production system.











Similar content being viewed by others
References
Brennan EB, Acosta-Martine V (2017) Cover cropping frequency is the main driver of soil microbial changes during six years of organic vegetable production. Soil Biol Biochem 109:188–204
Bünemann EK, Bongiorno G, Bai ZG, Creamer RE, De Deyn G, de Goede R, Fleskens L, Geissen V, Kuyper TW, Mäder P, Pulleman M, Sukkel W, van Groenigen JW, Brussaard L (2018) Soil quality – a critical review. Soil Biol Biochem 120:105–125
Chaparro JM, Sheflin AM, Manter DK, Vivanco JM (2012) Manipulating the soil microbiome to increase soil health and plant fertility. Plant Soil 48:489–499
Chávez-Romero Y, Navarro-Noya YE, Reynoso-Martínez SC, Sarria-Guzmán Y, Govaerts B, Verhulst N, Dendooven L, Luna-Guido M (2016) 16S metagenomics reveals changes in the soil bacterial community driven by soil organic C, N-fertilizer and tillage-crop residue management. Soil Till Res 159:1–8
Chen T, Lin S, Wu LK, Lin WX, Sampietro DA (2015) Soil sickness: current status and future perspectives. Allelopathy J 36:167–196
Chen SC, Duan GL, Ding K, Huang FY, Zhu YG (2018) DNA stable-isotope probing identifies uncultivated members of Pseudonocardia associated with biodegradation of pyrene in agricultural soil. FEMS Microbiol Ecol 94(3):fiy026
Daims H, Lücker S, Wagner M (2016) A new perspective on microbes formerly known as nitrite-oxidizing bacteria. Trends Microbiol 24:699–712
de Miera LES, Arroyo P, de Luis CE, Falagán J, Ansola G (2014) High-throughput sequencing of 16S RNA genes of soil bacterial communities from a naturally occurring CO2 gas vent. Int J Greenh Gas Con 29:176–184
Dumbrell AX, Nelson M, Helgason T, Dytham C, Fitter AH (2010) Relative roles of niche and neutral processes in structuring a soil microbial community. ISME J 4:337–345
Fan FL, Yin C, Tang YJ, Li ZJ, Song AL, Wakelin SA, Zou J, Liang YC (2014) Probing potential microbial coupling of carbon and nitrogen cycling during decomposition of maize residue by 13C-DNA-SIP. Soil Biol Biochem 70:12–21
Fan FQ, Zhang BY, Morrill PL, Husain T (2017) Profiling of sulfate-reducing bacteria in an offshore oil reservoir using phospholipid fatty acid (PLFA) biomarkers. Water Air Soil Pollut 228:410
Fang H, Han LX, Cui YL, Xue YF, Cai L, Yu YL (2016) Changes in soil microbial community structure and function associated with degradation and resistance of carbendazim and chlortetracycline during repeated treatments. Sci Total Environ 572:1203–1212
Fang H, Zhang HP, Han LX, Mei JJ, Ge QQ, Long ZN, Yu YL (2018) Exploring bacterial communities and biodegradation genes in activated sludge from pesticide wastewater treatment plants via metagenomic analysis. Environ Pollut 243:1206–1216
Franke-Whittle IH, Manici LM, Blaz Stres HI (2015) Rhizosphere bacteria and fungi associated with plant growth in soils of three replanted apple orchards. Plant Soil 395:317–333
Fu HD, Zhang GX, Zhang F, Sun ZP, Geng GM, Li TL (2017) Effects of continuous tomato monoculture on soil microbial properties and enzyme activities in a solar greenhouse. Sustainability 9:317
Handelsman J, Stabb EV (1996) Biocontrol of soilborne plant pathogens. Plant Cell 8:1855–1869
Hargreaves SK, Williams RJ, Hofmockel KS (2015) Environmental filtering of microbial communities in agricultural soil shifts with crop growth. PLoS One 10(7):e0134345
Huang H, Chou C, Erickson R (2006) Soil sickness and its control. Allelopathy J 18:1–21
Huang LF, Song LX, Xia XJ, Mao WH, Shi K, Zhou YH, Yu JQ (2013) Plant-soil feedbacks and soil sickness: from mechanisms to application in agriculture. J Chem Ecol 39:232–242
Jia Y, Huang H, Zhong M, Wang FH, Zhang LM, Zhu YG (2013) Microbial arsenic methylation in soil and rice rhizosphere. Environ Sci Technol 47:3141–3148
Kumari S, Regar RK, Manickam N (2018) Improved polycyclic aromatic hydrocarbon degradation in a crude oil by individual and a consortium of bacteria. Bioresour Technol 254:174–179
Lal R (2015) Restoring soil quality to mitigate soil degradation. Sustainability 7:5875–5895
Lemanceau P, Maron PA, Mazurier S, Mougel C, Pivato B, Plassart P, Ranjard L, Revellin C, Tardy V, Wipf D (2015) Understanding and managing soil biodiversity: a major challenge in agroecology. Agron Sustain Dev 35:67–81
Li TL, Yang LJ (2016) Overcoming continuous cropping obstacles-the difficult problem. Sci Agr Sin 49(5):916–918
Li CG, Li XM, Kong WD, Wu Y, Wang JG (2010) Effect of monoculture soybean on soil microbial community in the Northeast China. Plant Soil 330:423–433
Lin HR, Shi JY, Chen XL, Yang JJ, Chen YX, Zhao YD, Hu TD (2010) Effects of lead upon the actions of sulfate-reducing bacteria in the rice rhizosphere. Soil Biol Biochem 42:1038–1044
Liu XZ, Zhang LM, Prosser JI, He JZ (2009) Abundance and community structure of sulfate reducing prokaryotes in a paddy soil of southern China under different fertilization regimes. Soil Biol Biochem 41:687–694
Liu X, Zhang JL, Gu TY, Zhang WM, Shen QR, Yin SX, Qiu HZ (2014) Microbial community diversities and taxa abundances in soils along a seven-year gradient of potato monoculture using high throughput pyrosequencing approach. PLoS One 9(1):e86610
Liu X, Zhang Y, Ren XJ, Chen BH, Shen CW, Wang F (2019) Long-term greenhouse vegetable cultivation alters the community structures of soil ammonia oxidizers. J Soils Sediments 19:883–902
Lu LH, Yin SX, Liu X, Zhang WM, Gu TY, Shen QR, Qiu HZ (2013) Fungal networks in yield-invigorating and -debilitating soils induced by prolonged potato monoculture. Soil Biol Biochem 65:186–194
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting rhizobacteria. Annu Rev Microbiol 63:541–556
Luo GW, Rensing C, Chen H, Liu MQ, Wang M, Guo SW, Ling N, Shen QR (2018) Deciphering the associations between soil microbial diversity and ecosystem multifunctionality driven by long-term fertilization management. Funct Ecol 32:1103–1116
Ma L, Deng FC, Yang C, Guo CL, Dang Z (2018) Bioremediation of PAH-contaminated farmland: field experiment. Environ Sci Pollut Res 25:64–72
Mariotte P, Mehrabi Z, Bezemer TM, De Deyn GB, Kulmatiski A, Drigo B, Veen GF, van der Heijden MGA, Kardol P (2018) Plant–soil feedback: bridging natural and agricultural sciences. Trends Ecol Evol 33:129–142
Mendes LW, Kuramae EE, Navarrete AA, van Veen JA, Tsai SM (2014) Taxonomical and functional microbial community selection in soybean rhizosphere. ISME J 8:1577–1587
Mendes LW, Raaijmakers JM, de Hollander M, Mendes R, Tsai SM (2018) Influence of resistance breeding in common bean on rhizosphere microbiome composition and function. ISME J 12:212–224
Mohanram S, Kumar P (2019) Rhizosphere microbiome: revisiting the synergy of plant-microbe interactions. Ann Microbiol 69:307–320
Morrieën E, Hannula SE, Snoek LB, Helmsing NR, Zweers H, de Hollander M, Soto RL, Bouffaud ML, Buée M, Dimmers W, Duyts H, Geisen S, Girlanda M, Griffiths RI, Jørgensen HB, Jensen J, Plassart P, Redecker D, Schmelz RM, Schmidt O, Thomson BC, Tisserant E, Uroz S, Winding A, Bailey MJ, Bonkowski M, Faber JH, Martin F, Lemanceau P, de Boer W, van Veen JA, van der Putten WH (2017) Soil networks become more connected and take up more carbon as nature restoration progresses. Nat Commun 8:14349
Mougi A, Kondoh M (2012) Diversity of interaction types and ecological community stability. Science 337:349–351
Pepe-Ranney C, Campbell AN, Koechli CN, Berthrong S, Buckley DH (2016) Unearthing the ecology of soil microorganisms using a high resolution DNA-SIP approach to explore cellulose and xylose metabolism in soil. Front Microbiol 7:703
Perez PG, Ye J, Wang S, Wang XL, Huang DF (2014) Analysis of the occurrence and activity of diazotrophic communities in organic and conventional horticultural soils. Appl Soil Ecol 79:37–48
Philippot L, Raaijmakers JM, Lemanceau P, van der Putten WH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11:789–799
Raaijmakers JM, Paulitz TC, Steinberg C, Alabouvette C, Moënne-Loccoz Y (2009) The rhizosphere: a playground and battlefield for soilborne pathogens and beneficial microorganisms. Plant Soil 321:341–361
Reinhold-Hurek B, Hurek T (1997) Azoarcus spp. and their interactions with grass roots. Plant Soil 194:57–64
Santoyo G, del Carmen O-MM, Govindappa M (2012) Mechanisms of biocontrol and plant growth-promoting activity in soil bacterial species of Bacillus and Pseudomonas: a review. Biocontrol Sci Tenhn 8:855–872
Shen ZZ, Xue C, Taylor PWJ, Ou YN, Wang BB, Zhao Y, Ruan YZ, Li R, Shen QR (2018) Soil pre-fumigation could effectively improve the disease suppressiveness of biofertilizer to banana Fusarium wilt disease by reshaping the soil microbiome. Biol Fert Soils 54:793–806
Siegel-Hertz K, Edel-Hermann V, Chapelle E, Terrat S, Raaijmakers JM, Steinberg C (2018) Comparative microbiome analysis of a Fusarium Wilt suppressive soil and a Fusarium Wilt conducive soil from the Châteaurenard Region. Front Microbiol 9:568
Soares RA, Roesch LFW, Zanatta G, de Oliveira Camargo FA, Passaglia LMP (2006) Occurrence and distribution of nitrogen fixing bacterial community associated with oat (Avena sativa) assessed by molecular and microbiological techniques. Appl Soil Ecol 33:221–234
Uroz S, Buée M, Murat C, Frey-Klett P, Martin F (2010) Pyrosequencing reveals a contrasted bacterial diversity between oak rhizosphere and surrounding soil. Env Microbiol Rep 2:281–288
Wang JJ, Shen J, Wu YC, Tu C, Soininen J, Stegen JC, He JZ, Liu XQ, Zhang L, Zhang EL (2013) Phylogenetic beta diversity in bacterial assemblages across ecosystems: deterministic versus stochastic processes. ISME J 7:1310–1321
Wang R, Zhang HC, Sun LG, Qi GF, Chen S, Zhao XY (2017) Microbial community composition is related to soil biological and chemical properties and bacterial wilt outbreak. Sci Rep 7:343
Wang L, You LX, Zhang JM, Yang T, Zhang W, Zhang ZX, Liu PX, Wu S, Zhao F, Ma J (2018) Biodegradation of sulfadiazine in microbial fuel cells: reaction mechanism, biotoxicity removal and the correlation with reactor microbes. J Hazard Mater 360:402–411
Wartiainen I, Eriksson T, Zheng WW, Rasmussen U (2008) Variation in the active diazotrophic community in rice paddy—nifH PCR-DGGE analysis of rhizosphere and bulk soil. Appl Soil Ecol 39:65–75
Wei Z, Yang T, Friman VP, Xu Y, Shen Q, Jousset A (2015) Trophic network architecture of root-associated bacterial communities determines pathogen invasion and plant health. Nat Commun 6:8413
Wei Z, Hu J, Gu YA, Yin SX, Xu YC, Jousset A, Shen QR, Friman VP (2018) Ralstonia solanacearum pathogen disrupts bacterial rhizosphere microbiome during an invasion. Soil Biol Biochem 118:8–17
Wu DY, Raymond J, Wu M, Chatterji S, Ren QH, Graham JE, Bryant DA, Robb F, Colman A, Tallon LJ, Badger JH, Madupu R, Ward NL, Eisen JA (2009) Complete genome sequence of the aerobic CO-oxidizing Thermophile Thermomicrobium roseum. PLoS One 4(1):e4207
Wu LK, Chen J, Xiao ZG, Zhu XG, Wang JY, Wu HM, Wu YH, Zhang ZY, Lin WX (2018) Barcoded pyrosequencing reveals a shift in the bacterial community in the rhizosphere and rhizoplane of Rehmannia glutinosa under consecutive monoculture. Int J Mol Sci 19:50
Xiong W, Li ZG, Liu HJ, Xue C, Zhang RF, Wu HS, Li R, Shen QR (2015a) The effect of long-term continuous cropping of black pepper on soil bacterial communities as determined by 454 pyrosequencing. PLoS One 10(8):e0136946
Xiong W, Zhao QY, Zhao J, Xun WB, Li R, Zhang RF, Wu HS, Shen QR (2015b) Different continuous cropping spans significantly affect microbial community membership and structure in a vanilla-grown soil as revealed by deep pyrosequencing. Microb Ecol 70:209–218
Xiong W, Guo S, Jousset A, Zhao QY, Wu HS, Li R, Kowalchuk GA, Shen QR (2017) Bio-fertilizer application induces soil suppressiveness against Fusarium wilt disease by reshaping the soil microbiome. Soil Biol Biochem 114:238–247
Xu N, Tan GC, Wang HY, Gai XP (2016) Effect of biochar additions to soil on nitrogen leaching, microbial biomass and bacterial community structure. Eur J Soil Biol 74:1–8
Xu QC, Dai RB, Ruan Y, Rensing C, Liu MQ, Guo SW, Ling N, Shen QR (2018) Probing active microbes involved in Bt-containing rice straw decomposition. Appl Microbiol Biot 102:10273–10284
Yang W, Jing XY, Guan YP, Zhai C, Wang T, Shi DY, Sun WP, Gu SY (2019) Response of fungal communities and co-occurrence network patterns to compost amendment in black soil of Northeast China. Front Microbiol 10:1562
Yao HY, Jiao XD, Wu FZ (2006) Effects of continuous cucumber cropping and alternative rotations under protected cultivation on soil microbial community diversity. Plant Soil 284:195–203
Yao ZY, Xing JJ, Gu HP, Wang HZ, Wu JJ, Xu JM, Brookes PC (2016) Development of microbial community structure in vegetable-growing soils from open-field to plastic-greenhouse cultivation based on the PLFA analysis. J Soils Sediments 16:2041–2049
Zhai WW, Wong MT, Luo F, Hashmi MZ, Liu XM, Edwards EA, Tang XJ, Xu JM (2017) Arsenic methylation and its relationship to abundance and diversity of arsM genes in composting manure. Sci Rep 7:42198
Zhang RF, Vivanco JM, Shen QR (2017) The unseen rhizosphere root-soil-microbe interactions for crop production. Curr Opin Microbiol 37:8–14
Zhao QY, Xiong W, Xing YZ, Sun Y, Lin XJ, Dong YP (2018) Long-term coffee monoculture alters soil chemical properties and microbial communities. Sci Rep 8:6116
Zhao XR, Wu HY, Song XD, Yang SH, Dong Y, Yang JL, Zhang GL (2019) Intra-horizon differentiation of the bacterial community and its co-occurrence network in a typical Plinthic horizon. Sci Total Envriron 678:692–701
Zhou XG, Wu FZ (2012) Dynamics of the diversity of fungal and Fusarium communities during continuous cropping of cucumber in the greenhouse. FEMS Microbiol Ecol 80:469–478
Zhou XG, Gao DM, Liu J, Qiao PL, Zhou XL, Lu HT, Wu X, Liu D, Jin X, Wu FZ (2014) Changes in rhizosphere soil microbial communities in a continuously monocropped cucumber (Cucumis sativus L.) system. Eur J Soil Biol 60:1–8
Funding
We received support for this research from the National Natural Science Foundation of China (grant number: 41601250).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, X., Li, Y., Ren, X. et al. Long-Term Greenhouse Cucumber Production Alters Soil Bacterial Community Structure. J Soil Sci Plant Nutr 20, 306–321 (2020). https://doi.org/10.1007/s42729-019-00109-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42729-019-00109-9