Abstract
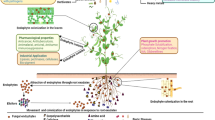
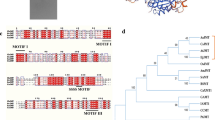
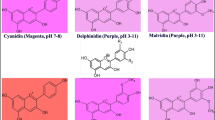
Phyllanthus emblica (L.) is one of the important medicinal plants possessing antioxidant, anti-tumor, hepatoprotective and immune-modulatory properties imparted by presence of high amount of flavonoid and vitamin C contents. Despite the medicinal value of P. emblica, molecular characterization of genes such as those involved in biosynthesis of flavonoids is lacking. Flavanone 3-hydroxylase (F3H) [EC 1.14.11.9] is an important regulatory enzyme of the flavonoid pathway which catalyzes the stereospecific hydroxylation of (2S)-naringenin to form (2R,3R)-dihydroflavonol. Herein, we isolated and investigated F3H gene from P. emblica, expressed functional recombinant proteins in microbial expression system and studied gene expression in different developmental stages. A full length PeF3H cDNA consisting of 1,469 bp (GenBank ID KF020698) has an open reading frame (ORF) of 1,095 bp, encoding a protein of 364 amino acids. Deduced amino acid analysis confirmed the presence of conserved amino acid residues which ligate ferrous iron and participates in 2-oxoglutarate binding (R-X-S). Three-dimensional protein structure modeling predicted that F3H had a jelly roll in the enzyme core, a typical structure shared by all 2-oxoglutarate-dependent dioxygenases including F3Hs. Phylogenetic analysis revealed that PeF3H was evolutionary closer F3H sequences of Triticum aestivum and Zea mays. Expression of PeF3H in an expression vector yielded a functional protein as HPLC analysis indicated conversion of naringenin to dihydrokaempferol after incubation with purified protein. Gene expression was analyzed in different developmental stages using q RT-PCR and was found to be highest in flowers as compared to leaves and fruits. In fruits, expression of PeF3H was increasing with the growth of the fruit.





Similar content being viewed by others
Abbreviations
- F3H:
-
Flavanone 3-hydroxylase
- RACE:
-
Rapid Amplification of cDNA Ends
- SMARTTM :
-
Switching Mechanism At 5′ End of RNA Transcript
- ORF:
-
Open Reading Frame
- BLAST:
-
Basic local alignment search tool
- GSP:
-
Gene specific primers.
References
Ahmed N, Maekawa M, Noda K (2009) Anthocyanin accumulation and expression pattern of anthocyanin biosynthesis gene in developing wheat coleoptiles. Biol Plant 53:223–228
Ahmed, R., Moushumi, S.J., Ahmed, H., Ali, M., Hasan, M., Reza, W.M.H., Haq, W.M., Jahan, R., Rahmatullah, M.: A Study of Serum Total Cholesterol and Triglyceride Lowering Activities of Phyllanthus Emblica L.(Euphorbiaceae) Fruits in Rats. Adv. Nat. Appl. Sci. 4: 168–170, 2010.
Aravind L, Koonin EV (2001) The DNA-repair protein AlkB, EGL-9, and leprecan define new families of 2-oxoglutarate-and iron-dependent dioxygenases. Genome Biol 2:1–0007
Aza-González C, Herrera-Isidrón L, Núñez-Palenius HG, De La Vega OM, Ochoa-Alejo N (2013) Anthocyanin accumulation and expression analysis of biosynthesis-related genes during chili pepper fruit development. Biol Plant 57:49–55
Britsch L, Grisebach H (1986) Purification and characterization of (2S)-flavanone 3-hydroxylase from Petunia hybrida. Eur J Biochem 156:569–577
Britsch L, Dedio J, Saedker H, Forkmann G (1993) Molecular characterization of flavanone 3 beta-hydroxylases. Consensus sequence, comparison with related enzymes and the role of conserved histidine residues. Eur J Biochem 217:745–754
Britsch L, Heller W, Grisebach H (1981) Conversion of flavanone to flavone, dihydroflavonol and flavonol with an enzyme system from cell cultures of parsley. Z. Naturforsch 36c:742–750
Britsch L, Ruhnau-Brich B, Forkmann G (1992) Molecular cloning, sequence analysis, and in vitro expression of flavanone 3 beta-hydroxylase from Petunia hybrida. J Biol Chem 267:5380–5387
Castellarin SD, Di Gaspero G, Marconi R, Nonis A, Peterlunger E, Paillard S, Adam-Blondon AF, Testolin R (2006) Colour variation in red grapevines (Vitis vinifera L.): genomic organisation, expression of flavonoid 3'-hydroxylase, flavonoid 3′,5′-hydroxylase genes and related metabolite profiling of red cyanidin-/blue delphinidin-based anthocyanins in berry skin. BMC Genomics 7:1–12
Davies KM (1993) A cDNA clone for flavanone 3-hydroxlase from Malus. Plant Physiol 103:291
Deboo, G.B., Albertsen, M.C., Taylor, L.P.: Flavanone 3-hydroxylase transcripts and flavonol accumulation are temporally coordinate in maize anthers. Plant J 7, 703–713
Decarolis E (1995) Deluca V (1994) 2-Oxoglutarate-dependent dioxygenase and related enzymes: biochemical characterization. Phytochemistry 36:1093–1107
Espley RV, Hellens RP, Putterill J, Stevenson DE, Kutty-Amma S, Allan AC (2007) Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J 49:414–427
Ghosal S (2000) Natural antioxidant compositions, method for obtaining same and cosmetic, pharmaceutical and nutritional formulations thereof. United States Patent No 6124268
Hausinger RP (2004) Fe(II)/alpha-ketoglutarate-dependent hydroxylases and related enzymes. Crit Rev Biochem Mol Biol 39:21–68
Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7:1071–1083
Jaakola L, Määttä K, Pirttilä AM, Törrönen R, Kärenlampi S, Hohtola A (2002) Expression of genes involved in anthocyanin biosynthesis in relation to anthocyanin, proanthocyanidin, and flavonol levels during bilberry fruit development. Plant Physiol 130:729–739
Judprasong K, Charoenkiatkul S, Thiyajai P, Sukprasansap M (2013) Nutrients and bioactive compounds of Thai indigenous fruits. Food Chem 140:507–512
Katsuyama Y, Funa N, Miyahisa I, Horinouchi S (2007) Synthesis of unnatural flavonoids and stilbenes by exploiting the plant biosynthetic pathway in Escherichia coli. Chem Biol 14:613–621
Kumar A, Singh K (2012) Isolation of High Quality RNA from Phyllanthus emblica and Its Evaluation by Downstream Applications. Mol Biotechnol 52:269–275
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Lo Piero AR, Puglisi I, Rapisarda P, Petrone G (2005) Anthocyanins accumulation and related gene expression in red orange fruit induced by low temperature storage. J Agri Food Sci 53:9083–9088
Lukacin R, Britsch L (1997) Identification of strictly conserved histidine and arginine residues as part of the active site in Petunia hybrida flavanone 3β-hydroxylase. Eur J Biochem 249:748–757
Martin C, Prescott A, Mackay S (1991) Bartlett. J, Vrijlandt, E.: Control of anthocyanin biosynthesis in flowers of Antirrhinum majus. Plant J 1:37–49
Meldgaard M (1992) Expression of chalcone synthase, dihydroflavonols reductase and flavanone-3-hydroxylase in mutants of barley deficient in anthocyanidin and proanthocyanidin biosynthesis. Theor Appl Genet 83:695–706
Morita Y, Saitoh M, Hoshino A, Nitasaka E, Iida S (2006) Isolation of cDNAs for R2R3-MYB, bHLH and WDR transcriptional regulators and identification of c and ca mutations conferring white flowers in the Japanese morning glory. Plant Cell Physiol 7:457–470
Patil SG, Deshmukh AA, Padol AR, Kale DB (2012) In vitro antibacterial activity of Emblica officinalis fruit extract by tube Dilution Method. Int J Toxicol Appl Pharmacol 2:49–51
Pelletier MK, Winkel-Shirley B (1996) Analysis of flavanone 3-hydroxylase in Arabidopsis seedlings. Plant Physiol 111:339–345
Prescott AG, John P (1996) Dioxygenases: molecular structure and role in plant metabolism. Annu Rev Plant Physiol 47:245–271
Prescott AG, Lloyd MD (2000) The iron (II) and 2-oxoacid-dependent dioxygenases and their role in metabolism. Nat Prod Rep 17:367–383
Punyasiri PAN, Abeysinghe ISB, Kumar V, Treutter D, Duy D, Gosch C, Martens S, Forkmann G, Fischer TC (2004) Flavonoid biosynthesis in the tea plant Camellia sinensis: properties of enzymes of the prominent epicatechin and catechin pathways. Arch Biochem Biophys 431:22–30
Quattrochio F, Wing JF, Leppen HTC, Mol JNM, Koes R (1993) Regulatory genes controlling anthocyanin pigmentation are functionally conserved among plant species and have distinct sets of target genes. Plant Cell 5:1497–1512
Quattrochio F, Wing JF, van der Woude K, Mol JNM, Koes R (1998) Analysis of bHLH and MYB domain proteins: species specific regulatory differences are caused by divergent evolution of target genes. Plant J 13:475–488
Rajak S, Banerjee SK, Sood S, Dinda AK, Gupta YK, Gupta SK, Maulik SK (2004) Emblica officinalis causes myocardial adaptation and protects against oxidative stress in ischemic‐reperfusion injury in rats. Phytotherapy Res 18:54–60
Roach PL, Clifton IJ, Fulop V, Harlos K, Barton GJ, Hajdu J, Andersson I, Schofield CJ, Baldwin JE (1995) Crystal structure of isopenicillin N synthase is the first from a new structural family of enzymes. Nature 375:700–704
Shen G, Pang Y, Wu W, Deng Z, Zhao L, Cao Y, Sun X, Tang K (2006) Cloning and Characterization of a Flavanone 3-Hydroxylase Gene from Ginkgo biloba. Biosci Rep 26:19–29
Singh K, Rani A, Kumar S, Sood P, Mahajan M, Yadav SK, Singh B, Ahuja PS (2008) An early gene of the flavonoid pathway, flavanone 3-hydroxylase, exhibits a positive relationship with the concentration of catechins in tea (Camellia sinensis). Tree Physiol 28:1349–1356
Sumalatha D (2013) Antioxidant and Antitumor activity of Phyllanthus emblica in colon cancer cell lines. Int J Curr Microbiol App Sci 2:189–195
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA 5: moleculare evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Wellmann F, Matern U, Lukacin R (2004) Significance of C-terminal sequence elements for Petunia flavanone 3beta-hydroxylase activity. FEBS Lett 561:149–154
Williams RJ, Spencer JPE, Rice-Evans C (2004) Flavonoids: antioxidants or signalling molecules? Free Rad Biol Med 36:838–849
Zhang YJ, Tanaka T, Iwamoto Y, Yang CR (2000) Kouno I (2000) Phyllaemblic acid, a novel highly oxygenated norbisabolane from the roots of Phyllanthus emblica. Tetrahedron Lett 41:1781–1784
Acknowledgments
Authors are thankful to Department of Science and Technology (DST) Govt. of India for their financial support to carry out this research work. Avneesh Kumar is also very thankful of Council of Scientific & Industrial Research (CSIR), India for providing scholarship during the research period.
Author information
Authors and Affiliations
Corresponding author
Additional information
Data archiving statement
The Sequence information of F3H has been submitted to NCBI Genbank database vide under accession no. KF020698
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PPTX 169 kb)
Rights and permissions
About this article
Cite this article
Kumar, A., Singh, B. & Singh, K. Functional characterization of flavanone 3-hydroxylase gene from Phyllanthus emblica (L.). J. Plant Biochem. Biotechnol. 24, 453–460 (2015). https://doi.org/10.1007/s13562-014-0296-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-014-0296-0