Abstract
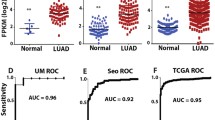
Tumorigenesis is a multistep process that attributes to the sequential accumulation of abnormal expression in key oncogenes or tumor suppressors. We aimed to identify stage-specific biomarkers to distinguish lung adenocarcinoma (LAC) stages in cancer progression. RNA-sequencing data of LAC and matched adjacent non-cancer tissues were downloaded from the Cancer Genome Atlas, including 29 pairs of samples from LAC at stage I, 14 from LAC at stage II, 13 from LAC at stage III, and 1 from LAC at stage IV. Differentially expressed genes (DEGs) were analyzed for each case at different stages of LAC. DEGs were further annotated based on transcription factor data information, tumor-associated gene database, and protein–protein interaction database. Functional annotation was performed for genes in PPI network by DAVID online tool. Our analysis identified 11 high-frequency DEGs in the stage I, 29 in the stage II, and 90 in the stage III of LAC. Among them, eight genes were significantly correlated with LAC stages and identified as biomarkers in LAC progression. ANGPTL5, C7orf16, EDN3, LOC150622, HOXA11AS, IL1F5, and USH1G significantly distinguished stage III from stages I and II. GJB6 was significantly enriched in the gap junction trafficking pathway, while C7orf16 and EDN3 were enriched in Wnt signaling pathway, cell cycle, and G protein-coupled receptor (GPCR) signaling. Up-regulated GJB6 especially in LAC stage II and down-regulated C7orf16 and EDN3 specifically in stage III were identified as biomarkers for distinguishing cancer stage in tumor progression through dysregulating gap junction, Wnt signaling, and GPCR signaling pathways.





Similar content being viewed by others
References
Subramanian J, Govindan R. Lung cancer in never smokers: a review. J Clin Oncol. 2007;25:561–70.
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893–917.
Kadara H, Kabbout M, Wistuba II. Pulmonary adenocarcinoma: a renewed entity in 2011. Respirology. 2012;17:50–65.
Zhang J, Ou JX, Bai CX. Tobacco smoking in China: prevalence, disease burden, challenges and future strategies. Respirology. 2011;16:1165–72.
Huang P, Cao K, Zhao H. Screening of critical genes in lung adenocarcinoma via network analysis of gene expression profile. Pathol Oncol Res. 2014.
Garber ME, Troyanskaya OG, Schluens K, Petersen S, Thaesler Z, Pacyna-Gengelbach M, et al. Diversity of gene expression in adenocarcinoma of the lung. Proc Natl Acad Sci U S A. 2001;98:13784–9.
Feldser DM, Kostova KK, Winslow MM, Taylor SE, Cashman C, Whittaker CA, et al. Stage-specific sensitivity to p53 restoration during lung cancer progression. Nature. 2010;468:572–5.
Kaushal V, Mukunyadzi P, Dennis RA, Siegel ER, Johnson DE, Kohli M. Stage-specific characterization of the vascular endothelial growth factor axis in prostate cancer: expression of lymphangiogenic markers is associated with advanced-stage disease. Clin Cancer Res. 2005;11:584–93.
Bhattacharjee A, Richards WG, Staunton J, Li C, Monti S, Vasa P, et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc Natl Acad Sci U S A. 2001;98:13790–5.
Chen X, Wang S-C, Cao L-H, Yang G-Q, Li M, Su J-C. Comparison between radial head replacement and open reduction and internal fixation in clinical treatment of unstable, multi-fragmented radial head fractures. Int Orthop. 2011;35:1071–6.
Tarazona S, Garcia-Alcalde F, Dopazo J, Ferrer A, Conesa A. Differential expression in RNA-seq: a matter of depth. Genome Res. 2011;21:2213–23.
Chen JS, Hung WS, Chan HH, Tsai SJ, Sun HS. In silico identification of oncogenic potential of fyn-related kinase in hepatocellular carcinoma. Bioinformatics. 2013;29:420–7.
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013;41:D808–15.
Kohl M, Wiese S, Warscheid B. Cytoscape: software for visualization and analysis of biological networks. In Data mining in proteomics. Springer; 201. pp. 291–303.
Da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Croft D, O’kelly G, Wu G, Haw R, Gillespie M, Matthews L, et al. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res. 2011;39:D691–7.
Solis LM, Behrens C, Raso MG, Lin HY, Kadara H, Yuan P, et al. Histologic patterns and molecular characteristics of lung adenocarcinoma associated with clinical outcome. Cancer. 2012;118:2889–99.
Li B, Ge Z, Song S, Zhang S, Yan H, Huang B, et al. Decreased expression of SOX7 is correlated with poor prognosis in lung adenocarcinoma patients. Pathol Oncol Res. 2012;18:1039–45.
Nguyen DX, Chiang AC, Zhang XH, Kim JY, Kris MG, Ladanyi M, et al. WNT/TCF signaling through LEF1 and HOXB9 mediates lung adenocarcinoma metastasis. Cell. 2009;138:51–62.
Basu M, Roy SS. Wnt/beta-catenin pathway is regulated by PITX2 homeodomain protein and thus contributes to the proliferation of human ovarian adenocarcinoma cell, SKOV-3. J Biol Chem. 2013;288:4355–67.
Barbazán J, Muinelo‐Romay L, Vieito M, Candamio S, Díaz‐López A, Cano A, et al. A multimarker panel for circulating tumor cells detection predicts patient outcome and therapy response in metastatic colorectal cancer. Int J Cancer. 2014.
Yaniv E, Borovsky Z, Mishan-Eisenberg G, Rachmilewitz J. Placental protein 14 regulates selective B cell responses. Cell Immunol. 2003;222:156–63.
Lee CL, Chiu PC, Lam KK, Chan RW, Chu IK, Koistinen R, et al. Glycodelin-A modulates cytokine production of peripheral blood natural killer cells. Fertil Steril. 2010;94:769–71.
Scholz C, Toth B, Brunnhuber R, Rampf E, Weissenbacher T, Santoso L, et al. Glycodelin A induces a tolerogenic phenotype in monocyte-derived dendritic cells in vitro. Am J Reprod Immunol. 2008;60:501–12.
Lee C-L, Lam KK, Koistinen H, Seppala M, Kurpisz M, Fernandez N, et al. Glycodelin-A as a paracrine regulator in early pregnancy. J Reprod Immunol. 2011;90:29–34.
Mishan-Eisenberg G, Borovsky Z, Weber MC, Gazit R, Tykocinski ML, Rachmilewitz J. Differential regulation of Th1/Th2 cytokine responses by placental protein 14. J Immunol. 2004;173:5524–30.
Lee C-L, Chiu PC, Lam KK, Siu S-O, Chu IK, Koistinen R, et al. Differential actions of glycodelin-A on Th-1 and Th-2 cells: a paracrine mechanism that could produce the Th-2 dominant environment during pregnancy. Hum Reprod. 2011;26:517–26.
Kunert-Keil C, Jeschke U, Simms G, Kasper M. Increased expression of glycodelin mRNA and protein in rat lungs during ovalbumin-induced allergic airway inflammation. Histochem Cell Biol. 2009;131:383–90.
Kunert-Keil C, Steinmüller F, Jeschke U, Gredes T, Gedrange T. Immunolocalization of glycodelin in human adenocarcinoma of the lung, squamous cell carcinoma of the lung and lung metastases of colonic adenocarcinoma. Acta Histochem. 2011;113:798–802.
Czyż J. The stage-specific function of gap junctions during tumourigenesis. Mol Cell Biochem. 2008;13:92–102.
Kar R, Batra N, Riquelme MA, Jiang JX. Biological role of connexin intercellular channels and hemichannels. Arch Biochem Biophys. 2012;524:2–15.
Cesen-Cummings K, Fernstrom MJ, Malkinson AM, Ruch RJ. Frequent reduction of gap junctional intercellular communication and connexin43 expression in human and mouse lung carcinoma cells. Carcinogenesis. 1998;19:61–7.
De Oliveira KD, Tedardi MV, Cogliati B, Dagli ML. Higher incidence of lung adenocarcinomas induced by DMBA in connexin 43 heterozygous knockout mice. Biomed Res Int. 2013;2013:618475.
Harada K, Nonaka T, Hamada N, Sakurai H, Hasegawa M, Funayama T, et al. Heavy-ion-induced bystander killing of human lung cancer cells: role of gap junctional intercellular communication. Cancer Sci. 2009;100:684–8.
Ozawa H, Mutai H, Matsunaga T, Tokumaru Y, Fujii M, Sakamoto K, et al. Promoted cell proliferation by connexin 30 gene transfection to head-and-neck cancer cell line. Anticancer Res. 2009;29:1981–5.
Princen F, Robe P, Gros D, Jarry-Guichard T, Gielen J, Merville M-P, et al. Rat gap junction connexin-30 inhibits proliferation of glioma cell lines. Carcinogenesis. 2001;22:507–13.
Sentani K, Oue N, Sakamoto N, Anami K, Naito Y, Aoyagi K, et al. Upregulation of connexin 30 in intestinal phenotype gastric cancer and its reduction during tumor progression. Pathobiology. 2010;77:241–8.
Klaus A, Birchmeier W. Wnt signalling and its impact on development and cancer. Nat Rev Cancer. 2008;8:387–98.
Teng Y, Wang XW, Wang YW, Wang J. [Effect of siRNA-mediated beta-catenin gene on Wnt signal pathway in lung adenocarcinoma A549 cell]. Zhonghua Yi Xue Za Zhi. 2010;90:988–92.
Delgado AP, Brandao P, Chapado MJ, Hamid S, Narayanan R. Open reading frames associated with cancer in the dark matter of the human genome. Cancer Genomics Proteomics. 2014;11:201–13.
Rennie C, Hulme H, Fisher P, Hall L, Agaba M, Noyes H, et al. A systematic, data-driven approach to the combined analysis of microarray and QTL data. 2010.
Cao Y, He M, Gao Z, Peng Y, Li Y, Li L, et al. Activating hotspot L205R mutation in PRKACA and adrenal Cushing’s syndrome. Science. 2014;344:913–7.
Welch A, Jacobs M, Wingo C, Cain B. Early progress in epigenetic regulation of endothelin pathway genes. Br J Pharmacol. 2013;168:327–34.
Kuzumaki N, Suzuki A, Narita M, Hosoya T, Nagasawa A, Imai S, et al. Multiple analyses of G-protein coupled receptor (GPCR) expression in the development of gefitinib-resistance in transforming non-small-cell lung cancer. PLoS One. 2012;7:e44368.
Samanta D, Kaufman J, Carbone DP, Datta PK. Long-term smoking mediated down-regulation of Smad3 induces resistance to carboplatin in non-small cell lung cancer. Neoplasia. 2012;14:644–55.
Acknowledgments
This study was supported by National Natural Science Foundation of China (grant No. 81372632).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liang, J., Lv, J. & Liu, Z. Identification of stage-specific biomarkers in lung adenocarcinoma based on RNA-seq data. Tumor Biol. 36, 6391–6399 (2015). https://doi.org/10.1007/s13277-015-3327-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3327-0