Abstract
Background
DNA tandem repeats (TRs) are often abundant and occupy discrete regions in eukaryotic genomes. These TRs often cause or generate chromosomal rearrangements, which, in turn, drive chromosome evolution and speciation. Tracing the chromosomal distribution of TRs could therefore provide insights into the chromosome dynamics and speciation among closely related taxa. The basic chromosome number in the genus Senna is 2n = 28, but dysploid species like Senna tora have also been observed.
Objective
To understand the dynamics of these TRs and their impact on S. tora dysploidization.
Methods
We performed a comparative fluorescence in situ hybridization (FISH) analysis among nine closely related Senna species and compared the chromosomal distribution of these repeats from a cytotaxonomic perspective by using the ITS1-5.8S-ITS2 sequence to infer phylogenetic relationships.
Results
Of the nine S. tora TRs, two did not show any FISH signal whereas seven TRs showed similar and contrasting patterns to other Senna species. StoTR01_86, which was localized in the pericentromeric regions in all S. tora, but not at the nucleolar organizer region (NOR) site, was colocalized at the NOR site in all species except in S. siamea. StoTR02_7_tel was mostly localized at chromosome termini, but some species had an interstitial telomeric repeat in a few chromosomes. StoTR05_180 was distributed in the subtelomeric region in most species and was highly amplified in the pericentromeric region in some species. StoTR06_159 was either absent or colocalized in the NOR site in some species, and StoIGS_463, which was localized at the NOR site in S. tora, was either absent or localized at the subtelomeric or pericentromeric regions in other species.
Conclusions
These data suggest that TRs play important roles in S. tora dysploidy and suggest the involvement of 45S rDNA intergenic spacers in “carrying” repeats during genome reshuffling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Repetitive elements (REs) comprise a considerable portion of plant genomes, even comprising > 85 % in some plant genomes (Schnable et al. 2009). Although considered ‘junk’ in the past, REs are now known as important players in genome function and structure, and species evolution (Fedoroff 2012; Wicker et al. 2007). REs are generated or produced by chromosomal rearrangements, which drive chromosome structure variations between closely related lineages (Murat et al. 2017; Schubert and Lysak 2011). Tracking the dynamics of various repeat families could therefore provide insights into the genome history of closely related taxa (Long et al. 2013; Waminal et al. 2018b).
REs can be categorized into two classes: tandem repeats (TRs) and dispersed repeats (Kubis et al. 1998). TRs follow a head-to-tail organization, whereas direct repeats, such as transposable elements. TRs can be further classified into three groups based on repeat unit length: microsatellites (2–5 bp repeats), minisatellites (6–100 bp repeats) and satellite DNA (satDNAs) (150–400 bp monomer length) (Mehrotra and Goyal 2014).
TRs are often distributed in distinct chromosomal regions, usually at pericentromeric and subtelomeric sites (Sharma et al. 2013), often in low copies forming a “library” of repeats (Fedoroff and Bennetzen 2013; Ruiz-Ruano et al. 2016). Due to sequence homologies in these chromosomal regions, they have been considered as hotspots for chromosomal rearrangements during genomic perturbations such as those resulting from a genome merger (Hartley and O’Neill 2019; Rosato et al. 2018; Schubert and Lysak 2011).
Often, these chromosomal rearrangements could amplify a single or few TR families, and when reproductive barriers are formed, new species may develop with an altered chromosomal number, organization, or TR abundance (Mandáková and Lysak 2018; Murat et al. 2017). Several basic chromosome numbers are present in a genus, which was described as dysploid (Winterfeld et al. 2020). Ascending dysploids, or species with more chromosomes, are formed when chromosomes are fragmented and when fragments maintain or develop centromeres. In contrast, descending dysploids are formed when chromosomes fuse (Winterfeld et al. 2020). As the differential abundance and distribution of TRs among species in a taxonomic group can vary (Perumal et al. 2017), TRs are used as cytotaxonomic markers to study phylogenetic relationships (Guerra 2008, 2012).
Fluorescence in situ hybridization (FISH) of TRs can provide essential information for understanding genome structure, chromosome evolution and phylogenetic relationships among related taxa (Iovene et al. 2008; Matsuda and Chapman 1995). This information complements the inference of phylogenetic relationships based on DNA markers such as the internal transcribed spacers (ITS) of 45S rDNA (Kim et al. 2015), which are used extensively because of their abundance of phylogenetically informative sites and easy amplification in the plant species (Farah et al. 2018). In addition, the intergenic spacers (IGS) of the 45S rDNA coding genes are known to “carry” different TRs during species evolution (Falquet et al. 1997; Almeida et al. 2012). Comparing the distribution of duplicated sequences in the S. tora IGS could provide important data for studying Senna evolution.
The genus Senna, formerly Cassia (family Fabaceae, subfamily Caesalpinioideae) comprises approximately 350 species of herbs, shrubs, and trees (Tucker 1996; Monkheang et al. 2011). Senna has varied economic and medicinal applications, and has been used not only as a natural pesticide but also for treating skin diseases, gastrointestinal disorders, and inflammation (Ongchai et al. 2019; Singh et al. 2013). A desire to further exploit these health benefits has prompted the genome sequencing of Senna tora, as a widespread and representative species, to better understand Senna biology and evolution.
The predominant chromosome number of the genus Senna is 2n = 28 (Rice et al. 2015), but species with descending dysploid karyotypes of 2n = 22–26 have also been identified, such as S. tora with 2n = 26 (Cordeiro and Felix 2018; Pellerin et al. 2019). To understand the dysploidization in S. tora, we examined its genome TR composition in our previous work (Waminal et al. 2021). We identified eight S. tora TRs, many of which showed unusual chromosomal distributions. For example, StoTR02_7_tel, which is the Arabidopsis-type telomeric repeat, showed highly amplified loci in the pericentromeric regions in all S. tora chromosomes in addition to signals at chromosome termini. Besides, StoTR05_180, which was observed in the subtelomeric region of all S. occidentalis chromosomes was absent in these regions but highly amplified in the pericentromeric regions in S. tora chromosomes.
The classification of Senna is not yet fully understood, and no comparative cytogenetics have been performed to examine chromosome evolution in the genus. Here, to understand the dynamics of S. tora TRs and their role in S. tora dysploidy, we performed comparative FISH among nine Senna species. We analyzed the chromosomal distribution patterns of these TRs in a cyto-phylogenetic context based on the ITS1-5.8S-ITS2 sequences.
Materials and methods
Plant material
Root tips were collected from germinated seeds of nine Senna species, which were provided by the National Plant Germplasm System (USDA, USA), Department of Herbal Crop Research (NIHHS, RDA, Korea), and Rare Palm Seeds (Germany). Roots were treated with 2 mM 8-hydroxyquinoline for 5 h at 18 °C to arrest cells at metaphase, and then fixed in Carnoy’s solution and stored in 70 % ethanol until used for chromosome preparation.
Chromosome preparation
Chromosome spreads were prepared according to Waminal et al. (2012) and Eliazar et al. (2019). Briefly, the meristematic tips were immersed in an enzyme solution consisting of 1 % pectolyase Y-23 (Duchefa Biochemie, Netherlands) and 2 % cellulase R-10 (Duchefa Biochemie, Netherlands) for 60–90 min at 37 °C. Next, chilled Carnoy’s solution was added, and after centrifugation, the precipitate was pipetted and dropped onto pre-warmed glass slides in a humid chamber at 70 °C and then air-dried.
Probes preparation and fluorescence in situ hybridization (FISH)
S. tora TRs were identified using low-coverage sequences and short-read clustering with TAREAN (Novák et al. 2017) in our previous study (Waminal et al. 2018a, 2021). All pre-labeled oligonucleotide probes (PLOPs) used in this study are listed in Table 1.
For the FISH procedure, a total of 40 µL hybridization mixture containing 100 % formamide, 50 % dextran sulfate, 20× SSC, 50 % dextran sulfate, 20× SSC, 50 ng/µL of each probe, and Sigma water were added to each slide and then denatured at 80 °C and kept in a humid chamber at 37 °C overnight. After hybridization, the slides were washed with 2× SSC at room temperature (RT) and then dehydrated through ethanol 70 %, 90 %, and 100 % for 3 min each. Finally, the slides were counterstained with a DAPI-Vectashield solution and captured under Cytovision ver 7.2 software with an Olympus BX53 fluorescence microscope system, equipped with a Leica DFC365 FS CCD camera. The images were enhanced using Adobe Photoshop CS6. Chromosomes were measured using the IdeoKar 1.2 software (Mirzaghaderi and Marzangi 2015), and chromosome typing was performed according to the method of Levan et al. (1964). Chromosomes were arranged in pairs depending on their FISH signals, length, and other chromosome features as described in our previous work (Pellerin et al. 2019; Youn and Kim 2018).
Genomic DNA extraction and sequencing of ITS sequences
Genomic DNA of nine Senna species was extracted from young leaves by using the cetyltrimethylammonium bromide (CTAB) method (Allen et al. 2006). The ITS primer pairs (F: GTCGCTCCTACCGATTGAA; R: TCTTTTCCTCCGCTTATTGA) were designed based on the 45S rDNA sequence of S. tora by using Primer3 software. PCR was performed by initial denaturation at 95 °C for 3 min, followed by 30 cycles of denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s, and extension at 72 °C for 30 s, followed by a final extension at 72 °C for 5 min. Amplicons were ethanol purified and sequenced by Bionics (South Korea).
Phylogenetic analysis
The Senna ITS sequences were aligned using CLC Main Workbench version 5.5, then confirmed through MEGA X. Genetic distance was calculated using the Kimura 2 parameter formula (K2P), and the phylogenetic tree was constructed by MEGA X.
Results
Six of the eight S. tora TRs were detected in the nine Senna species
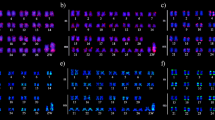
All Senna species in this study had 2n = 28 chromosomes (Fig. 1). In contrast, S. tora has only 2n = 26 chromosomes (Pellerin et al. 2019). To understand the impact of TRs in the karyotype dysploidy of S. tora, we performed FISH to analyze the presence or absence of signals and variations in chromosomal distributions to examine the dynamics of TRs identified in the S. tora genome.
FISH of S. tora TRs on root metaphase chromosomes of the nine Senna species. Six of the eight TRs, which showed signals from initial FISH screening, and StoIGS_463, are shown here. White arrows indicate the ITRs in S. candolleana, S. corymbosa, S. floribunda, S. occidentalis, and S. multiglandulosa. For signal patterns of individual probes, see supplementary data. Scale bar = 10 µm
Out of the eight S. tora TRs (Waminal et al. 2021), two were not detected by FISH in all nine Senna species (StoTR03_178 and StoTR04_55), while the other six were either present in all or only a few species (Table 2). StoTR02_7_tel, StoTR05_180, Sto_5S, and Sto_45S_CDS were detected in all species, whereas StoTR01_86 and StoTR06_159 were observed only in eight and four species, respectively (Fig. 1; Table 1). In addition, StoIGS_463 was also observed in only four species.
StoTR02_7_Tel and StoTR05_180 were mostly colocalized
The StoTR02_7_Tel, which is an Arabidopsis-type telomere repeat sequence, was highly amplified in the interstitial regions of S. tora chromosomes in addition to the canonical sites at chromosome termini (Pellerin et al. 2019; Waminal et al. 2021). Likewise, StoTR05_180 was also mostly colocalized with StoTR02_7_Tel in S. tora in pericentromeric regions but absent in subtelomeric sites.
In the present study, all nine Senna species had StoTR02_7_Tel signals at the terminal region of all chromosomes (Fig. 1). However, in addition to these canonical sites, interstitial telomeric repeat (ITR) signals were also detected in S. candolleana, S. corymbosa, S. floribunda, S. multiglandulosa, and S. occidentalis (Figs. 2, 3 and 4; Table 2). In S. candolleana, chromosomes 1 and 7 had ITRs in the long arm. In S. corymbosa, ITRs were detected in the proximal region of the short arm of chromosomes 1, 2, and 3. In S. floribunda, three chromosomes had ITRs in the short arm; those in chromosomes 6 and 13 were in proximal regions while those in chromosome 11 were more interstitial. In S. occidentalis, ITRs were located at proximal regions in the short arm of chromosomes 1, 3, and 4.
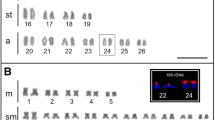
FISH karyograms of the root metaphase chromosomes of the nine Senna species. StoTR01_86 was mostly colocalized at 45S rDNA sites in all species except S. siamea. StoTR05_180 was at subtelomeric sites in all species except in S. sulfurea, where it was localized at pericentromeric regions. The white arrows indicate the ITR in S. candolleana, S. corymbosa, S. floribunda, S. occidentalis, and S. multiglandulosa. The orange arrow shows the weak signal of StoTR01_86 in S. didymobotrya. For karyogram of each species showing individual TR distribution, see supplementary data. Scale bar = 10 µm
Karyotype idiograms showing the distribution of S. tora TRs in nine Senna species. Note the colocalization of StoTR01_86 and StoTR06_159 at the 45S rDNA loci in some species, and the unique distribution of StoTR05_180 at the pericentromeric regions of all chromosomes in S. sulfurea, contrary to subtelomeric location in other species
Meanwhile, StoTR05_180 signals were detected as colocalized signals with StoTR02_7_Tel in all species, except in S. sulfurea (Figs. 2, 3 and 4). In S. sulfurea, StoTR05_180 did not colocalize with StoTR02_7_Tel. StoTR02_7_Tel was exclusively distributed at chromosome termini in S. sulfurea, whereas StoTR05_180 was absent in these regions but was amplified in the pericentromeric regions with varied intensities in all chromosomes (Fig. 2).
The number of 45S rDNA loci was more diverse than that of 5S rDNA
Similar to S. tora, all nine species had only one pair of Sto_5S rDNA, whereas Sto_45S rDNA varied from one to four loci (Table 1). The 5S rDNA signals were localized in the short arms of the respective chromosomes in S. alata, S. candolleana, S. corymbosa, S. multiglandulosa, S. sulfurea, and S. siamea, and in the long arms in S. didymobotrya, S. floribunda, and S. occidentalis (Figs. 2, 3; Table 2).
Cyto-phylogenetic analysis among Senna species. The ten species, including S. tora, were grouped into four (I–IV). StoTR05_180 at the subtelomeric site is shared in all ten, and likely the primitive distribution pattern in, Senna. StoTR01_86 was shared by species in Groups II–IV. Independent chromosomal rearrangements involving several repeats and chromosomes have taken place in species Groups II–IV. Species in Group IV shared rearrangements involving the displacement of StoTR05_180 to the pericentromeric regions of chromosomes (pink box)
The Sto_45S_CDS signals were distributed only in the subtelomeric regions of the short arms of the respective chromosomes. One pair of 45S rDNA was detected in S. corymbosa, S. floribunda, S. multiglandulosa, and S. occidentalis, two in S. candolleana, three in S. alata, S. siamea, and S. sulfurea, and four in S. didymobotrya (Figs. 2, 5; Table 2).
StoTR01_86 and StoTR06_159 colocalized at the 45S rDNA loci
The chromosomal distribution of StoTR01_86 in S. tora was in total contrast with that observed in the Senna species in this study. Although StoTR01_86 was localized in the pericentromeric regions in all S. tora chromosomes (Waminal et al. 2021), it was absent in the pericentromeric region in all currently investigated Senna species but colocalized at all 45S rDNA loci, except in S. siamea, which did not show any StoTR01_86 signal at all (Figs. 2, 3 and 4; Table 2).
On the other hand, StoTR06_159 was colocalized with 45S rDNA in S. tora, and also in S. alata, S. corymbosa, S. floribunda, and S. multiglandulosa (Figs. 2, 3 and 4; Table 2). However, the minor extra-NOR StoTR06_159 locus observed in S. tora was not detected in species with StoTR06_159 signals. None of the other species showed any StoTR06_159 signals (Table 2).
StoIGS_463 localized at extra-NOR loci
StoIGS_463 is a 463 bp duplicated sequence with two copies identified in the IGS region of S. tora 45S rDNA (Waminal et al. 2021). FISH also confirmed the exclusive localization of StoIGS_463 at the NOR site in S. tora. However, FISH on the nine Senna species showed no signals at NOR sites, but rather at subtelomeric or interstitial chromosomal regions in some species and were completely absent in some species (Figs. 2, 3; Table 2). While some distinct paired signals were observed, dispersed signals were also detected in many chromosomes, such as those in S. corymbosa, indicative of transposable element FISH signals.
Of the nine species, only S. alata, S. candolleana, S. corymbosa, and S. floribunda showed StoIGS_463 FISH signals (Figs. 2, 3 and 4; Table 2). Aside from chromosomes without signals, three StoIGS_463 distribution patterns were observed in these four species: (1) at the subtelomeric and interstitial regions, (2) only in the subtelomeric regions, and (3) only in the interstitial regions. In S. alata, there were ten, one, and three chromosomes that had the first, second, and third StoIGS_463 distribution patterns, respectively (Fig. 4; Table 2). In S. candolleana, only the first and third patterns were observed with eleven and three chromosomes bearing each pattern, respectively (Fig. 4; Table 2). In S. corymbosa, chromosome 4 did not show any StoIGS_463 signal, while four, five, and four chromosomes showed patterns i, ii and iii, respectively (Fig. 4; Table 2). Finally, in S. floribunda, there were 11, two, and one chromosomes that showed patterns i, ii and iii, respectively (Fig. 4; Table 2).
Chromosome rearrangement patterns support ITS-based phylogenetic tree
Since many TR loci formed from chromosomal rearrangements are not readily visible by FISH when these loci are shorter than the FISH detection threshold, grouping species based solely on FISH signal patterns could be misleading. Therefore, we compared the FISH distribution with the phylogenetic tree inferred using the entire ITS sequences (ITS1-5.8S-ITS2) of the Senna species, including S. tora data from our previous work (Waminal et al. 2021).
The Senna ITS length ranged from 632 bp in S. tora to 663 bp in S. corymbosa and had a mean of 649 bp. The ITS1 and ITS2 had relatively higher GC contents than those of 5.8S, and the GC content of the entire sequence ranged from 58.54–62.84 % and averaged 61.45 % (Table 3).
These ITS sequences divided the Senna species into four (Fig. 4). Group I comprised only S. siamea, which showed the fewest FISH signals of the S. tora TRs, and the most primitive chromosomal distribution pattern, StoTR05_180, at the subtelomeric regions of all chromosomes. Groups II–IV all had StoTR01_86 signals, mostly at the 45S rDNA loci. Independent chromosomal rearrangements involving the other repeats have taken place in the species in these groups. The absence of FISH signals may either be short arrays that make these loci undetectable using FISH or indicate a lack of actual rearrangements. Additional chromosomal rearrangements involving StoTR05_180 occurred in Group IV. These rearrangements displaced the subtelomeric location of StoTR05_180 into the pericentromeric region, where they have been highly amplified in both S. sulfurea and S. tora. Additional rounds of rearrangements may have occurred in S. tora, as shown by several chromosomes with interstitial StoTR05_180 signals.
Discussion
In our study, we have compared the chromosomal distribution of the TRs identified from the S. tora genome with nine other Senna species in order to understand the impacts and evolutionary dynamics of these TRs in the dysploidization of the S. tora karyotype. Compared with the nine Senna species assessed in this study, S. tora had the most extensive chromosomal rearrangements and the only one with a descending dysploidy karyotype (Fig. 4), suggesting the involvement of these TRs in shaping the extant S. tora genome.
Interstitial telomeric repeats are evidence for Senna chromosomal rearrangements
In most eukaryotes, telomeric repeats are usually located at the terminal end of chromosomes and have a key function in preventing chromosomal damage (Muraki et al. 2012). However, telomeric repeats have also been found in interstitial regions, also known as interstitial telomeric repeats (ITRs) (Bolzan 2012). Several mechanisms have been postulated to explain the formation of ITRs. These include interchromosomal telomere fusion, DNA polymerase slippage, and double-strand break repair (Uchida et al. 2002; Lin and Yan 2008).
Of the ten Senna species, including S. tora, six had readily detectable ITR signals indicating shared chromosomal rearrangements involving StoTR02_7_Tel among these Senna species. The lack of ITR signals in the other species may indicate fixation and reduction of these repeat loci, rendering them undetectable by FISH, as has been previously observed in other plants (He et al. 2013). It is also possible that these StoTR02_7_Tel-mediated chromosomal rearrangements occurred independently in each species. The extreme contrast of the StoTR02_7_Tel distribution between S. siamea and S. tora suggests that S. siamea has a more primitive karyotype and that the massive chromosomal rearrangements in S. tora may have evolved relatively recently.
Concerted conversion of StoTR05_180 to (peri)centromeric TRs in S. sulfurea and S. tora
Species in Groups I–III carried StoTR05_180 in the subtelomeric regions, whereas those in Group IV (S. sulfurea and S. tora) have somehow managed to transpose these TR loci from the subtelomeric to the pericentromeric region in all chromosomes. We proposed the hypothesized that this transposition has been occurred involving all chromosomes of the group IV ancestral karyotype. As a result, the evidence indicated that StoTR05_180 presents in the pericentromeric region of all chromosomes in S. sulfurea and S. tora. The other scenario is likely given the relatively random process of chromosomal rearrangement events.
One possible mechanism for this concerted TR array transposition may involve chromoplexy, which is a massive chromosomal rearrangement event that involves several chromosomes (Comai and Tan 2019; Pellestor and Gatinois 2020). Microhomologies between telomeric and (peri)centromeric regions make these regions hotspots for chromosomal inversions (He et al. 2013).
While both S. sulfurea and S. tora had lost subtelomeric StoTR05_180 loci but amplified pericentromeric loci, only S. tora developed a novel centromeric repeat, StoTR03_178 (Fig. 5), which may have likely evolved from StoTR05_180 (Waminal et al. 2021). Disruption of the epigenetic makeup immediately after chromosomal rearrangements may have enabled StoTR05_180 to function as a novel centromeric repeat. This, in S. tora, may have eventually caused centromere positioning, and fixation of StoTR03_178 variants.
The 45S rDNA IGS as repeat carrier during chromosomal rearrangements
45S rDNA is associated with genome rearrangements (Havlová et al. 2016). The sub-repeat elements in the IGS are considered important players in the dynamics of IGS (Jo et al. 2011). The elimination and reorganization of the IGS repeat elements were observed in Nicotiana tabacum after allopolyploidization which caused IGS length variation among Nicotiana species (Volkov et al. 1999). Although there is no exact mechanism to clearly explain the movement of TRs in and out of the 45S rDNA IGS, some authors have observed that TRs in the IGS moved out and amplified in another chromosomal region (Almeida et al. 2012). However, others also noted the opposite direction of IGS TR evolution in Phaseolus vulgaris, such that TRs moved into the 45S rDNA IGS from another region in the genome (Falquet et al. 1997). Our data suggest that the 45S rDNA IGS can act as TR “carrier” during chromosomal rearrangements, and that fragments of transposed TRs could sometimes get lodged and fixed in a taxon. Moreover, this observation suggests that TRs in the IGS are more like “footprints” of the recent involvement of the IGS in chromosomal rearrangements.
StoTR01_86, which was at the 45S rDNA locus in most Senna species except S. siamea and S. tora, suggests that 45S rDNA is involved in chromosomal rearrangement after the divergence of the Senna species from the ancestral karyotype S. siamea, which resulted in the fixation of StoTR01_86 in Group II-III species. Eventually, chromosomal rearrangement specific to S. tora may have caused more recent expulsion of the StoTR01_86 from the IGS and moved it into the pericentromeric regions of all chromosomes (Fig. 4) (Waminal et al. 2021).
StoTR06_159 also illustrates the movement of TRs in and out of the 45S rDNA IGS (Fig. 4). Moreover, StoIGS_463 explicitly showed the active role of the 45S rDNA IGS in homing repeats, which in this case is the subtelomeric ancestral repeat of StoIGS_463.
FISH data suggest the involvement of TRs in S. tora dysploidy
In Senna, the basic chromosome number is x = 14 (Waminal et al. 2021). Several species, however, have x = 11, 12, 13 resulting from descending dysploidization events (Elaine et al. 2005). Although various Senna species showed FISH signals from S. tora TRs, only S. tora had extensive genome rearrangements and dysploid karyotype (Fig. 4), suggesting the involvement of TRs in S. tora karyotype dysploidy.
Moreover, StoTR03_178 was observed in the interstitial region of S. tora chromosome 7, suggesting a relatively recent chromosome fusion (Waminal et al. 2021). Although we are not yet certain which ancestral chromosomes fused to form the S. tora chromosome 7, it is likely that orthologous S. sulfurea chromosomes 1 and 4 may have been involved. This hypothesis is based on the presence of subtelocentric S. sulfurea chromosomes 1 and 4, which were not detected in S. tora (Fig. 3). Moreover, subtelocentric chromosomes are often involved in interchromosomal fusion (Schubert and Lysak 2011).
Conclusions
We compared the chromosomal distribution of the eight S. tora TRs and a 45S rDNA IGS duplicated sequence, StoIGS_463, among nine Senna species. We have seen the dynamics of these TRs, which showed both shared and independent evolution. Importantly, we have shown cytogenetic evidence of the bidirectionality of TR movement into or out of the 45S rDNA IGS region, suggesting a role for 45S rDNA as a repeat carrier during chromosomal rearrangements. Moreover, these data also provide cytogenetic visualization of the expansion, contraction, and reorganization of repeat families in a repeat “library” of a lineage. Further studies should focus on characterizing the 45S rDNA IGS sequences of all Senna species to gain more insight into the role of the IGS in genome rearrangements.
References
Allen GC, Flores-Vergara MA, Krasynanski S, Kumar S, Thompson WF (2006) A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1:2320–2325. https://doi.org/10.1038/nprot.2006.384
Almeida C, Fonsêca A, dos Santos KGB, Mosiolek M, Pedrosa-Harand A (2012) Contrasting evolution of a satellite DNA and its ancestral IGS rDNA in Phaseolus (Fabaceae). Genome 55:683–689. https://doi.org/10.1139/g2012-059
Bolzan AD (2012) Chromosomal aberrations involving telomeres and interstitial telomeric sequences. Mutagenesis 27:1–15. https://doi.org/10.1093/mutage/ger052
Comai L, Tan EH (2019) Haploid induction and genome instability. Trends Genet TIG 35:791–803. https://doi.org/10.1016/j.tig.2019.07.005
Cordeiro JMP, Felix LP, Fabaceae (2018) Intra- and interspecific karyotypic variations of the genus Senna Mill (Caesalpinioideae). Acta Bot Bras 32:128–134. https://doi.org/10.1590/0102-33062017abb0274
Elaine B, Miotto STS, Schifino-Wittmann MT, Castro BD (2005) Cytogenetics and cytotaxonomy of Brazilian species of Senna Mill. (Cassieae - Caesalpinioideae - Leguminosae). Caryologia 58:152–163. https://doi.org/10.1080/00087114.2005.10589445
Eliazar AP, Waminal NE, Kim T-H, Kim HH (2019) FISH karyotype comparison between wild and cultivated Perilla species using 5S and 45S rDNA probes. Plant Breed Biotechnol 7:237–244. https://doi.org/10.9787/PBB.2019.7.3.237
Falquet J, Ceusot F, Dron M (1997) Molecular analysis of Phaseolus vulgaris rDNA unit and characterization of a satellite DNA homologous to IGS subrepeats. Plant Physiol Biochem 35:611–622
Farah AH, Lee SY, Gao Z, Yao TL, Madon M, Mohamed R (2018) Genome size, molecular phylogeny, and evolutionary history of the tribe Aquilarieae (Thymelaeaceae), the natural source of agarwood. Front Plant Sci 9:712. https://doi.org/10.3389/fpls.2018.00712
Fedoroff NV (2012) Transposable elements, epigenetics, and genome evolution. Science 338:758–767. https://doi.org/10.1126/science.338.6108.758
Fedoroff NV, Bennetzen JL (2013) Transposons, genomic shock, and genome evolution. Plant transposons and genome dynamics in evolution. John Wiley & Sons, Ltd, pp 181–201
Guerra M (2008) Chromosome numbers in plant cytotaxonomy: concepts and implications. Cytogenet Genome Res 120:339–350. https://doi.org/10.1159/000121083
Guerra M (2012) Cytotaxonomy: the end of childhood. Plant Biosyst Int J Deal Asp Plant Biol 146:703–710. https://doi.org/10.1080/11263504.2012.717973
Hartley G, O’Neill RJ (2019) Centromere repeats: hidden gems of the genome. Genes 10:223. https://doi.org/10.3390/genes10030223
Havlová K, Dvořáčková M, Peiro R, Abia D, Mozgová I, Vansáčová L, Gutierrez C, Fajkus J (2016) Variation of 45S rDNA intergenic spacers in Arabidopsis thaliana. Plant Mol Biol 92:457–471. https://doi.org/10.1007/s11103-016-0524-1
He L, Liu J, Torres GA, Zhang H, Jiang J, Xie C (2013) Interstitial telomeric repeats are enriched in the centromeres of chromosomes in Solanum species. Chromosome Res 21:5–13. https://doi.org/10.1007/s10577-012-9332-x
Iovene M, Wielgus S, Simon P, Buell C, Jiang J (2008) Chromatin structure and physical mapping of chromosome 6 of potato and comparative analyses with tomato. Genetics 180:1307–1317. https://doi.org/10.1534/genetics.108.093179
Jo S-H, Park H-M, Kim S-M, Kim HH, Hur C-G, Choi D (2011) Unraveling the sequence dynamics of the formation of genus-specific satellite DNAs in the family solanaceae. Heredity 106:876–885. https://doi.org/10.1038/hdy.2010.131
Kim SJ, Cho KS, Yoo KO, Lim KB, Hwang YJ, Chang DC, Kim KS (2015) Sequence analysis of the internal transcribed spacer (ITS) region of the nuclear ribosomal DNA (nrDNA) Chrysanthemum species in Korea. Hortic Environ Biotechnol 56:44–53. https://doi.org/10.1007/s13580-015-0085-2
Kubis S, Schmidt T, Heslop-Harrison JS (PAT) (1998) Repetitive DNA elements as a major component of plant genomes. Ann Bot 82:45–55. https://doi.org/10.1006/anbo.1998.0779
Levan A, Fredga K, Sandberg AA (1964) Nomenclature for centromeric position on chromosomes. Hereditas 52:201–220. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Lin KW, Yan J (2008) Endings in the middle: current knowledge of interstitial telomeric sequences. Mutat Res Mutat Res 658:95–110. https://doi.org/10.1016/j.mrrev.2007.08.006
Long Q, Rabanal FA, Meng D, Huber CD, Farlow A, Platzer A, Zhang Q, Vilhjálmsson BJ, Korte A, Nizhynska V et al (2013) Massive genomic variation and strong selection in Arabidopsis thaliana lines from Sweden. Nat Genet 45:884–890. https://doi.org/10.1038/ng.2678
Mandáková T, Lysak MA (2018) Post-polyploid diploidization and diversification through dysploid changes. Curr Opin Plant Biol 42:55–65. https://doi.org/10.1016/j.pbi.2018.03.001
Matsuda Y, Chapman VM (1995) Application of fluorescence in situ hybridization in genome analysis of the mouse. Electrophoresis 16:261–272. https://doi.org/10.1002/elps.1150160142
Mehrotra S, Goyal V (2014) Repetitive sequences in plant nuclear DNA: types, distribution, evolution and function. Genomics Proteomics Bioinformatics 12:164–171. https://doi.org/10.1016/j.gpb.2014.07.003
Mirzaghaderi G, Marzangi K (2015) IdeoKar: an ideogram constructing and karyotype analyzing software. Caryologia 68:31–35. https://doi.org/10.1080/00087114.2014.998526
Monkheang P, Sudmoon R, Tanee T, Noikotr K, Bletter N, Chaveerach A (2011) Species diversity, usages, molecular markers and barcode of medicinal Senna species (Fabaceae, Caesalpinioideae) in Thailand. J Med Plants Res 5:6173–6181. https://doi.org/10.5897/JMPR11.1075
Muraki K, Nyhan K, Han L, Murnane JP (2012) Mechanisms of telomere loss and their consequences for chromosome instability. Front Oncol 2:135. https://doi.org/10.3389/fonc.2012.00135
Murat F, Armero A, Pont C, Klopp C, Salse J (2017) Reconstructing the genome of the most recent common ancestor of flowering plants. Nat Genet 49:490–496. https://doi.org/10.1038/ng.3813
Novák P, Ávila Robledillo L, Koblížková A, Vrbová I, Neumann P, Macas J (2017) TAREAN: a computational tool for identification and characterization of satellite DNA from unassembled short reads. Nucleic Acids Res 45:e111. https://doi.org/10.1093/nar/gkx257
Ongchai S, Chokchaitaweesuk C, Kongdang P, Chomdej S, Buddhachat K (2019) In vitro chondroprotective potential of Senna alata and Senna tora in porcine cartilage explants and their species differentiation by DNA barcoding-high resolution melting (Bar-HRM) analysis. PLOS ONE 14:e0215664. https://doi.org/10.1371/journal.pone.0215664
Pellerin RJ, Waminal NE, Kim HH (2019) FISH mapping of rDNA and telomeric repeats in 10 Senna species. Hortic Environ Biotechnol 60:253–260. https://doi.org/10.1007/s13580-018-0115-y
Pellestor F, Gatinois V (2020) Chromoanagenesis: a piece of the macroevolution scenario. Mol Cytogenet 13:3. https://doi.org/10.1186/s13039-020-0470-0
Perumal S, Waminal NE, Lee J, Lee J, Choi BS, Kim HH, Grandbastien M-A, Yang TJ (2017) Elucidating the major hidden genomic components of the A, C, and AC genomes and their influence on Brassica evolution. Sci Rep 7:17986. https://doi.org/10.1038/s41598-017-18048-9
Rice A, Glick L, Abadi S, Einhorn M, Kopelman NM, Salman-Minkov A, Mayzel J, Chay O, Mayrose I (2015) The chromosome counts database (CCDB) – a community resource of plant chromosome numbers. New Phytol 206:19–26. https://doi.org/10.1111/nph.13191
Rosato M, Álvarez I, Feliner GN, Rosselló JA (2018) Inter- and intraspecific hypervariability in interstitial telomeric-like repeats (TTTAGGG)n in Anacyclus (Asteraceae). Ann Bot 122:387–395. https://doi.org/10.1093/aob/mcy079
Ruiz-Ruano FJ, López-León MD, Cabrero J, Camacho JPM (2016) High-throughput analysis of the satellitome illuminates satellite DNA evolution. Sci Rep 6:28333. https://doi.org/10.1038/srep28333
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F, Pasternak S, Liang C, Zhang J, Fulton L, Graves TA et al (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115. https://doi.org/10.1126/science.1178534
Schubert I, Lysak MA (2011) Interpretation of karyotype evolution should consider chromosome structural constraints. Trends Genet TIG 27:207–216. https://doi.org/10.1016/j.tig.2011.03.004
Sharma A, Wolfgruber TK, Presting GG (2013) Tandem repeats derived from centromeric retrotransposons. BMC Genom 14:142. https://doi.org/10.1186/1471-2164-14-142
Singh SK, Singh S, Yadav A (2013) A review on Cassia species: pharmacological, traditional and medicinal aspects in various countries. Am J Phytomedicine Clin Ther 1:291–312
Tucker SC (1996) Trends in evolution of floral ontogeny in Cassia sensu stricto, Senna, and Chamaecrista (Leguminosae: Caesalpinioideae: Cassieae: Cassiinae); a study in convergence. Am J Bot 83:687–711. https://doi.org/10.1002/j.1537-2197.1996.tb12758.x
Uchida W, Matsunaga S, Sugiyama R, Kawano S (2002) Interstitial telomere-like repeats in the Arabidopsis thaliana genome. Genes Genet Syst 77:63–67. https://doi.org/10.1266/ggs.77.63
Volkov RA, Borisjuk NV, Panchuk II, Schweizer D, Hemleben V (1999) Elimination and rearrangement of parental rDNA in the allotetraploid Nicotiana tabacum. Mol Biol Evol 16:311–320. https://doi.org/10.1093/oxfordjournals.molbev.a026112
Waminal NE, Park HM, Ryu KB, Kim JH, Yang TJ, Kim HH (2012) Karyotype analysis of Panax ginseng C.A.Meyer, 1843 (Araliaceae) based on rDNA loci and DAPI band distribution. Comp Cytogenet 6:425–441. https://doi.org/10.3897/CompCytogen.v6i4.3740
Waminal NE, Pellerin RJ, Kim N-S, Jayakodi M, Park JY, Yang TJ, Kim HH (2018a) Rapid and efficient FISH using pre-labeled oligomer probes. Sci Rep 8:8224. https://doi.org/10.1038/s41598-018-26667-z
Waminal NE, Perumal S, Liu S, Chalhoub B, Kim HH, Yang TJ, Liu S, Snowdon R, Chalhoub B (2018b) Quantity, distribution, and evolution of major repeats in Brassica napus. In: Liu S, Snowdon R, Chalhoub B (eds) The Brassica napus Genome. Springer International Publishing, Cham, pp 111–129
Waminal NE, Pellerin RJ, Kang S-H, Kim HH (2021) Chromosomal mapping of tandem repeats revealed massive chromosomal rearrangements and insights into Senna tora dysploidy. Front Plant Sci 12:629898. https://doi.org/10.3389/fpls.2021.629898
Wicker T, Sabot F, Hua-Van A, Bennetzen JL, Capy P, Chalhoub B, Flavell A, Leroy P, Morgante M, Panaud O et al (2007) A unified classification system for eukaryotic transposable elements. Nat Rev Genet 8:973–982. https://doi.org/10.1038/nrg2165
Winterfeld G, Ley A, Hoffmann MH, Paule J, Röser M (2020) Dysploidy and polyploidy trigger strong variation of chromosome numbers in the prayer-plant family (Marantaceae). Plant Syst Evol 306:36. https://doi.org/10.1007/s00606-020-01663-x
Youn SM, Kim HH (2018) Chromosome karyotyping of Senna covesii and S. floribunda based on triple-color FISH mapping of rDNAs and telomeric repeats. Plant Breed Biotechnol 6:51–56. https://doi.org/10.9787/PBB.2018.6.1.51
Acknowledgements
This study was funded by a grant from the National Research Foundation of Korea (NRF 2017R1A2B2004778).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ta, T.D., Waminal, N.E., Nguyen, T.H. et al. Comparative FISH analysis of Senna tora tandem repeats revealed insights into the chromosome dynamics in Senna. Genes Genom 43, 237–249 (2021). https://doi.org/10.1007/s13258-021-01051-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01051-w