Abstract
This study reports the isolation of myxobacteria from soil collected from plains in north India. Based on the morphology and 16S rDNA sequence, the isolated myxobacteria were identified as Corallococcus sp., Pyxidicoccus sp., Myxococcus sp., Cystobacter sp. and Archangium sp. The myxobacteria were functionally characterized to assess their ability to produce antibacterial and anticancer metabolites. The isolates were found to be functionally versatile as they produced extracellular bioactive molecules that exhibited high frequency of activities against Bacillus cereus, Mycobacterium smegmatis, Enterobacter cloacae and Pseudomonas syringae. The strains also showed cytotoxic activity against the human cancer cell lines of liver, pancreas, prostrate, bone and cervix. These results indicate the importance of isolating diverse strains of myxobacteria from unexplored habitats to find novel bioactive compounds. Moreover, the bioactive molecules explored in this study are predominantly hydrophilic compounds, obviating the limitations of solubility-related aspect of drug discovery.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For decades, microbes have been the major contributor to natural products for drug discovery and development. The microbial natural products being used as drugs are: antibacterial agents, such as the penicillins, cephalosporins, aminoglycosides, tetracyclines and various polyketides; cholesterol lowering agents, such as mevastatin and lovastatin; immunosuppressive agents, such as the cyclosporins and rapamycin; and anthelmintics and antiparasitic drugs, such as the ivermectins (Buss and Waigh 1995). The emergence of antibiotic-resistant pathogenic microorganisms and rapid development of resistance to chemotherapeutic drugs have necessitated the discovery of structurally diverse and mechanistically distinct antimicrobial and anticancer compounds. Most of the microbial bioactive compounds discovered so far originated from actinomycetes, but the focus is currently on the less exploited microbes as viable sources of secondary metabolites; viz., marine bacteria, cyanobacteria, endosymbionts and myxobacteria. Myxobacteria are aerobic Gram-negative, unicellular, rod-shaped δ-proteobacteria, distributed in a wide range of habitats including soil, bark of trees, decaying plant materials, and freshwater and marine environment (Ravenschlag et al. 1999; Reichenbach 1999; Garcia et al. 2009; Dawid 2000). They have a very large genome size compared to other bacteria, and a number of structurally unique biologically active secondary metabolites are produced by this group of bacteria (Wenzel and Muller 2009; Han et al. 2013; Reichenbach 2001). Many compounds isolated from myxobacteria act on unique cellular targets, which, at the time of their discovery, were not targeted by other secondary metabolites (Bode and Muller 2006; Herrmann et al. 2017).
Myxobacteria have been the source of several new antibacterials, e.g., thuggacins from Sorangium cellulosum (active against Mycobacterium tuberculosis), crocacin from Chondromyces crocatus (active against Gram-positive bacteria), indiacens from Sandaracinus amylolyticus (mildly inhibit Gram-positive and Gram-negative bacteria), disciformycins from Pyxidicoccus fallax (active against methicillin- and vancomycin-resistant Staphylococcus aureus), Coralmycins (potent activity against Gram-negative bacteria), etc. (Irschik et al. 2007; Kunze et al. 1994; Steinmetz et al. 2012; Surup et al. 2014; Kim et al. 2016). Several new antifungal compounds have also been reported from myxobacteria: pedein A and B, miuraenamides, aurafurons A and B and cyrmenins (Kunze et al. 2005, 2008; Iizuka et al. 2006; Sasse et al. 2003). A recent assessment of the antibacterial compounds form myxobacteria has revealed that most of these compounds are active against Gram-positive bacteria (Schaberle et al. 2014).
Several secondary metabolites produced by myxobacteria are promising anticancer agents. Epothilone, a paclitaxel mimetic, obtained from Sorangium cellulosum So ce56, has been approved for breast cancer treatment (Burris 2008). In 2007, ixabepilone, an analog of epothilone B, was approved by FDA for the treatment of metastatic or locally advanced taxane-resistant breast cancer (Tan and Toppmeyer 2008). Tubulysins, produced by Archangium gephyra Ar 315, Angiococcus disciformis An d48 and Cystobacter sp. SBCb004, are potent growth inhibitors compared to taxol, epothilone and vincristine and are effective in multidrug-resistant cell lines (Sasse et al. 2000; Steinmetz et al. 2004; Chai et al. 2010). Argyrin A and cruentaren A produced by Archangium gephyra and Byssovorax cruenta, respectively, are highly cytotoxic against various human tumor cell lines (Nickeleit et al. 2008; Kunze et al. 2007). Recently, nannocystin from Nannocystis sp. has been found to show activity against various cancer cell lines by targeting eukaryotic translation (Krastel et al. 2015).
Although myxobacteria are an attractive source of new chemical class of compounds, they have not been fully exploited in the quest for new bioactive compounds, because their isolation and purification are tedious. The fact that they are very sensitive to mechanical stress and are prone to lysis make their handling very difficult (Reichenbach and Dworkin 1992). The ability of myxobacteria to synthesize a particular compound is specific to a strain, which makes myxobacteria an attractive proposition for bioprospection (Reichenbach and Hofle 1999). In view of their immense potential as a source of natural compounds having new chemical structures with unique mechanism of action and no report on functional diversity of myxobacteria from Indian habitats, the present study was undertaken to investigate myxobacteria to assess them as source of novel pharmaceutically active molecules.
Our study shows that myxobacteria isolated from soil in India inhibit Bacillus cereus, Mycobacterium smegmatis and Gram-negative bacteria (Pseudomonas syringae and Enterobacter cloacae) suggesting their potential to be exploited as a source of antimicrobial compounds. The extracted molecules also showed antiproliferative activity against the cancer cell lines of cervix, liver, pancreas, prostate and bone. Most of the compounds tested in laboratories fail to make it to the clinic because of bioavailability limitations. To address this constraint, in this study, we focused on the assessment of bioactive potential of water-soluble compounds secreted by myxobacteria.
Materials and methods
All chemicals and media components were procured from HiMedia, Sisco Research Laboratory, Lobachemie, Genei-Merck or Sigma-Aldrich. All plastic wares and glass wares were purchased from HiMedia, Tarsons or Borosil.
Isolation, purification and taxonomy of myxobacteria
Myxobacteria were isolated by baiting technique from the soil samples collected near decaying plants and animals in the plains of north India (Reichenbach and Hofle 1999). Escherichia coli and/or autoclaved baker's yeast was streaked on cycloheximide (100 µg/ml) incorporated water CleriGel plate as three parallel streaks, cross streak or dot. Cycloheximide (100 µg/ml)-treated soil samples were placed on E. coli/autoclaved yeast and incubated at 30 °C for 3–20 days. For purification, colony morphology and/or fruiting bodies were viewed under stereomicroscope to confirm the identity of myxobacteria according to the criteria indicated by Reichenbach and Dworkin (1992). The fruiting bodies and cells from the edge of the swarms were picked up and streaked on CY agar (g/l: Casitone, 10; yeast extract, 1; CaCl2·2H2O, 1; agar 20) or SP agar (DSZM medium 222) and incubated at 30 °C. This step was repeated till the isolates were obtained in pure culture. Thereafter, the isolates were maintained on SP medium.
Molecular characterization of myxobacteria
Genomic DNA isolation
Myxobacteria were grown in SP medium at 30 °C for 48 h on a rotary shaker. Genomic DNA was extracted using HiYield™ Genomic DNA Mini Kit (Real Biotech Corporation, Taiwan).
PCR amplification of 16S rDNA
Genomic DNA of myxobacterial isolates was used as template for PCR amplification of 16S rRNA gene with bacterial universal forward primer 8-27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and reverse primer 1492R (5′-GGTTACCTTGTTACGACTT-3′). The PCR reaction mixture (50 µl) contained 5 µl of 10× Taq reaction buffer (Genei, Bangalore, India), 0.2 µM each primer, 200 µM each dNTP, 20 ng of template DNA and 1.5 U Taq DNA polymerase (Genei, Bangalore, India). PCR amplification was carried out using MyCycler (Biorad). Thermal cycling conditions were as follows: (1) initial denaturation (2 min at 95 °C), (2) five cycles of denaturation (95 °C for 45 s), annealing (45 °C for 45 s) and primer extension (72 °C for 45 s), (3) five cycles of denaturation (95 °C for 45 s), annealing (50 °C for 45 s) and primer extension (72 °C for 45 s), (4) 30 cycles of denaturation (95 °C for 45 s), annealing (55 °C for 45 s) and primer extension (72 °C for 45 s), (5) final extension step at 72 °C for 10 min. The amplified product was resolved by agarose electrophoresis, and the fragment was excised out from the gel and purified by HiYield™ Gel/PCR DNA Kit (Real Biotech Corporation, Taiwan).
16S rDNA sequence analysis
16S rDNA was sequenced at Bioserve Biotechnology Pvt. Ltd., Hyderabad, India, and the identification of the isolates was confirmed on the basis of 16S rRNA gene sequence using the Ribosomal Database Project (RDP; http://rdp.cme.msu.edu) (Cole et al. 2014). Multiple sequence alignment and phylogenetic analysis were performed using neighbor-joining (N-J) method with 1000 bootstrapping replications of the MEGA 6.06 software package (Tamura et al. 2013).
Preparation of extract
A seed culture was prepared by growing myxobacteria isolates in 10 ml CY broth at 30 °C for 36–48 h in an incubator shaker. To prepare extract for evaluating the antibacterial properties, 5 ml of seed culture was inoculated into 500 ml CY broth and incubated at 30 °C at 180 rpm for 72 h. Thereafter, the culture supernatant was incubated with 2% (w/v) Amberlite XAD16 N (Sigma) at 30 °C for 16 h. The resin was extracted with 200 ml of methanol at 30 °C for 12 h at 180 rpm. The extract was dried on a vacuum rotary evaporator (Buchi) at 40 °C. The dried powdered extract was suspended in water at 0.25 mg/ml, incubated at 30 °C overnight and centrifuged at 10,000 rpm; the supernatant was designated as water extract (WE). The insoluble residual matter was dissolved in 250 µl DMSO and designated as DMSO extract (DE).
To prepare extracts for checking the anticancer activity, myxobacteria were grown in 300 ml of CY broth at 30 °C for 36 h in an incubator shaker. Thereafter, 300 ml of ethyl acetate was added to each flask and extraction was carried out at 30 °C for 2 h at 180 rpm. The upper organic phase thus obtained was separated and concentrated on a rotary evaporator and suspended in water as above.
Water-soluble fractions of the extracts were sterilized through a 0.22 µ filter and stored at 4 °C till further use.
Antimicrobial activity
The antibacterial activity was evaluated by agar plate diffusion assay. The extracts from the isolated myxobacteria were screened for antibacterial activity against Bacillus cereus MTCC 430, Enterobacter cloacae MTCC 509, Pseudomonas syringae MTCC 1604 and Mycobacterium smegmatis MTCC6. 100 µl of each WE or 50 µl of each DE was added to the wells (5 mm) on Mueller–Hinton (MH) agar plate seeded with 100 µl of activated test cultures. The plates were incubated at 30 °C for 24 h and the zones of inhibition around the bacterial extracts were measured and compared with the negative control (water for WE and 10% DMSO for DE). Kanamycin (100 µg/ml) was included as positive control in the assay.
In vitro cytotoxicity against human cell lines
Human cancer cell lines (HepG2, HeLa, MIA PaCa-2, MG-63 and PC-3) were procured from NCCS, Pune, India. Cells were grown in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) with 10,000 U penicillin, 10 mg streptomycin and 25 μg amphotericin B per ml at 37 °C in a CO2 incubator (5% CO2; 90% RH).
In vitro cytotoxicity of different isolates was determined by sulforhodamine B (SRB) assay. 3 × 103 cells were added to each well of 96 well plates and allowed to attach and grow for 24 h. 10 µl of WE was added to each well and the plate was incubated for 48 h. Cells were also incubated with 10 μM doxorubicin (DOX) as positive control. The cells were then fixed with 50 μl of ice-cold 50% (w/v) trichloroacetic acid at 4 °C for 1 h (Skehan et al. 1990). To the washed and air dried plate, 100 μl SRB (0.057% w/v SRB in 1% acetic acid solution) solution was added to each well and the plate was incubated at room temperature for 30 min. The unbound SRB solution was removed by washing with 1% acetic acid solution and plates were air dried. 200 μl of 10 mM Tris base solution (pH 10.5) was added to each well and after shaking on a gyratory shaker for 5 min, the optical density was measured at 510 nm using a microplate reader (Vichai and Kirtikara 2006). Inhibition of cell growth was calculated as: % cell growth = (OD of treated cells/OD of control) × 100; % cell inhibition = 100 − % of cell growth. Cytotoxicity was also checked on Chinese hamster ovary (CHO) cells to assess the specificity of extracts on cancer cells.
Mass spectrometry of the extracts
Extracts were fractionated by HPLC performed on an Agilent 6410 LC/MS–MS instrument (Agilent Technologies, USA) equipped with a column Agilent ZORBAX Eclipse XAD C18, (4.6 × 100 mm) eluted at 0.5 ml min flow rate. The mobile phase consisted of solvent A (water with 0.01% formic acid) and solvent B (acetontrile). A gradient program was used as follow: 0–8 min, 5–70% B; 8–12 min, 70% B; 12–13 min, 70–5% B; and 13–15 min, 5% B. The effluent from the column was diverted to the ESI interface of a 6410B triple quad LC/MS system (Agilent Technologies, USA.
The analysis was performed using an electrospray ionization (ESI) source in positive and negative modes. The operation conditions were as follows: scan range of 50–1800 amu, ion source temperature 300 °C, nebulizer 50 psi, gas flow 11 L/min, capillary voltage 3000 and a step size of 0.1 amu. Agilent Mass Hunter software (version B.04.00) was used for data acquisition and processing. Data were manually sorted to list such information as the retention time, m/z values for adduct ions.
Statistical analysis
All the experiments were carried out in triplicate and the values were represented as their mean ± standard deviation.
Results
Taxonomy of the isolated myxobacteria
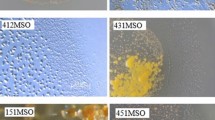
Fifty myxobacterial strains were isolated from pure culture from various soil samples in India. These isolates were identified by observing the morphology of vegetative cells, swarms and fruiting bodies. The isolates that had identical colony/swarm morphology or fruiting bodies from the same soil sample were discarded to avoid replication of the isolated strains. Among these myxobacteria, nine isolates showing potent inhibitory activities against the test microorganisms and human cancer cell lines were selected for molecular characterization. The amplified 16S rDNA of the selected isolates was sequenced and compared with the 16S rDNA sequence of myxobacteria represented in the RDP database (Table 1). Using MEGA6, a phylogenetic tree was generated by comparing the amplified 16S rDNA sequences with the 16S rDNA sequences of the myxobacterial type strains (Fig 1). All the isolated myxobacteria belong to two families (Cystobacteraceae and Myxococcaceae) in the suborder Cystobacterineae within the order myxococcales. The isolated myxobacteria belonged to five genera: Corallococcus, Pyxidicoccus, Cystobacter, Archangium and Myxococcus. The swarm morphology of the purified myxobacterial strains is shown in Online Resource 1.
Antimicrobial activity of myxobacteria
The results in Table 2 show that seven of the isolated myxobacterial strains examined in this study exhibited antimicrobial activity against one or more of the test microorganisms. The extract from S104 and S145 showed broad-spectrum antibacterial activity and inhibited all the test organisms. WE and DE of GNDU 172 showed appreciable activity against B. subtilis and E. cloacae, whereas S225 did not exhibit inhibitory activity against any of the test organisms. The DE of both S213 and S223 exhibited a similar spectrum of antibacterial activity, being active against B. cereus, P. syringae and M. smegmatis, whereas S223 WE in addition strongly inhibited B. cereus. WE and DE of the isolate S229 showed potent inhibitory activity against B. cereus and P. syringae, respectively. The DE of the isolate S233 was specifically active against M. smegmatis (Fig. 2).
Anticancer activity of myxobacteria
Out of the selected isolates, eight exhibited antiproliferative activity against one or more cancer cell lines, viz., cervical (HeLa), HepG2 (liver), MG-63 (osteosarcoma), MIA PaCa-2 (Pancreatic) and PC-3 (prostate) tested in this study (Table 3). The extract from S145 was active (more than 50% growth inhibition) against all tested cell lines except HepG2 (liver) cell line. S145 showed 58, 53, 60 and 86% inhibition of HeLa, MG-63, MIA PaCa-2 and PC-3 cell lines, respectively. GNDU172 and S225 produced antiproliferative agents against MIA PaCa-2 (88 and 72% inhibition, respectively) and PC-3 (59 and 67% inhibition, respectively) cell lines. The extract of S199 showed inhibition of 76% against PC-3 and 54% against HeLa cell line. S213 and S223 produced potent anticancer agents against MIA PaCa-2 (more than 80% inhibition) and PC-3 (64 and 85% inhibition, respectively). S229 showed 57% inhibition of MG-63 in addition to being active against MIA PaCa-2 and PC-3 exhibiting 73 and 86% inhibition, respectively. The isolate S233 was active against most of the tested cell lines where it exhibited 61, 53, 84 and 76% activity against HeLa, HepG2, MIA PaCa-2 and PC-3 cell lines respectively.
Discussion
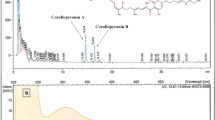
There is a persistent requirement for new bioactive compounds so that the demand for new natural products in therapy can be met. Myxobacteria have great potential to contribute to the pool of natural products for drug discovery. Myxobacteria were isolated from soil collected near decaying plants and animals from plains in north India by E. coli baiting method, because it has been reported to be a productive isolation method for soil myxobacteria (Gaspari et al. 2005). The isolates were identified by swarm morphology or fruiting bodies and subjected to molecular characterization by 16S rDNA sequencing. A phylogenetic tree was constructed based on the sequences of the 16S rDNA of the isolated myxobacteria, strains showing the highest similarity with the isolated myxobacteria and the type strains from the database which showed that isolates were distributed within the suborder Cystobacterineae. Though the isolates S104 and S145 showed the greatest similarity in the RDP database to C. coralloides, S145 resembles C. exiguus based on the morphology of its fruiting bodies. Stackebrandt and Pauker (2005) have reported that Corallococcus spp., C. coralloides and C. exiguus, share high 16rRNA gene sequence similarity and can be distinguished based on their fruiting bodies. The isolates included in this study that belong to the same genera showed differences in the functional characteristics with respect to their ability to produce bioactive metabolites. The isolates obtained in pure cultures exhibited inhibitory activities against M. smegmatis, B. cereus and Gram-negative bacteria. Five out of the nine myxobacterial isolates selected for functional characterization produced inhibitory agents against Gram-negative test bacteria (E. cloacae and P. syringae), whereas six isolates showed activity against M. smegmatis. An assessment of the available antibiotic compounds till date has revealed that 30% of these inhibit only Gram-positive bacteria and only 1.5% of the compounds exhibit activity specifically against Gram-negative bacteria (Berdy 2005). Studies on myxobacteria have revealed that they secrete natural products that are inhibitory predominantly to Gram-positive bacteria, whereas very few myxobacterial isolates show inhibitory activity against Gram-negative bacteria (Schaberle et al. 2014). On the contrary, more than 50% of myxobacteria isolated from Israel by Gaspari et al. (2005) synthesized compounds showing inhibitory activity against Gram-negative bacteria, suggesting that myxobacteria have the capability to meet the medical need for compounds acting on Gram-negative pathogens. The antibacterial spectrum of myxobacterial isolated in this study is different from the inhibitory activities reported earlier for the same genera. Corallopyronins, reported from C. coralliodes, do not inhibit Gram-negative bacteria, whereas the extract reported in this study from S104 showed potent activity against E. cloacae and weak activity against P. syringae (Schaberle et al. 2014). The comparison of the m/z of the peaks in mass spectra of the water-soluble fraction of methanol extracts from isolates S104 and S145 with the bioactive compounds synthesized by C. coralloides listed in the Dictionary of Natural Products (http://dnp.chemnetbase.com) did not identify any known compounds, suggesting that myxobacteria isolated in this study produce novel bioactive metabolites (Online Resource 2). This is the first report of the inhibitory activity of Pyxidicoccus sp. (GNDU172) against E. cloacae. Two molecules, gulmirecins and disciformycins, reported from P. fallax are active only against staphylococci (Schieferdecker et al. 2014). Our isolates S213 and S223; identified as Cystobacter sp., showed weak activity in DE against P. syringae. On the contrary, Baumann et al. (2014) reported cystobactamids from Cystobacter sp. that efficiently inhibited the growth of Gram-negative bacteria E. coli and Acinetobacter baumannii. Another light- and oxygen-sensitive secondary metabolite with no aqueous solubility, Roimatacene, synthesized by Cystobacter ferrugineus Cb G35 inhibits Gram-negative bacteria (Schaberle et al. 2014). We have used the extracts stored at 4 °C over a time period of more than 3 months and did not encounter any decrease in the activity of the compounds (data not shown). It is possible that the isolates in this study are novel strains that produce previously unknown metabolites having new chemical structures that can be developed into potent lead structures against Gram-negative pathogens. A study on the secondary metabolome of M. xanthus has suggested that intraspecific screens of species of myxobacteria are likely to yield novel secondary metabolites (Krug et al. 2008). These compounds can be attributed to the orphan biosynthetic pathways discovered in the genomes of myxobacteria (Gross 2007).
The isolated myxobacteria produced anticancer compounds that showed high frequency of inhibitory activity against the human cancer cell lines of the pancreas and prostate. The extracts of some isolates inhibited more than one cancer cell line tested in this study. It is possible that the compounds produced by these bacteria target a common underlying mechanism of inhibition of growth of cancer cells. This study reports for the first time the anticancer activity of secondary metabolites from C. exiguus. Pyxidicoccus sp. in this study produces compounds that are inhibitory to pancreatic (MIA PaCa-2) and prostate (PC-3) cancer cell lines, whereas a secondary metabolite from P. fallax has been associated with inhibition of leukemic cells by Schieferdecker et al. (2015). In the present investigation, the extract from M. fulvus (S199) exhibited the inhibition of PC-3 (prostate cancer) and HeLa (cervical cancer) cells which have not been documented in the cytotoxicity profile of the metabolite isolated from M. fulvus in a study by Ahn et al. (1999). The anticancer peptides, tubulysins, which are very effective against drug-resistant cancers, have been isolated from Cystobacter sp. and Archangium sp. but these peptides have no aqueous solubility and require the development of targeted delivery efforts (Murray et al. 2015). In this study, the extracts from Cystobacter spp. (S213, S223 and S229) and Archangium sp. (S233) that exhibit anticancer activity constitute water-soluble compounds and are unlikely to be previously described tubulysins. Horstmann et al. (2011) reported inhibition of cervix and prostate cancer cell lines by secondary metabolites of Cystobacter sp. and Archangium sp. and in this study MG-63 (bone), and MIA PaCa-2 (pancreatic) cancer cell lines were also inhibited by extracts from these myxobacteria.
It is evident from the results of this study that the isolated myxobacteria isolated from Indian habitats can contribute novel compounds to the pool of natural products for therapy against pathogenic microbes and human cancers. Further studies are being focused on the production and isolation of bioactive molecules, to determine their chemical nature and to deduce the mechanism of action for antibacterial and anticancer activities.
References
Ahn JW, Woo SH, Lee CO, Cho KY, Kim BS (1999) KR025, a new cytotoxic compound from Myxococcus fulvus. J Nat Prod 62:495–496
Baumann S, Herrmann J, Raju R, Steinmetz H, Mohr KI, Huttel S, Harmrolfs K, Stadler M, Muller R (2014) Cystobactamids: myxobacterial topoisomerase inhibitors exhibiting potent antibacterial activity. Angew Chem Int Ed Engl 53:14605–14609. doi:10.1002/anie.201409964
Berdy J (2005) Bioactive microbial metabolites. J Antibiot (Tokyo) 58:1–26. doi:10.1038/ja.2005.1
Bode HB, Muller R (2006) Analysis of myxobacterial secondary metabolism goes molecular. J Ind Microbiol Biotechnol 33:577–588. doi:10.1007/s10295-006-0082-7
Burris HA 3rd (2008) Preclinical investigations with epothilones in breast cancer models. Semin Oncol 35:S15–S21. doi:10.1053/j.seminoncol.2008.02.002
Buss AD, Waigh RD (1995) Natural products as leads for new pharmaceuticals. In: Wolff ME (ed) Burger’s medicinal chemistry and drug discovery. Principles and practice. Wiley, New York, pp 983–1033. doi:10.1002/0471266949.bmc018
Chai Y, Pistorius D, Ullrich A, Weissman KJ, Kazmaier U, Muller R (2010) Discovery of 23 natural tubulysins from Angiococcus disciformis An d48 and Cystobacter SBCb004. Chem Biol 17:296–309. doi:10.1016/j.chembiol.2010.01.016
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, Brown CT, Porras-Alfaro A, Kuske CR, Tiedje JM (2014) Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42(Database issue):D633–D642. doi:10.1093/nar/gkt1244
Dawid W (2000) Biology and global distribution of myxobacteria in soils. FEMS Microbiol Rev 24:403–427. doi:10.1111/j.1574-6976.2000.tb00548.x
Garcia RO, Krug D, Muller R (2009) Discovering natural products from myxobacteria with emphasis on rare producer strains in combination with improved analytical methods. Methods Enzymol 458:59–91. doi:10.1016/S0076-6879(09)04803-4
Gaspari F, Paitan Y, Mainini M, Losi D (2005) Myxobacteria isolated in Israel as potential source of new anti-infectives. J Appl Microbiol 98:429–439. doi:10.1111/j.1365-2672.2004.02477.x
Gross H (2007) Strategies to unravel the function of orphan biosynthesis pathways: recent examples and future prospects. Appl Microbiol Biotechnol 75:267–277. doi:10.1007/s00253-007-0900-5
Han K, Li ZF, Peng R, Zhu LP, Zhou T, Wang LG, Li SG, Zhang XB, Hu W, Wu ZH, Qin N, Li YX (2013) Extraordinary expansion of a Sorangium cellulosum genome from an alkaline milieu. Sci Rep 3:2101. doi:10.1038/srep02101
Herrmann J, Fayad AA, Muller R (2017) Natural products from myxobacteria: novel metabolites and bioactivities. Nat Prod Rep 34:135–160. doi:10.1039/c6np00106h
Horstmann N, Essig S, Bockelmann S, Wieczorek H, Huss M, Sasse F, Menche D (2011) Archazolid A-15-O-β-d-glucopyranoside and iso-archazolid B: potent V-ATPase inhibitory polyketides from the myxobacteria Cystobacter violaceus and Archangium gephyra. J Nat Prod 74:1100–1105. doi:10.1021/np200036v
Iizuka T, Fudou R, Jojima Y, Ogawa S, Yamanaka S, Inukai Y, Ojika M (2006) Miuraenamides A and B, novel antimicrobial cyclic depsipeptides from a new slightly halophilic myxobacterium: taxonomy, production, and biological properties. J Antibiot (Tokyo) 59:385–391. doi:10.1038/ja.2006.55
Irschik H, Reichenbach H, Hofle G, Jansen R (2007) The thuggacins, novel antibacterial macrolides from Sorangium cellulosum acting against selected Gram-positive bacteria. J Antibiot (Tokyo) 60:733–738. doi:10.1038/ja.2007.95
Kim YJ, Kim HJ, Kim GW, Cho K, Takahashi S, Koshino H, Kim WG (2016) Isolation of coralmycins A and B, potent anti-Gram negative compounds from the Myxobacteria Corallococcus coralloides M23. J Nat Prod 79:2223–2228. doi:10.1021/acs.jnatprod.6b00294
Krastel P, Roggo S, Schirle M, Ross NT (2015) Nannocystin A: an elongation factor 1 inhibitor from myxobacteria with differential anti-cancer properties. Angew Chem Int Ed Engl 54:10149–10154. doi:10.1002/anie.201505069
Krug D, Zurek G, Revermann O, Vos M, Velicer GJ, Muller R (2008) Discovering the hidden secondary metabolome of Myxococcus xanthus: a study of intraspecific diversity. Appl Environ Microbiol 74:3058–3068. doi:10.1128/AEM.02863-07
Kunze B, Jansen R, Hofle G, Reichenbach H (1994) Crocacin, a new electron transport inhibitor from Chondromyces crocatus (myxobacteria). Production, isolation, physico-chemical and biological properties. J Antibiot (Tokyo) 47:881–886. doi:10.7164/antibiotics.47.881
Kunze B, Reichenbach H, Muller R, Hofle G (2005) Aurafuron A and B, new bioactive polyketides from Stigmatella aurantiaca and Archangium gephyra (Myxobacteria). Fermentation, isolation, physico-chemical properties, structure and biological activity. J Antibiot (Tokyo) 58:244–251. doi:10.1038/ja.2005.28
Kunze B, Sasse F, Wieczorek H, Huss M (2007) Cruentaren A, a highly cytotoxic benzolactone from Myxobacteria is a novel selective inhibitor of mitochondrial F1-ATPases. FEBS Lett 581:3523–3527. doi:10.1016/j.febslet.2007.06.069
Kunze B, Bohlendorf B, Reichenbach H, Hofle G (2008) Pedein A and B: production, isolation, structure elucidation and biological properties of new antifungal cyclopeptides from Chondromyces pediculatus (Myxobacteria). J Antibiot (Tokyo) 61:18–26. doi:10.1038/ja.2008.104
Murray BC, Peterson MT, Fecik RA (2015) Chemistry and biology of tubulysins: antimitotic tetrapeptides with activity against drug resistant cancers. Nat Prod Rep 32:654–662. doi:10.1039/c4np00036f
Nickeleit I, Zender S, Sasse F, Geffers R, Brandes G, Sorensen I, Steinmetz H, Kubicka S, Carlomagno T, Menche D, Gutgemann I, Buer J, Gossler A, Manns MP, Kalesse M, Frank R, Malek NP (2008) Argyrin a reveals a critical role for the tumor suppressor protein p27(kip1) in mediating antitumor activities in response to proteasome inhibition. Cancer Cell 14:23–35. doi:10.1016/j.ccr.2008.05.016
Ravenschlag K, Sahm K, Pernthaler J, Amann R (1999) High bacterial diversity in permanently cold marine sediments. Appl Environ Microbiol 65:3982–3989 PMID: 10473405
Reichenbach H (1999) The ecology of the myxobacteria. Environ Microbiol 1:15–21. doi:10.1046/j.1462-2920.1999.00016.x
Reichenbach H (2001) Myxobacteria, producers of novel bioactive substances. J Ind Microbiol Biotechnol 27:149–156. doi:10.1038/sj.jim.7000025
Reichenbach H, Dworkin M (1992) The myxobacteria. In: Balows A, Truper HG, Dworkin M, Harder W, Schleifer KH (eds) The prokaryotes, 2nd edn. Springer Verlag, New York, pp 3416–3487. doi:10.1007/978-1-4757-2191-1
Reichenbach H, Hofle G (1999) Myxobacteria as producers of secondary metabolites. In: S Grabley, R Thiericke (eds) Drug discovery from nature. Springer, Berlin, Germany, pp 149–179. doi:10.1046/j.1462-2920.1999.00016.x
Sasse F, Steinmetz H, Heil J, Hofle G, Reichenbach H (2000) Tubulysins, new cytostatic peptides from myxobacteria acting on microtubuli. Production, isolation, physico-chemical and biological properties. J Antibiot (Tokyo) 53:879–885. doi:10.7164/antibiotics.53.879
Sasse F, Leibold T, Kunze B, Hofle G, Reichenbach H (2003) Cyrmenins, new beta-methoxyacrylate inhibitors of the electron transport. Production, isolation, physico-chemical and biological properties. J Antibiot (Tokyo) 56:827–831. doi:10.7164/antibiotics.56.827
Schaberle TF, Lohr F, Schmitz A, Konig GM (2014) Antibiotics from myxobacteria. Nat Prod Rep 31:953–972. doi:10.1039/c4np00011k
Schieferdecker S, Konig S, Weigel C, Dahse HM, Werz O, Nett M (2014) Structure and biosynthetic assembly of gulmirecins, macrolide antibiotics from the predatory bacterium Pyxidicoccus fallax. Chemistry 20:15933–15940. doi:10.1002/chem.201404291
Schieferdecker S, Konig S, Koeberle A, Dahse HM, Werz O, Nett M (2015) Myxochelins target human 5-lipoxygenase. J Nat Prod 78:335–338. doi:10.1021/np500909b
Skehan P, Storeng R, Scudiero D, Monks A, McMahon J, Vistica D, Warren JT, Bokesch H, Kenney S, Boyd MR (1990) New colorimetric cytotoxicity assay for anticancer-drug screening. J Natl Cancer Inst 82:1107–1112. doi:10.1093/jnci/82.13.1107
Stackebrandt E, Pauker O (2005) Gene sequence heterogeneity of Corallococcus coralloides strains isolated from geographically diverse locations. Environ Microbiol 7:1017–1023. doi:10.1111/j.1462-2920.2005.00773.x
Steinmetz H, Glaser N, Herdtweck E, Sasse F et al (2004) Isolation, crystal and solution structure determination, and biosynthesis of tubulysins-powerful inhibitors of tubulin polymerization from myxobacteria. Angew Chem Int Ed Engl 43:4888–4892. doi:10.1002/anie.200460147
Steinmetz H, Mohr KI, Zander W, Jansen R, Gerth K, Muller R (2012) Indiacens A and B: prenyl indoles from the myxobacterium Sandaracinus amylolyticus. J Nat Prod 75:1803–1805. doi:10.1021/np300288b
Surup F, Viehrig K, Mohr KI, Herrmann J, Jansen R, Muller R (2014) Disciformycins A and B: 12-membered macrolide glycoside antibiotics from the myxobacterium Pyxidicoccus fallax active against multiresistant staphylococci. Angew Chem Int Ed Engl 53:13588–13591. doi:10.1002/anie.201406973
Tamura K, Stecher G, Peterson D, Filipski A (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729. doi:10.1093/molbev/mst197
Tan AR, Toppmeyer DL (2008) Ixabepilone in metastatic breast cancer: complement or alternative to taxanes. Clin Cancer Res 14:6725–6729. doi:10.1158/1078-0432.CCR-07-4704
Vichai V, Kirtikara K (2006) Sulforhodamine B colorimetric assay for cytotoxicity screening. Nat Protoc 1:1112–1116. doi:10.1371/journal.pone.0091694
Wenzel SC, Muller R (2009) Myxobacteria—‘microbial factories’ for the production of bioactive secondary metabolites. Mol BioSyst 5:567–574. doi:10.1039/b901287g
Acknowledgements
The authors are grateful to the Council for Scientific and Industrial Research (CSIR), New Delhi, India, for providing financial assistance. SK is grateful to CSIR and UGC for the fellowship. AKY is grateful to DST for the fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no commercial/financial conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kumar, S., Yadav, A.K., Chambel, P. et al. Molecular and functional characterization of myxobacteria isolated from soil in India. 3 Biotech 7, 112 (2017). https://doi.org/10.1007/s13205-017-0722-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-0722-9