Abstract
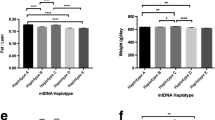
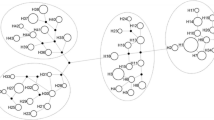
The mitochondrial genome plays an important role in energy production and cells functions control. Therefore, it can affect the traits which are significant for selection of farm animals. The published data allow assuming that the mtDNA variability in pigs is one of the ways to assess and predict the breeding and productive qualities. The aim of our work was to determine the mtDNA haplotypes by the D-loop region sequencing and evaluate their associations with the selection features in pigs. Studies were carried out on landrace sows (n = 2626) bred in the Russian Federation. We estimated the Muscle Depth and the Percent Meat. As a result, three mtDNA haplogroups (C, D, and E) were identified. In the studied group of pigs, haplotype E was determined in 1378 sows (52.5%) and had the highest frequency. Haplotype D was detected in 1064 sows (40.6%). In general, haplotypes of European origin were more common in our population. Among the haplotypes of Asian origin (including A, B, and C), only haplotype C was identified in our group of pigs. It was determined in 182 sows (6.9%). Haplotypes A and B were not identified. Analysis of variance (ANOVA) has demonstrated the significance of differences in the haplotypes of the mtDNA regarding the studied traits of meat productivity in sows. These facts confirm the prospects of mtDNA studies as determinants of pig productivity. Our results have shown that some haplotypes of mtDNA are largely related to phenotypic traits.


Similar content being viewed by others
References
Bruggerhoff K, Zakhartchenko V, Wenigerkind H, Reichenbach HD, Prelle K, Schernthaner W, Alberio R, Kuchenhoff H, Stojkovic M, Brem G, Hiendleder S, Wolf E (2002) Bovine somatic cell nuclear transfer using recipient oocytes recovered by ovum pick-up: effect of maternal lineage of oocyte donors. Biol Reprod 66(2):367–373. https://doi.org/10.1095/biolreprod66.2.367
Cagnone G, Tsai TS, Srirattana K, Rossello F, Powell DR, Rohrer G, Cree L, Trounce IA, St John JC (2016) Segregation of naturally occurring mitochondrial DNA variants in a mini-pig model. Genetics 202(3):931–944. https://doi.org/10.1534/genetics.115.181321
Cianciulli A, Calvello R, Mitolo V, Panaro M (2018) Intron evolution of chicken, mouse, and human mitochondrial carrier genes. Rend Fis Acc Lincei 29:437. https://doi.org/10.1007/s12210-018-0713-8
Ciesielski GL, Oliveira MT, Kaguni LS (2016) Animal mitochondrial DNA replication. Enzymes 39:255–292. https://doi.org/10.1016/bs.enz.2016.03.006
Clop A, Amills M, Noguera JL, Fernández A, Capote J, Ramón MM, Kelly L, Kijas JM, Andersson L, Sànchez A (2004) Estimating the frequency of Asian cytochrome B haplotypes in standard European and local Spanish pig breeds. Genet Sel Evol 36:97–104. https://doi.org/10.1186/1297-9686-36-1-97
Derr JN, Hedrick PW, Halbert ND, Plough L, Dobson LK, King J, Duncan C, Hunter DL, Cohen ND, Hedgecock D (2012) Phenotypic effects of cattle mitochondrial DNA in American bison. Conserv Biol 26(6):1130–1136. https://doi.org/10.1111/j.1523-1739.2012.01905.x
Fernandez AI, Alves E, Fernandez A, de Pedro E, Lopez-Garcia MA, Ovilo C, Rodriguez MC, Silio L (2008) Mitochondrial genome polymorphisms associated with longissimus muscle composition in Iberian pigs. J Anim Sci 86(6):1283–1290. https://doi.org/10.2527/jas.2007-0568
Ferreira IM, Braga TF, Guimarães EC, Durval MC, Mendes LB, Soares JS, Matarim DL, Kruger BC, Franco MM, Antunes RC (2019) A DdeI polymorphism in the growth hormone gene is associated with higher lean meat yield and pH 45 min in postmortem pigs. Genet Mol Res. https://doi.org/10.4238/gmr18257
Giuffra E, Kijas JM, Amarger V, Carlborg O, Jeon JT et al (2000) The origin of the domestic pig: independent domestication and subsequent introgression. Genetics 154:1785–1791
Gorlov IF, Kolosov YA, Shirokova NV, Getmantseva LV, Slozhenkina MI, Mosolova NI, Bakoev NF, Leonova MA, Kolosov AY, Zlobina EY (2018) GDF9 gene polymorphism and its association with litter size in two Russian sheep breeds. Rend Fis Acc Lincei 29:61. https://doi.org/10.1007/s12210-017-0659-2
Guo Y, Hou L, Zhang X, Huang M, Mao H, Chen H, Ma J, Chen C, Ai H, Ren J, Huang L (2015) A meta analysis of genome-wide association studies for limb bone lengths in four pig populations. BMC Genet 16:95. https://doi.org/10.1186/s12863-015-0257-1
Häggman J, Uimari P (2017) Novel harmful recessive haplotypes for reproductive traits in pigs. J Anim Breed Genet 134(2):129–135. https://doi.org/10.1111/jbg.12240
Kadenbach B (2018) Regulation of mitochondrial respiration and ATP synthesis via cytochrome C oxidase. Rend Fis Acc Lincei 29:421. https://doi.org/10.1007/s12210-018-0710-y
Kelly RD, Rodda AE, Dickinson A, Mahmud A, Nefzger CM, Lee W, Forsythe JS, Polo JM, Trounce IA, McKenzie M, Nisbet DR, St John JC (2013) Mitochondrial DNA haplotypes define gene expression patterns in pluripotent and differentiating embryonic stem cells. Stem Cells 31(4):703–716. https://doi.org/10.1002/stem.1313
Kijas JMH, Andersson L (2001) A phylogenetic study of the origin of the domestic pig estimated from the near-complete mtDNA genome. J Mol Evol 52:302–308. https://doi.org/10.1007/s002390010158
Kim KI, Lee JH, Li K et al (2002) Phylogenetic relationships of Asian and European pig breeds determined by mitochondrial DNA D-loop sequence polymorphism. Anim Genet 33:19–25. https://doi.org/10.1046/j.1365-2052.2002.00784.x
Kim JM, Choi BD, Kim BC, Park SS, Hong KC (2009) Associations of the variation in the porcine myogenin gene with muscle fibre characteristics, lean meat production and meat quality traits. J Anim Breed Genet 126(2):134–141. https://doi.org/10.1111/j.1439-0388.2008.00724.x
Kumar JK, Singh AP, Srivastav AB, Sarkhel BC (2018) Molecular characterization of the complete mitochondrial genome sequence of Indian wild pig (Sus scrofa cristatus). J Anim Biotechnol 30:186–191. https://doi.org/10.1080/10495398.2018.1469506
Larson G, Dobney K, Albarella U et al (2005) Worldwide phylogeography of wild boar reveals multiple centers of pig domestication. Science 307(5715):1618–1621. https://doi.org/10.1126/science.1106927
Lee WT, Sun X, Tsai TS, Johnson JL, Gould JA, Garama DJ, Gough DJ, McKenzie M, Trounce IA, St John JC (2017) Mitochondrial DNA haplotypes induce differential patterns of DNA methylation that result in differential chromosomal gene expression patterns. Cell Death Discov 3:17062. https://doi.org/10.1038/cddiscovery.2017.62
Mannen H, Morimoto ML, Oyamat K, Mukai F, Tsuji S (2003) Identification of mitochondrial DNA substitutions related to meat quality in Japanese black cattle. J Anim Sci 81(1):68–73. https://doi.org/10.2527/2003.81168x
Meadus WJ, Duff P, Juarez M, Roberts JC, Zantinge JL (2018) Identification of marbling gene loci in commercial pigs in canadian herds. Agriculture 8:122. https://doi.org/10.20944/preprints201806.0330.v1
Mignon-Grasteaua S, Boissy A, Bouix J, Faurea J-M, Fisherd AD, Hinch GN, Jensen P, Le Neindre P, Mormède P, Prunet P, Vandeputte M, Beaumonta C (2005) Genetics of adaptation and domestication in livestock. Livest Prod Sci. https://doi.org/10.1016/j.livprodsci.2004.11.001
Nakavisut S, Crump RE (2006) Graser Body length and its genetic relationships with production and reproduction traits in pigs. In: Proceedings of the 8th World Congress on Genetics Applied to Livestock Production, Belo Horizonte, MG, Brasil pp 06–45
Nisztuk-Pacek S, Slaska B, Zieba G, Rozempolska-Rucinska I (2018) Two mitochondrial genes are associated with performance traits in farmed raccoon dogs (Nyctereutes procyonoides) Czech. J Anim Sci 63(3):110–118
Okumura N, Kurosawa Y, Kobayashi E (2001) Genetic relationship amongst the major non-coding regions of mitochondrial DNAs in wild boars and several breeds of domesticated pigs. Anim Genet 32:139–147. https://doi.org/10.1046/j.1365-2052.2001.00757.x
Sato M, Sato K (2013) Maternal inheritance of mitochondrial DNA by diverse mechanisms to eliminate paternal mitochondrial DNA. Biochim Biophys Acta 1833:1979–1984. https://doi.org/10.1016/j.bbamcr.2013.03.010
Sell-Kubiak E, Duijvesteijn N, Lopes MS, Janss LL, Knol EF, Bijma P, Mulder HA (2015) Genome-wide association study reveals novel loci for litter size and its variability in a large white pig population. BMC Genom 16:1049. https://doi.org/10.1186/s12864-015-2273-y
St John JC, Tsai T (2018) The association of mitochondrial DNA haplotypes and phenotypic traits in pigs. BMC Genet 19(1):41. https://doi.org/10.1186/s12863-018-0629-4
Sutarno JM, Greeff J, Lymbery AJ (2002) Mitochondrial DNA polymorphisms and fertility in beef cattle. Theriogenology 57(6):1603–1610. https://doi.org/10.1016/s0093-691x(02)00664-7
Tamassia M, Nuttinck F, May-Panloup P, Reynier P, Heyman Y, Charpigny G, Stojkovic M, Hiendleder S, Renard JP, Chastant-Maillard S (2004) In vitro embryo production efficiency in cattle and its association with oocyte adenosine triphosphate. Biol Reprod 71(2):697–704. https://doi.org/10.1095/biolreprod.103.026104
Tsai TS, Rajasekar S, St John JC (2016) The relationship between mitochondrial DNA haplotype and the reproductive capacity of domestic pigs (Sus scrofa domesticus). BMC Genet 17(1):67. https://doi.org/10.1186/s12863-016-0375-4
Wang K, Liu D, Hernandez-Sanchez J, Chen J, Liu C, Wu Z, Fang M, Li N (2015) Genome wide association analysis reveals new production trait genes in a male duroc population. PLoS One 10(9):e0139207. https://doi.org/10.1371/journal.pone.0139207
Wu GS, Yao YG, Qu KX, Ding ZL, Li H, Palanichamy MG, Duan ZY, Li N, Chen YS, Zhang YP (2007) Population phylogenomic analysis of mitochondrial DNA in wild boars and domestic pigs revealed multiple domestication events in East Asia. Genome Biol 8(11):R245. https://doi.org/10.1186/gb-2007-8-11-r245
Wu J, Platero Luengo A, Gil MA, Suzuki K, Cuello C, Morales Valencia M, Parrilla I, Martinez CA, Nohalez A, Roca J, Martinez EA, Izpisua Belmonte JC (2016) Generation of human organs in pigs via interspecies blastocyst complementation. Reprod Domest Anim 51:18–24. https://doi.org/10.1111/rda.12796
Xing K, Zhu F, Zhai LW, Chen SK, Tan Z, Sun YY, Hou ZC, Wang CD (2016) Identification of genes for controlling swine adipose deposition by integrating transcriptome, whole-genome resequencing, and quantitative trait loci data. Sci Rep 6:23219. https://doi.org/10.1038/srep23219
Yen NT, Lin CS, Ju CC, Wang SC, Huang MC (2007) Mitochondrial DNA polymorphism and determination of effects on reproductive trait in pigs. Reprod Domest Anim 42(4):387–392. https://doi.org/10.1111/j.1439-0531.2006.00797.x
Yu G, Xiang H, Wang J, Zhao X (2013) The phylogenetic status of typical Chinese native pigs: analyzed by Asian and European pig mitochondrial genome sequences. J Anim Sci Biotechnol 4(1):9. https://doi.org/10.1186/2049-1891-4-9
Yu G, Xiang H, Tian J, Yin J, Pinkert CA, Li Q, Zhao X (2015) Mitochondrial haplotypes influence metabolic traits in porcine transmitochondrial cybrids. Sci Rep 5:13118. https://doi.org/10.1038/srep13118
Zhang X, Li Z, Yang H, Liu D, Cai G, Li G, Mo J, Wang D, Zhong C, Wang H, Sun Y, Shi J, Zheng E, Meng F, Zhang M, He X, Zhou R, Zhang J, Huang M, Zhang R, Li N, Fan M, Yang J, Wu Z (2018) Novel transgenic pigs with enhanced growth and reduced environmental impact. eLife. https://doi.org/10.1186/2049-1891-4-9
Acknowledgements
This study was funded by the Grant of the President of Russian Federation for supporting of young Russian scientists-candidates of sciences (contract No. MK-1443.2018.11).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The authors declare that animal tissue samples were collected by trained personnel under strict veterinary rules. Sampling was performed in accordance with the ethical guidelines of the L.K. Ernst Federal Science Center for Animal Husbandry.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kolosova, M.A., Getmantseva, L.V., Bakoev, S.Y. et al. Associations of mtDNA haplotypes with productive traits in pigs. Rend. Fis. Acc. Lincei 30, 807–813 (2019). https://doi.org/10.1007/s12210-019-00853-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12210-019-00853-1