Abstract
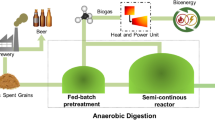
This study investigated the anaerobic co-digestion (AcoD) of brewery by-products for biomethane and bioenergy recovery, focusing on operational performance evaluation, kinetic analysis, microbial metataxonomic, and metabolic function prediction. The biochemical methane potential was conducted under mesophilic (35 °C) and methanogenic conditions (pH 7.5) by mixing brewery wastewater and sludge from the brewery wastewater treatment plant (1:1, v/v), following the addition (2.5 – 12.5 %, w/v) of brewer’s spent grains (BSG). The results demonstrate that the highest methane yield (88.02 mL CH4/g TVS) was obtained with 12.5 % BSG, which was 20.66-fold higher than the control reactor operated with wastewater and sludge (4.26 mL CH4/g TVS). The bioenergy recovery from biomethane could generate electricity (0.348 kWh/kg TVS) and heat (1556 MJ/kg TVS), avoiding greenhouse gas emissions (0.114 kg CO2-eq/kg TVS). The microbial community dynamics revealed a predominance of Halobacterota, Chloroflexi, and Euryarchaeota phylum. The genera Methanosaeta and Methanobacterium, and the Anaerolineaceae family predominated in the AcoD process. The metabolic function prediction showed the presence of genes (K01895, K00193, K00625, and K00925) associated with the direct activation of acetate in the acetoclastic pathway and methane production. Finally, the data obtained provide a perspective on using brewery by-products for bioenergy production in a biorefinery concept, reducing the environmental impacts and contributing to the circular bioeconomy transition of the beer industry.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The beer industry produces a high amount of wastewater and solid by-products, including brewer’s spent grains (BSG), spent hops, and yeast, making it one of the most water- and energy-intensive industrial activities [1]. In 2020, worldwide beer production was estimated at 1.82 billion hL/year [2]. In the case of a brewery located in Seville (Spain), 4,225,245 hL of beer was produced in 2020, generating 75,984 tons BSG and 981,017 m3 wastewater. For this, 32,258 MWh electricity and 222,179 GJ heat were necessary annually to supply the energy demand. This industrial sector faces the challenge of novel and sustainable technologies that can treat the by-products and produce renewable energy, aiming to meet the Sustainable Development Goals toward a circular bioeconomy transition.
BSG contains (dry weight) fibers (20–70%), hemicellulose (20–35%), cellulose (12–25%), lignin (10–28%), and proteins (19–30%) as major compounds [3]. Brewery wastewater can present high organic matter (50 g O2/L) and alkalinity (6 g CaCO3/L) as the most expressive compounds [4]. The management of BSG and wastewater generated can be an alternative for the recovery of bioenergy in the form of methane and hydrogen using anaerobic digestion technology, being cost-effective and feasible raw materials for a biorefinery [5]. However, due to the high lignocellulose content, BSG is difficult to degrade anaerobically as a mono-substrate and can lead to ammonia inhibition due to the high protein content [6]. Due to the different physicochemical characteristics, the mixture of substrates can be a suitable approach to improve biogas production due to the nutrient balance in the reactor. Therefore, anaerobic co-digestion (AcoD) of BSG with other substrates generated in the brewery (e.g., wastewater and sludge from the wastewater treatment plant) can become a reliable alternative to sustainable waste management in a biorefinery advocating a circular bioeconomy.
In a biorefinery concept, the by-products generated in conventional processes (e.g., BSG, wastewater, and sludge generated by breweries) can be used as feedstock to produce bioenergy and value-added products. A biorefinery approach can be used to design sustainable bioprocesses for energy recovery and material recycling, minimizing negative environmental effects and maximizing the use of renewable resources. In addition, anaerobic digestion technology produces biogas, which can be converted into biomethane, electricity, and thermal energy. From an environmental standpoint, using biogas to reduce greenhouse gas (GHG) emissions is a positive strategy to ameliorate the environmental impacts of the food processing industry [5].
Notwithstanding, a diverse and stable microbial community in anaerobic reactors is necessary for the conversion of substrates into biogas, especially in the case of methanogens, where the microorganisms are the most sensitive anaerobes to process fluctuations [7]. Active microorganisms can serve as an important indicator in the operation performance of reactors, and identifying the dynamics of microbial populations can support decision-making to understand better, regulate, and develop controls for the anaerobic digestion process [8]. Next-generation deoxyribonucleic acid (DNA) sequencing analyses allow reconciling the taxonomic signature of the community in understanding the physicochemical characteristics of AcoD, since specific metabolic pathways of the microbial population can affect the overall effectiveness of biogas production [9]. Several tools have been developed to determine the functional inference of 16S rRNA data, such as PICRUSt2 and Tax4Fun, based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) to elucidate the metabolic functions of microorganisms in the process [10].
Based on the abovementioned, this study evaluated the AcoD of brewery by-products for biomethane and bioenergy recovery. In order to combine available industrial by-products in a biorefinery concept, the AcoD of brewery wastewater with various BSG concentrations was evaluated, using the sludge produced during the treatment of brewery wastewater as inoculum for the anaerobic microorganisms. This study focuses on identifying the profile of the microbial community associated with metabolic function and its correlation with operational parameters, methane production, and bioenergy potential.
Materials and Methods
Inoculum and Feedstock
Ambev Brewery (Jaguariúna, SP, Brazil) provided the brewery solid waste (BSG, wet basis). The BSG was oven-dried (105 °C, 8 h), packed, and stored (−18 °C) for later use. The raw brewery wastewater (without treatment) was provided by Heineken Brewery (Seville, Spain). The mesophilic inoculum (granular sludge) used for AcoD was obtained from the treatment of brewery wastewater in up-flow anaerobic sludge blanket digestion (UASB) reactors under mesophilic temperature (35 °C) (Heineken, Seville, Spain). Table 1 displays the initial characterization of the inoculum and substrates used in the experiment.
Biochemical Methane Potential
The biochemical methane potential (BMP) evaluated the addition of BSG in the AcoD of wastewater and sludge from the brewery wastewater treatment. The batch reactor (250 mL) was operated with 120 mL of working volume and 130 mL of headspace for biogas storage. The started-up was conducted with a mixing ratio of 1:1 (v/v) between mesophilic inoculum (60 mL) and raw wastewater (60 mL), following the addition of BSG (2.5, 5, 7.5, and 12.5%, in dry weight). The BMP tests were coded as follows: BMP-1, control reactor (AcoD of inoculum and wastewater, 1:1 v/v); BMP-2, addition of 2.5% BSG in the reactor (4.79 g BSG/L); BMP-3, the addition of 5% BSG in the reactor (9.6 g BSG/L); BMP-4, addition of 7.5% BSG in the reactor (14.39 g BSG/L); and BMP-5, addition of 12.5% BSG in the reactor (24 g BSG/L). The endogen control (negative control) was composed of inoculum (60 mL) and water (60 mL) (without brewery wastewater and BSG). All the experiment was conducted in triplicate (n=3). Table 1 describes the composition of the reactors without inoculum.
The pH of the reactors was adjusted to 7.5 (methanogenic conditions) with the addition of sodium hydroxide (6 mol/L). The reactors were flushed with N2 gas for 30 s to ensure anoxic conditions. The BMP was conducted in an orbital shaking incubator at 85 rpm and 35 °C (mesophilic temperature). The operational performance was conducted before and after BMP tests. The biogas volume and composition were measured daily until the methane production was < 1% of accumulated methane.
Operational Performance
Analytical Methods
The Standard Methods for the Examination of Water and Wastewater [11] were used to characterize the inoculum, substrates, mixtures, and digestate. The following parameters were determined: pH, total solids (TS), total volatile solids (TVS), total fixed solids (TFS), total chemical oxygen demand (TCOD), soluble chemical oxygen demand (SCOD), alkalinity, ammonia (NH3), ammoniacal nitrogen (N-NH3), ammonium (NH4+). The TVS and COD removal were determined according to Eqs. 1 and 2, respectively. The volatile fatty acids (VFA) were determined by gas chromatography equipped with a flame ionization detector, according to the methodology previously described by Sganzerla et al. [4]. The concentration of acetic, propionic, isobutyric, butyric, isovaleric, valeric, caproic, and hexanoic acids was performed using standard calibration curves, and the total content of VFA was calculated and expressed as equivalent in acetic acid.
Biogas Volume and Composition
The pressure of the reactors was measured daily. Under normal conditions, the pressure can be converted to volume using the ideal law of gases (Eq. 3).
where P is the absolute pressure measured in the reactors (kPa), V is the volume (m3), n is the amount of substance (mol CH4), T is the temperature (K), and R is the universal gas constant (8.3145 L kPa/K mol).
The biogas composition was determined daily with a Geotech Biogas 5000 gas analyzer (GasDetect, Copenhagen, Denmark). The volume of methane produced in the endogen control was subtracted from the volume obtained from BMP-1 until BMP-5. The accumulated methane volume (Eq. 4) and the methane yield (Eq. 5) were determined.
where V is the volume of biogas (mL), n is the number of days analyzed, CH4 is the methane content in the biogas (%), MY is the methane yield, and TVS is the content of volatile solids in the reactor.
Kinetic Analysis
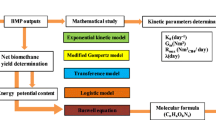
The methane production was evaluated by kinetic analysis using the modified Gompertz (Eq. 6), Cone (Eq. 7), and first-order kinetics (Eq. 8) models. The kinetic analysis was conducted using SigmaPlot® software (Systat Inc., Palo Alto, CA, USA).
where M is the cumulative methane volume (mL); P is the methane production potential (mL); Rm is the maximum methane production rate (mL/h); e = 2.718; λ is the lag phase time (h); t represents the fermentation time (h); kmethane is the hydrolysis rate constant (1/h); and n is the shape factor.
Bioenergy Recovery and Avoided Greenhouse Gas Emissions
The biomethane produced by AcoD can be applied in a co-generator to produce electricity (Eq. 9) and heat (Eq. 10) [12].
where MY is the methane yield (m3 CH4/kg TVS); \({\textrm{LCV}}_{{\textrm{CH}}_4}\) is the lower calorific value of methane (35.59 MJ/m3); ηe is the engine efficiency (%), and CF is the conversion factor from MJ to MWh (1 MWh = 3600 MJ).
The amount of GHG emissions that could be reduced by switching from grid-supplied electricity to locally produced energy from the burning of biogas were estimated according to Eqs. 11 for electricity and Eq. 12 for heat [13]
where EFElectricity is the emission factor of CO2-eq for 2019 Brazilian electric energy generation (0.075 ton CO2-eq/MWh), and EFHeat is the emission factor for heat, assumed as the default value of natural gas (0.056 ton CO2-eq/GJ) [14].
Microbial Community Dynamics and Metabolic Function Prediction
At the end of the BMP, samples (50 mL) were collected and stored at -20 °C for DNA extraction. The samples were centrifuged (9000 rpm, 5 min), and the pellet was used for extraction and purification processes. Using the DNeasy PowerSoil Kit (QIAGEN) and following the manufacturer's instructions, genomic DNA was extracted. The Qubit® fluorometer measured the genetic material (Thermo Fisher Scientific). The DNA samples were evaluated by next-generation sequencing of the V3-V4 region from the 16S rRNA gene for the Archaea and Bacteria domains [15], via MiSeq Sequencing System (Illumina Inc., USA), using primers 341F (CCTACGGGRSGCAGCAG) and 806R (GGACTACHVGGTWTCTAAT) with V2×500 cycles and 100k reads per sample.
The bioinformatics analysis was processed using the FastQC v.0.11.8 to assess nucleotide quality and QIIME 2 pipeline. The Amplicon Sequence Variant (ASV) and the relative abundance values were used to perform the taxonomic signature. The taxonomic annotation was processed using the SILVA database and the feature-classifier plugin from the classify sklearn method (v. 138) [16]. The sequences were deposited at NCBI database under the bioproject accession PRJNA870110. Functional prediction analyses were conducted using the software PICRUSt2 in the predefined settings (default), using the ASV generated in the denoising step [17]. The resulting data were normalized by the number of 16S rRNA copies, while gene inference was conducted using the Kyoto Encyclopedia of Genes and Genomes (KEGG) Orthology (KO) database [18]. BURRITO software was used to obtain the metabolic functions associated with the identified genes, following the KEGG Brite Hierarchy [19].
Statistical Analysis
The statistical analysis was assessed to determine the differences in the TVS removal, COD removal, and methane yield during AcoD of brewery by-products. Analysis of variance (ANOVA) was employed to assess statistically significant factors and interactions between the variables. Tukey's test evaluated significant differences between the samples (p ≤ 0.05). The statistical analysis was conducted using Statistica® software (version 10.0, StatSoft Inc., Tulsa, OK, USA).
Results and Discussion
Characterization and Operational Performance
Table 2 shows the characterization of the BMP tests at the beginning and end of AcoD. The AcoD was evaluated by the pH, alkalinity, solids (TS, TVS, and TFS), ammonia nitrogen (NH3, N-NH3, and NH4+), chemical oxygen demand (TCOD and SCOD), and VFA.
pH plays an essential role in the growth of microorganisms that affect the AcoD process. Therefore, it was necessary to maintain the pH in stable ranges (between 7 and 8) to provide a suitable environment for the development of methanogenic bacteria. During the methanogenic AcoD of brewery by-products the pH was maintained within the appropriate scope. The decrease in pH during AcoD was associated with the production of organic acids by acidogenic bacteria, which were subsequently converted to methane. The stable pH range reported in this study provided a suitable environment for developing a microbial community favorable for methane production. The activity of methanogens producing alkalinity during the AcoD process in the form of carbon dioxide, ammonia, and bicarbonate counteracts pH fluctuations related to VFA concentration [20]. Alkalinity values increased in all reactors during the AcoD, reaching values between 3020 and 4841 mg CaCO3/L. The increase in alkalinity during the methanogenic anaerobic digestion of BSG was associated with the biodegradation of the lignocellulose and organic matter in the anaerobic reactor [21, 22]. In addition, the increase in alkalinity was proportional to the addition of BSG in the reactors, with the highest values corresponding to BMP-5. These results can be explained by the protein content of BSG, which was hydrolyzed during AcoD and released ammonia, causing an increase in alkalinity [23]. In addition, there are low VFA concentrations at the end of the process (Table 3), which can be associated with the high buffer capacity of the system, presenting a low inhibition potential for methanogens [24]. Acetic, propionic, and butyric acids were consumed during the AcoD, demonstrating that the VFA was converted into methane during the methanogenesis stage [25]. In this study, acetic acid was the major VFA in the reactors, corroborating with previous studies on the anaerobic digestion of BSG [22].
The hydrolysis of the proteins causes an increase in NH3, N-NH3, and NH4+ (Table 2). The critical ammonia nitrogen concentration initiating inhibition has been reported to range from 1 to 14 g/L [26]. The N-NH3 content is crucial to avoid inhibition of methanogenic activity, and values of approximately 200 mg/L are recommended for sufficient provision of nitrogen for microbial growth [26]. In this study, the highest value recorded at the end of AcoD was for BMP-5 (868.7 mg N-NH3/L), which can be associated with the high level of acclimatization by the microorganisms, corroborating the methane produced. In a previous study, the ammonia nitrogen in the anaerobic reactor operated with BSG ranged from 106.4 to 260.68 mg/L [22], corroborating the results obtained in the present study. Otherwise, the combined NH3 and NH4+ content, known as total ammoniacal nitrogen, must be higher than 500 mg/L and lower than 3000 mg/L [27] to avoid a possible inhibition of methanogenic activity. This condition was fulfilled in all BMP of brewery by-products.
Regarding the composition of solids, the reactors that started with brewery wastewater (BMP-1) had a low TS content (4.02%). This value increased proportionally with the addition of BSG, reaching 6.74% TS for the reactor with 25% BSG. One of the most critical parameters during the AcoD is the removal of TVS, a parameter widely applied to express the efficiency of anaerobic processes. High TVS removal indicated that the microbial community used the available nutrients in the substrate to produce biogas. In this study, the TVS removal was approximately 40% (Fig. 1a). In the case of the reactor with the highest TVS content (BMP-5), the reduction was 42.26%, which was statistically equal to that of the reactor with the lowest TVS content (BMP-1). The TVS removal obtained in this study was similar to other studies on the anaerobic digestion of sludge, which obtained 37–42% of TVS removal [28]. Previous studies on methanogenic anaerobic digestion of BSG demonstrated that solid biodegradation could reach values higher than 50% [29], demonstrating that BSG is a suitable feedstock for biogas production.
COD is another critical parameter representing substrate solubilization, governing AcoD effectiveness. COD is defined as the amount of oxygen required to oxidize an organic compound, and it is widely used to determine the quality parameters in wastewater treatment facilities and as a balanced unit in anaerobic reactors. Decreasing COD is desirable during AcoD, indicating that complex organic matter is being converted into biogas and other aqueous products that are more suitable for use as renewable fertilizers [30]. In this study, the reactors started up with a TCOD content ranging from 3985.20 (BMP-5) to 5116.80 mg/L (BMP-1), which can be considered suitable values for anaerobic reactors [31]. At the end of AcoD, the TCOD and SCOD removal reached almost 100% for all the reactors (Fig. 1b). Values above 80% of COD removal are considered positive for anaerobic digestion [32]. This fact demonstrates that AcoD can be a suitable process to reduce the organic matter of brewery by-products under the concentrations evaluated. The methanogenic microorganisms consumed the organic matter and converted it into soluble compounds, VFA, and methane, demonstrating the effectiveness of AcoD for the treatment of brewery by-products.
Methane Production and Yield
The start-up of the AcoD corresponds to biogas production and organic matter degradation by microbial organisms. Fig. 2 shows the methane production and yield from the AcoD of brewery by-products. On the first day of AcoD, methane production was noticeable. These results can be associated with the exponential growth of methanogenic microorganisms and the adaptation under methanogenic and mesophilic conditions, as they can produce biomethane immediately after start-up [24]. After the continuous consumption of substrate by microbiota under anaerobic conditions, a decrease in methane production was recorded after approximately 10 days of AcoD. The production of methane finished after 17 days of digestion. In this study, the reactor with the highest amount of BSG (BMP-5) resulted in a better methane profile. The result indicated that BSG is an easier biodegradable substrate for AcoD under methanogenic and mesophilic conditions. In a previous study on the AcoD of brewery by-products under acidogenic and thermophilic conditions, the addition of BSG promoted the inhibition of hydrogen production due to ammonia production [4].
The maximum methane yield obtained was 88.03 mL CH4/g TVS for BMP-5 (Fig. 2c). These results were similar to the AcoD of activated sludge with granular sludge, cow manure, and food waste [33] and higher than previous studies on the dry anaerobic digestion of BSG without wastewater (26.72 – 39.51 mL CH4/g TVS) [14, 29]. The addition of biodegradable substrate in AcoD enhanced the biogas yield due to improved nutrient balance and diluting toxic compounds in the reactor. Moreover, the supplementation of nutrients contained in BSG was suitable for microbial growth due to the balance of nutrients, and it is more beneficial to the degradation of organic matter. The results from this study indicate that BSG is highly anaerobically digestible with brewery wastewater and makes a co-digestion process viable to increase methane production.
For a feasible industrial-scale application of AcoD, the methane yield should be higher than 100 mL CH4/g TVS [34]. However, for the industrial-scale application of AcoD of brewery by-products, the methane yield in a laboratory-scale semi-continuous process should be evaluated. The results obtained in this batch study are the first approach to determine the BMP and initial operational conditions, especially the amount of BSG that should be added in the start-up of semi-continuous reactors. Hence, further studies should be conducted to optimize the hydraulic retention time and organic loading rate in the temperature-phase semi-continuous process, aiming to increase the methane yield at feasible ranges for industrial-scale implementation.
Kinetic Analysis
The kinetic parameters offer additional insightful data on the biodegradation patterns of the substrates and methane production. The efficiency of methane production during AcoD has been widely determined by the modified Gompertz, Cone, and first-order kinetic models. In this study, these kinetic models were applied to predict methane production (Table 4).
The methane production achieved at the end of the experimental BMP assays was compared with the maximum methane production obtained from the kinetic models. The parameter P showed the estimated methane production for each test performed and was entirely accurate in all three models, as the percentage difference between the estimated values and the experimental values obtained was lower than 20 % in all tests. Based on these results, it can be affirmed that all the kinetics models accurately predicted the behavior of methane production in the AcoD of brewery by-products, with the modified Gompertz model being the most accurate for the tested substrate.
The maximum methane production rate (Rm) increased according to the concentration of BSG in the reactors, presenting its maximum value for BMP-4 (1.292 mL/h) and decreasing slightly for BMP-5. The maximum values of Rm corresponded to the test with the lowest starting organic load. Values of λ were zero in all BMP tests. This value indicates that methane production began immediately upon assembly of the assays, without initial inhibition or need for acclimatization by the microbial community. Concerning R2, the first-order model and the Cone model presented values higher than 0.99, indicating a high correlation between the predicted and experimental data. However, the modified Gompertz model showed high variable R2 values, with the maximum for BMP-1 (0.993) and the minimum for BMP-5 (0.807). RMSE indicates the association between the observed and predicted data by kinetic fitting. The modified Gompertz model presented the highest RMSE values, and the RMSE increased according to the addition of BSG in the reactor. Although all kinetic models showed valuable data for the AcoD of brewery by-products, based on the R2 and RMSE, the model with the highest efficiency in data fitting was the Cone model. It should be noted that each study presents peculiarities and that there is no agreement in the literature on the most appropriate model in each case. Thus, several models should be studied to predict the production of methane during AcoD [32].
Bioenergy Potential and Environmental Benefits
Biomass, as a renewable energy source, plays an essential role in reducing GHG emissions, especially for heat and electricity generated from biogas in co-generator. Based on the methane yield, the recovery of electricity, heat, and avoided GHG emissions were estimated (Table 5). The process operated with the highest amount of BSG (17.5%) presented the highest methane potential and, consequently, the highest production of electricity (0.348 kWh/kg TVS) and heat (1.556 MJ/kg TVS). The replacement of conventional energy by the bioenergy generated from biogas could mitigate up to 1.14×10–1 kg CO2-eq/kg TVS, with 2.61×10–2 kg CO2-eq/kg TVS for the use of electricity and 8.77×10–2 kg CO2-eq/kg TVS for the use of heat. Therefore, the AcoD of brewery by-products for methane production could be an initial approach to promote the beer industry’s circular bioeconomy transition, with the sustainable management of industrial by-products and bioenergy recovery. This approach represents a positive strategy for self-production and energy consumption and is a promising technological route to be applied in a biorefinery.
Microbial Community in Anaerobic Co-diestion Process
A total of 82,843 and 50,537 reads were retrieved for the inoculum and BMP-5, respectively, after the denoising step. Shannon index 3.077 (inoculum) and 4.147 (BMP-5) indicated increased diversity in the microbial consortium in AcoD. The Chao1 index was the same (178) for both samples. The diversity analysis revealed 9 different phyla, 7 from Bacteria and 2 from Archaea domain. Different profiles were observed in the structure of the microbial community of the inoculum and the sample of the BMP-5 assay (Fig. 3). In the inoculum, the phylum Desulfobacterota predominated with an abundance greater than 50%, followed by Halobacterota (20%) and Euryarchaeota (8.14%). Desulfobacterium comprises a variety of sulfate-reducing bacteria commonly found in sewage treatment plant sludg. Sulfate-reducing bacteria compete with methanogenic Archaea, due to a higher growth rate and better substrate affinity [25]. In the AcoD (BMP-5), there was a predominance of the phylum Halobacterota (29.84%), Chloroflexi (14.53%), Euryarchaeota (13%), Synergistota (11.87%) and reduction of Desulfobacterota (16.97%) and consequently greater volume of methane produced (Fig. 2). Chloroflexi members can hydrolyze complex compounds, such as grains used in the brewery, transforming them into VFA, carbon dioxide, and hydrogen. The heterotrophic growth of this phylum is related to the acetic acid content in the system. In addition, this group can also form granular structure sludge, an important factor in amplifying the methane production capacity [35].
The phylum Halobacterota was found with greater abundance in BMP-5 than in the inoculum, which may be due to the presence of acetic acid since the presence of this VFA was not detected in the inoculum (Table 3). This group comprises methanogens that use acetic acid as an electron donor [36]. This phylum is responsible for approximately 2/3 of the total methane production. In contrast, phylum Euryarchaeota species (13%) comprise hydrogenotrophic species responsible for approximately 1/3 of the total methane yield [36]. The Synergistota phylum is associated with the consumption of butyric and propionic acids, helping the system's stability. The group of synergistic bacteria is primarily involved in methane production and converting glucose, cellobiose, and amino acids into VFA [37].
A great richness was observed with more than 170 genera, with only 15 with an abundance greater than 1% (Fig. 3). Anaerobic digestion is a multi-step process that requires a consortium of syntrophic bacteria and methanogenic archaea. The inoculum microbial community was dominated by Syntrophobacter (48%). This group of syntrophic bacteria has a positive association with methanogenic archaea, and some species can transform propionate into acetate, favoring acetoclastic methanogenesis [38]. In the AcoD process (BMP-5) there was a reduction in the abundance of Syntrophobacter (16.46%), Methanosaeta (28.7%), and Methanobacterium (13.02%) (Fig. 3c). Anaerolineaceae, anaerobic filamentous bacteria, which hydrolyzes different types of carbohydrates or polypeptides, was favored by BMP-5 conditions (13.74%). Anaerolineaceae and Syntrophobacter also participate in interspecies direct electron transfer, which may also contribute to methanogenesis [38].
Methanosaeta is an obligate acetic acid consumer Archaea, dominant at low concentrations of free ammonia and acetate [39], corroborating with the concentrations presented for these compounds (Table 2 and Table 3) since there was a 99% reduction of acetic acid at the end of the process (Table 3). Acetoclastic methanogenesis is considered an important methane production pathway since archaea live in a symbiotic relationship with syntrophic oxidant bacteria that produce acetate and H2, and methanogens convert it into methane, which prevents inhibition of syntrophic bacteria by the excess of acetate [40]. Methanobacterium is a hydrogenotrophic methanogen that uses H2 and CO2 to produce CH4 and tends to increase with high carbohydrate content and decreasing lipid and protein content [41]. The abundance of Methanosaeta was 2-fold higher than Methanobacterium, indicating that Methanosaeta was the main genus involved in methane production. These results corroborate the literature, where a predominance of the genus Methanosaeta in effluent from the brewery was associated with the production of biogas [42]. The genus of anaerobic bacteria Aminiphilus showed a relative abundance of 4.79% in the BMP-5 sample. Recent studies describe that this group can ferment a variety of VFA and amino acids and settle in neutral environments [43], corroborating with the experimental results obtained in this study (Table 2). In addition, Aminiphilus has been reported in brewery effluent UASB reactors [44].
Metabolic Function Prediction
A total of 5,906 different KO objects were obtained for inoculum and BMP5. The main metabolic pathways of the reactors were predicted using the KEGG Orthology database. Fig. 4a shows several genes related to the methanogenesis process for the studied microbial communities. Genes associated with the acetoclastic pathway, including acetyl-CoA synthetase (K01895), acetyl-CoA decarbonylase/synthase (K00193), acetate kinase (K00925), phosphate acetyltransferase (K00625), which are crucial for the direct activation of acetate [45] were found in greater abundance in BPM5. Enzymes involved in hydrogenotrophic methanogenesis, such as formylmethanofuran dehydrogenase (K00201/ K00202), coenzyme F420 hydrogenase subunit beta (K00441), 5,10-methylenetetrahydromethanopterin reductase (K00320) and formylmethanofuran--tetrahydromethanopterin N-formyltransferase (K00672) were also predicted in greater abundances in BMP5, which evidences the participation of H2-consuming archaea in the AD process. Fig. 4a shows Ids of the genes (K00169; K00170; K00171; K00172; K00672; K01499; K00319; K00320; K0041) that encode enzymes of the hydrogenotrophic pathway also described for the genus Methanosaeta [38]. While Methanobacterium, a typical hydrogenotrophic, also contains genes (K01895; K00193-K00197) related to acetoclastic methanogenesis, as evidenced by Gaio et al. [46] in UASB reactors treating domestic sewage. Genes common to acetoclastic and hydrogenotrophic metabolic pathways were also detected in greater abundance in BMP5. As well as other Ids (KO) described in the literature as participants in methane metabolism [10]. Genes related to the methylotrophic pathway showed an abundance close to zero (data not shown in Fig. 4a), following the Archaea genera identified in this study.
Among the main functions identified in the KEGG category, it was observed that the abundances were quite similar in both samples (Fig. 4b). Cellular functions related to translation (17%) and membrane transport (15%) were predominant in the inoculum and BPM-5. Metabolism of cofactors and vitamins, carbohydrates, lipids, and amino acids were equivalent to 19% in both reactors, which indicates the provision of available substrates for acidogenesis and subsequent methane production. The metabolism of cofactors and vitamins can promote the catalytic reaction of functional cellular enzymes, thereby accelerating the anaerobic reaction process [47]. Energy metabolism (equivalent to 4%) is inevitably required by cells to maintain basic metabolism [48]. In comparison, the subcategory methane metabolism was 22% higher in BMP-5. Cell motility was among (1.77 – 2.4), possibly related to some genes that encode methanogenesis [49]. The predominance of various metabolisms can be attributed to the composition of BSG (hemicellulose, cellulose, lignin, and proteins) used in AD processes [50]. The taxonomic analysis of microbial communities associated with the main metabolic functions involved in the encoding of important enzymes in the BSG bioconversion process shows the possible anaerobic digestion routes used.
Conclusion
Brewery by-products were subjected to AcoD to assess the BMP under mesophilic and methanogenic conditions. Bioenergy production with the methane generated from the AcoD combined with the knowledge of the microorganisms involved in the process could contribute to avoiding GHG emissions and decrease the carbon footprint of the beer industry. Finally, BSG can be used as a co-substrate in the AcoD of brewery wastewater and sludge from wastewater treatment plants, improving biomethane and bioenergy production, advocating the circular bioeconomy transition of the beer industry in a biorefinery concept.
Data Availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Berry Z, Loughrin J, Burris S et al (2022) Improving Anaerobic Digestion of Brewery and Distillery Spent Grains through Aeration across a Silicone Membrane. Sustainability 14:2755. https://doi.org/10.3390/su14052755
Zeko-Pivač A, Tišma M, Žnidaršič-Plazl P et al (2022) The Potential of Brewer’s Spent Grain in the Circular Bioeconomy: State of the Art and Future Perspectives. Front Bioeng Biotechnol 10:870744. https://doi.org/10.3389/fbioe.2022.870744
Amoriello T, Ciccoritti R (2021) Sustainability: Recovery and Reuse of Brewing-Derived By-Products. Sustainability 13:2355. https://doi.org/10.3390/su13042355
Sganzerla WG, Sillero L, Forster-Carneiro T et al (2022) Determination of Anaerobic Co-fermentation of Brewery Wastewater and Brewer’s Spent Grains for Bio-hydrogen Production. Bioenergy Res:1–11. https://doi.org/10.1007/s12155-022-10486-2
Sganzerla WG, Ampese LC, Mussatto SI, Forster-Carneiro T (2021) A bibliometric analysis on potential uses of brewer’s spent grains in a biorefinery for the circular economy transition of the beer industry. Biofuels Bioprod Biorefin 15:1965–1988. https://doi.org/10.1002/bbb.2290
Bachmann SAL, Calvete T, Féris LA (2022) Potential applications of brewery spent grain: Critical an overview. J Environ Chem Eng 10:106951. https://doi.org/10.1016/j.jece.2021.106951
Tyagi VK, Bhatia A, Kubota K et al (2021) Microbial community dynamics in anaerobic digesters treating organic fraction of municipal solid waste. Environ Technol Innov 21:101303. https://doi.org/10.1016/j.eti.2020.101303
Adames LV, Jacobus AP, Sakamoto IK et al (2022) Bioenergy Recovery from Anaerobic Co-Digestion of Crude Glycerol and Domestic Sewage In-Series Reactor: Microbial Characterization and System Performance. Bioenergy Res 15:2145–2158. https://doi.org/10.1007/s12155-022-10417-1
Silva Rabelo CAB, Camargo FP, Sakamoto IK, Varesche MBA (2023) Metataxonomic characterization of an autochthonous and allochthonous microbial consortium involved in a two-stage anaerobic batch reactor applied to hydrogen and methane production from sugarcane bagasse. Enzyme Microb Technol 162:110119. https://doi.org/10.1016/j.enzmictec.2022.110119
Bedoya K, Hoyos O, Zurek E et al (2020) Annual microbial community dynamics in a full-scale anaerobic sludge digester from a wastewater treatment plant in Colombia. Sci Total Environ 726:138479. https://doi.org/10.1016/j.scitotenv.2020.138479
APHA - American Public Health Association (2017) Standard Methods for the Examination of Water and Wastewater, 23rd ed. American Public Health Association, American Water Works Association, and Water Environment Federation, Washington
Campello LD, Barros RM, Tiago Filho GL, dos Santos IFS (2021) Analysis of the economic viability of the use of biogas produced in wastewater treatment plants to generate electrical energy. Environ Dev Sustain 23:2614–2629. https://doi.org/10.1007/s10668-020-00689-y
Silva dos Santos IF, Braz Vieira ND, de Nóbrega LGB et al (2018) Assessment of potential biogas production from multiple organic wastes in Brazil: Impact on energy generation, use, and emissions abatement. Resour Conserv Recycl 131:54–63. https://doi.org/10.1016/j.resconrec.2017.12.012
Buller LS, Sganzerla WG, Lima MN et al (2022) Ultrasonic pretreatment of brewers’ spent grains for anaerobic digestion: Biogas production for a sustainable industrial development. J Clean Prod 355:131802. https://doi.org/10.1016/j.jclepro.2022.131802
Caporaso JG, Lauber CL, Walters WA et al (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624. https://doi.org/10.1038/ismej.2012.8
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian Classifier for Rapid Assignment of rRNA Sequences into the New Bacterial Taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Douglas GM, Maffei VJ, Zaneveld JR et al (2020) PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 38:685–688. https://doi.org/10.1038/s41587-020-0548-6
Kanehisa M, Sato Y, Kawashima M et al (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44:457–462. https://doi.org/10.1093/nar/gkv1070
McNally CP, Eng A, Noecker C et al (2018) BURRITO: An Interactive Multi-Omic Tool for Visualizing Taxa–Function Relationships in Microbiome Data. Front Microbiol 9:365. https://doi.org/10.3389/fmicb.2018.00365
Raja Ram N, Nikhil GN (2022) A critical review on sustainable biogas production with focus on microbial-substrate interactions: bottlenecks and breakthroughs. Bioresour Technol Rep 19:101170. https://doi.org/10.1016/j.biteb.2022.101170
Sganzerla WG, Costa JM, Tena-Villares M et al (2022) Dry Anaerobic Digestion of Brewer’s Spent Grains toward a More Sustainable Brewery: Operational Performance, Kinetic Analysis, and Bioenergy Potential. Fermentation 9:2. https://doi.org/10.3390/fermentation9010002
Sganzerla WG, Ampese LC, Mussatto SI, Forster-Carneiro T (2022) Subcritical water pretreatment enhanced methane-rich biogas production from the anaerobic digestion of brewer’s spent grains. Environ Technol 1–19. https://doi.org/10.1080/09593330.2022.2157756
Zhang G, Shi Y, Chen W et al (2021) Methane production from waste activated sludge by combining calcium peroxide pretreatment with zero valent iron bio-enhancement: Performance and mechanisms. J Clean Prod 320:128773. https://doi.org/10.1016/j.jclepro.2021.128773
Shahbaz M, Ammar M, Hussain S et al (2022) Mesophilic batch co-digestion of municipal solid waste and food waste: improved digester performance and stability at optimum organic loading rates. J Chem Technol Biotechnol 97:633–642. https://doi.org/10.1002/jctb.6909
Yang S, Xue W, Liu P et al (2022) Revealing the methanogenic pathways for anaerobic digestion of key components in food waste: Performance, microbial community, and implications. Bioresour Technol 347:126340. https://doi.org/10.1016/j.biortech.2021.126340
Yellezuome D, Zhu X, Wang Z, Liu R (2022) Mitigation of ammonia inhibition in anaerobic digestion of nitrogen-rich substrates for biogas production by ammonia stripping: A review. Renew Sustain Energy Rev 157:112043. https://doi.org/10.1016/j.rser.2021.112043
Ariunbaatar J, Scotto Di Perta E, Panico A et al (2015) Effect of ammoniacal nitrogen on one-stage and two-stage anaerobic digestion of food waste. Waste Manag 38:388–398. https://doi.org/10.1016/j.wasman.2014.12.001
Agabo-García C, Pérez M, Rodríguez-Morgado B et al (2019) Biomethane production improvement by enzymatic pre-treatments and enhancers of sewage sludge anaerobic digestion. Fuel 255:115713. https://doi.org/10.1016/j.fuel.2019.115713
Sganzerla WG, Tena-Villares M, Buller LS et al (2022) Dry Anaerobic Digestion of Food Industry by-Products and Bioenergy Recovery: A Perspective to Promote the Circular Economy Transition. Waste Biomass Valoriz 13:2575–2589. https://doi.org/10.1007/s12649-022-01682-4
Atelge MR, Krisa D, Kumar G et al (2020) Biogas Production from Organic Waste: Recent Progress and Perspectives. Waste Biomass Valoriz 11:1019–1040. https://doi.org/10.1007/s12649-018-00546-0
Pereira da Silva F, Lutterbeck CA, Colares GS et al (2021) Treatment of university campus wastewaters by anaerobic reactor and multi-stage constructed wetlands. J Water Process Eng 42:102119. https://doi.org/10.1016/j.jwpe.2021.102119
Blasius JP, Contrera RC, Maintinguer SI, Alves de Castro MCA (2020) Effects of temperature, proportion and organic loading rate on the performance of anaerobic digestion of food waste. Biotechnol Rep 27:e00503. https://doi.org/10.1016/j.btre.2020.e00503
Gaur RZ, Suthar S (2017) Anaerobic digestion of activated sludge, anaerobic granular sludge and cow dung with food waste for enhanced methane production. J Clean Prod 164:557–566. https://doi.org/10.1016/j.jclepro.2017.06.201
Westerholm M, Liu T, Schnürer A (2020) Comparative study of industrial-scale high-solid biogas production from food waste: Process operation and microbiology. Bioresour Technol 304:122981. https://doi.org/10.1016/j.biortech.2020.122981
Owusu-Agyeman I, Eyice Ö, Cetecioglu Z, Plaza E (2019) The study of structure of anaerobic granules and methane producing pathways of pilot-scale UASB reactors treating municipal wastewater under sub-mesophilic conditions. Bioresour Technol 290:121733. https://doi.org/10.1016/j.biortech.2019.121733
Fan Q, Fan X, Fu P et al (2022) Microbial community evolution, interaction, and functional genes prediction during anaerobic digestion in the presence of refractory organics. J Environ Chem Eng 10:107789. https://doi.org/10.1016/j.jece.2022.107789
Bao R, Wei Y, Guan R et al (2023) High-solids anaerobic co-digestion performances and microbial community dynamics in co-digestion of different mixing ratios with food waste and highland barley straw. Energy 262:125529. https://doi.org/10.1016/j.energy.2022.125529
Du Q, Mu Q, Wu G (2021) Metagenomic and bioanalytical insights into quorum sensing of methanogens in anaerobic digestion systems with or without the addition of conductive filter. Sci Total Environ 763:144509. https://doi.org/10.1016/j.scitotenv.2020.144509
Harirchi S, Wainaina S, Sar T et al (2022) Microbiological insights into anaerobic digestion for biogas, hydrogen or volatile fatty acids (VFAs): a review. Bioengineered 13:6521–6557. https://doi.org/10.1080/21655979.2022.2035986
Kurade MB, Saha S, Salama E-S et al (2019) Acetoclastic methanogenesis led by Methanosarcina in anaerobic co-digestion of fats, oil and grease for enhanced production of methane. Bioresour Technol 272:351–359. https://doi.org/10.1016/j.biortech.2018.10.047
Zhang Y, Guo B, Zhang L et al (2021) Microbial community dynamics in granular activated carbon enhanced up-flow anaerobic sludge blanket (UASB) treating municipal sewage under sulfate reducing and psychrophilic conditions. Chem Eng J 405:126957. https://doi.org/10.1016/j.cej.2020.126957
Enitan AM, Kumari S, Odiyo JO et al (2018) Principal component analysis and characterization of methane community in a full-scale bioenergy producing UASB reactor treating brewery wastewater. Phys Chem Earth, Parts A/B/C 108:1–8. https://doi.org/10.1016/j.pce.2018.06.006
Ban Q, Zhang L, Li J (2019) Functional bacterial and archaeal dynamics at slightly acidic conditions revealed by pyrosequencing in a propionate-fed UASB reactor. Desalin water Treat 160:125–134. https://doi.org/10.5004/dwt.2019.24249
Beraud-Martínez LK, Gómez-Gil B, Franco-Nava MÁ et al (2021) A metagenomic assessment of microbial communities in anaerobic bioreactors and sediments: Taxonomic and functional relationships. Anaerobe 68:102296. https://doi.org/10.1016/j.anaerobe.2020.102296
Li Y, Jing Z, Pan J et al (2022) Multi-omics joint analysis of the effect of temperature on microbial communities, metabolism, and genetics in full-scale biogas reactors with food waste. Renew Sustain Energy Rev 160:112261. https://doi.org/10.1016/j.rser.2022.112261
Gaio J, Lora NL, Iltchenco J et al (2022) Seasonal characterization of the prokaryotic microbiota of full-scale anaerobic UASB reactors treating domestic sewage in southern Brazil. Bioprocess Biosyst Eng 46(1):69–87. https://doi.org/10.1007/s00449-022-02814-9
Li Y, Chen Z, Peng Y et al (2022) Deeper insights into the effects of substrate to inoculum ratio selection on the relationship of kinetic parameters, microbial communities, and key metabolic pathways during the anaerobic digestion of food waste. Water Res 217:118440. https://doi.org/10.1016/j.watres.2022.118440
Ito K, Suda T (2014) Metabolic requirements for the maintenance of self-renewing stem cells. Nat Rev Mol Cell Biol 15:243–256. https://doi.org/10.1038/nrm3772
Town JR, Dumonceaux TJ (2016) Laboratory-scale bioaugmentation relieves acetate accumulation and stimulates methane production in stalled anaerobic digesters. Appl Microbiol Biotechnol 100:1009–1017. https://doi.org/10.1007/s00253-015-7058-3
Li Y, Achinas S, Zhao J et al (2020) Co-digestion of cow and sheep manure: Performance evaluation and relative microbial activity. Renew Energy 153:553–563. https://doi.org/10.1016/j.renene.2020.02.041
Funding
Funding for open access publishing: Universidad de Cádiz/CBUA This work was supported by the Brazilian Science and Research Foundation (CNPq) (productivity grant 302451/2021–8); Coordination for the Improvement of Higher Education Personnel (CAPES, Brazil) (Finance code 001); São Paulo Research Foundation (FAPESP, Brazil) (grant numbers 2018/14938–4 for TFC and 2019/26925–7 for WGS); Andalusian Research, Development, and Innovation Plan (PAIDI, Spain) (grant number P18-RT-1348, CIREc-AD Waste to Energy); and Asociación Universitaria Iberoamericana de Postgrado (AUIP) (Programa de Becas de Movilidad entre Universidades Andaluzas e Iberoamericanas 2022).
Author information
Authors and Affiliations
Contributions
W G S: conceptualization, investigation, data curation, formal analysis, writing—original draft. M T: conceptualization, investigation, writing—original draft. L S: methodology, visualization, writing—original draft. F E M: investigation, data curation. I V M S: writing—original draft. J G: writing—original draft. S P: writing—review and editing. Tânia Forster-Carneiro: writing—review and editing, supervision. R S: writing—review and editing, supervision. M P: writing—review and editing, supervision.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sganzerla, W.G., Tena, M., Sillero, L. et al. Application of Anaerobic Co-digestion of Brewery by-Products for Biomethane and Bioenergy Production in a Biorefinery Concept. Bioenerg. Res. 16, 2560–2573 (2023). https://doi.org/10.1007/s12155-023-10605-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12155-023-10605-7