Abstract
mCherry is one of the most successfully applied monomeric red fluorescent proteins (RFPs) for in vivo and in vitro imaging. However, questions pertaining to the photostability of the RFPs remain and rational further engineering of their photostability requires information about the fluorescence quenching mechanism in solution. To this end, NMR spectroscopic investigations might be helpful, and we present the near-complete backbone NMR chemical shift assignment to aid in this pursuit.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Biotechnological context
Monomeric Red Fluorescent Proteins (mRFPs) are widely used as genetically encodable tags for studying cellular processes. Their distinctive fluorescence results from the chemical rearrangement of amino acids, giving rise to the formation of an acylimine, further modified in different mRFPs. The DsRed precursor (λexcmax = 558nm, λemmax = 583nm) has the disadvantage of being tetrameric and having low photostability and a slow folding rate. For these reasons, several monomeric variants have been engineered to improve performance in terms of photostability and structural stability and to cover a different range of wavelengths. By using random mutagenesis (Shaner et al. 2004) the series of mFruits was obtained, which comprises mCherry (λexcmax = 587nm, λemmax = 610nm), mStrawberry (λexcmax = 574nm, λemmax = 596nm), and mOrange (λexcmax = 548nm, λemmax = 562nm). Crystal structures of these proteins have been obtained (Shu et al. 2006) and these all show the canonical β-barrel structure harboring an α-helix comprising the residues involved in the formation of the chromophore. The β-barrel in the monomeric mCherry variant (PDB code 2h5q) comprises 11 strands with the chromophore formed by the contiguous residues Methionine-Tyrosine-Glycine (collectively referred to as position 66 in the PDB entry and in our sequence numbering). We report here the near-complete assignment of backbone NMR chemical shifts for mCherry. The N-terminus (residues − 4 to 3) and C-terminus (residues 224–231) are not present in the crystallographic structure and are dynamically disordered. The NMR assignments presented here will be used to address the photostability of the mFruits.
Method and experiments
Protein expression and purification
A pET11a plasmid containing the gene of mCherry from Discosona sp. (Uniprot code Q5S3G8) containing an N-terminal poly-His tag MGHHHHHHG was transformed in BL21(DE3) E. coli cells. A single colony was picked and grown overnight at 37°C in Luria-Bertani media, then spun down and resuspended in 50 mL M9 media to grow overnight. After spinning down, 5 parallel growth flasks were prepared; in each, 4 mL of culture was added to 200 mL M9 medium containing the relevant isotopes (1 gr/L NH4Cl and 3 gr/L 13C6-D-glucose). After growing the cells at 37°C for 4h, 0.001M IPTG was added, and the temperature lowered to room temperature (26°C). Expression was stopped after 24h and the protein pellet was purified on Ni-NTA columns, dialyzed in 50mM Tris pH8.0, 2mM DTT, 0.5mM EDTA and then dialyzed in 20mM Tris, 50mM NaCl pH 7.0. Validation of correct expression of the full-length protein was obtained by ESI-TOF mass spectrometry from which a labelling efficiency of 97% was obtained. The protein used for NMR analysis was inclusive of the poly-His tag.
NMR spectroscopy
The NMR sample consisted of 1.5 mM U-15N,13C protein dissolved in 20 mM Tris-HCl, 50 mM NaCl, pH 7.0, 5% D2O and spectra were acquired at 303K with a Bruker Avance III HD spectrometer operating at 950MHz proton frequency, equipped with z-axis gradients and a cryogenic probe head (TCI). The following experiments were collected using Bruker library pulse sequences: 2D 15N-1H HSQC, 3D HNCO, HN(CA)CO, HNCACB, HN(CO)CACB, H(NCA)NH, HN(CO)CA, H(NCOCA)NH, iHNCA, using TROSY (Pervushin et al., 1997) and BEST (Lescop et al., 2007) methodology.
Data were processed with NMRPipe (Delaglio et al. 1995) and spectral assignments were made with NMRFAM-Sparky (Lee et al. 2015). Spectral offsets of 0.5*93 Hz were applied to the amide 15N and 1H dimensions of all TROSY spectra to align them with the correct decoupled chemical shift values of the 2D 15N-1H HSQC spectrum. Chemical shift referencing followed the IUPAC recommendation of the protein NMR community (Markley et al., 1998).
Extent of assignments and data deposition
Despite the presence of more than 236 residues, the 15N-1H TROSY-HSQC spectrum of mCherry is well resolved (Fig.1), and the sensitivity of 3D BEST-TROSY spectra were adequate for the assignment. Strong signals in the H(NCA)NH and H(NCOCA)NH spectra were helpful for assigning the more dynamic regions, including the termini.
Residue numbering “_Atom_chem_shift.Auth_seq_ID” in the BMRB deposition (ID 51489) follows that reported in the crystal structure (PDB code 2h5q), which runs from Met(-4) to Lys231. The cyclized residues Met-Tyr-Gly are collectively referred to as number 66 and the sequence then continues with Ser69. In the crystal structure, the N-terminal region is disordered and the first residues were not observed. NMR spectra allow the observation of 6 residues in the N-terminal region (from Ser(-2) to Asp3) that were lacking in the electron density map. Their NMR chemical shifts confirm that this region is disordered. Similarly, the last eight residues are not observed in the crystal structure but are assigned in our work and are confirmed disordered. Residues 66–76 containing the chromophore could not be assigned. Also, no assignment was obtained for residues 37, 53–54, 80–83, 130 and 221–222. The completeness of assignments is: 1H (199), 15N (199), 13Cα (211), 13Cβ (187), and 13C’ (210). Excluding the His-tag, the protein sequence contains 234 amino acids, of which 26 Gly (having no 13Cβ) and 12 Pro (lacking 1H and yielding no 15N assignments in the triple resonance experiments). The extent of completeness for the aforementioned nuclei is then 86%, 86%, 90%, 90%, and 90%, respectively.
An intial secondary structure analysis was obtained with the TALOS + software (Shen et al. 2009) and compared with the classification obtained with the STRIDE software (Heinig and Frishman 2004) from the crystallographic structure in Fig.2. Good overall agreement is observed.
(a) Secondary structure of mCherry obtained by using the present NMR chemical shift assignment used as input for the TALOS + software. Blue regions refer to β-strands and red regions to α-helices. (b) Secondary structure of mCherry obtained from the crystal structure (PDB code 2h5q) used as input for the STRIDE software. Blue regions refer to β-strands and red regions to α-helices
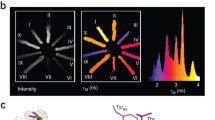
Although canonical helix and strand conformations are easily detected by programs like TALOS+, the combination of 1H, 15N, 13Cα, 13Cβ, and 13C’ chemical shifts contains more information about the backbone conformation. It is possible to recover the eight common structural motifs defined as Dictionary of Protein Secondary Structure (DSSP) by the program CheSPI (Nielsen and Mulder 2021). A CheSPI analysis for mCherry is shown in Fig.3. The top panel (a) shows the much richer structural classification, where colors depend on backbone geometry and structural context (see legend). Furthermore, the heights of the bars (CheZOD score) reflects the dynamic information content of the shift information, with a values of 8 marking the border between order and disorder, and values below 3 indicative of ‘random coil’ dynamic averaging. As can be seen in the central panel (b), the canonical secondary structure elements are well retrieved, but some regions diverge from this. The bottom panel (c) shows a summary in which also coil (grey line), turn (green arc), and 310-helices (magenta squiggle) are identified, in addition to α-helix (red squiggle) and β-strand (blue arrow).
CheSPI analysis for mCherry. (a) On the CheSPI color scale, well-formed strands and helices are defined by blue and red colors, respectively, while coil color depends on context; turns are shown in green, and disordered, ‘random coil’, residues are displayed as grey. Hues change from red through orange to yellow at the C-terminal ends of helices and green at the ends of β-strands. For a more comprehensive explanation of the PCA analysis underlying CheSPI colors, the reader is referred to the paper of Nielsen and Mulder (Nielsen and Mulder 2021). (b) Stacked bar plot of CheSPI populations of “extended” (blue), “helical” (red), “turn” (green), and “non-folded” (grey), local structures (c) CheSPI DSSP-8 assignment. Cartoon of the most confident CheSPI prediction: coil (grey line), turn (green arc), 310-helix (magenta squiggle), α-helix (red squiggle), β-strand (blue arrow)
Data Availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request. The chemical shifts have been deposited with the BioMagResBank (https://bmrb.io/) and are available under entry number 51489.
References
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. https://doi.org/10.1007/BF00197809
Heinig M, Frishman D (2004) STRIDE: a web server for secondary structure assignment from known atomic coordinates of proteins. Nucleic Acids Res 32:W500–W502. https://doi.org/10.1093/nar/gkh429
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinforma Oxf Engl 31:1325–1327. https://doi.org/10.1093/bioinformatics/btu830
Nielsen JT, Mulder FAA (2021) CheSPI: chemical shift secondary structure population inference. J Biomol NMR 75:273–291. https://doi.org/10.1007/s10858-021-00374-w
Shaner NC, Campbell RE, Steinbach PA, Giepmans BNG, Palmer AE, Tsien RY (2004) Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat Biotechnol 22:1567–1572. https://doi.org/10.1038/nbt1037
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223. https://doi.org/10.1007/s10858-009-9333-z
Shu X, Shaner NC, Yarbrough CA, Tsien RY, Remington SJ (2006) Novel chromophores and buried charges control color in mFruits. Biochemistry 45:9639–9647. https://doi.org/10.1021/bi060773l
Acknowledgements
We thank professor Art Pardi (University of Colorado, Boulder, USA) for contributing to the early stages of this work, prior to his retirement, and Pia Friis (JILA, Boulder, USA) for assistance with sample production. This work was partially supported by the NSF Physics Frontier Center at JILA (PHY 1734006 to R.J.). R.J. is a member of the Quantum Physics Division of the National Institute of Standards and Technology (NIST). Certain commercial equipment, instruments, or materials are identified in this paper in order to specify the experimental procedure adequately. Such identification is not intended to imply recommendation or endorsement by the NIST, nor is it intended to imply that the materials or equipment identified are necessarily the best available for the purpose. Access to the NMR spectrometers at the Danish Center for Ultrahigh-Field NMR Spectroscopy (Ministry of Higher Education and Science grant AU- 2010-612-181) is gratefully acknowledged and partial support to M.S. from the Regione Lazio ‘Contributo premiale per i ricercatori 2022’, DE G05411. We thank Dr. Woonghee Lee (University of Colorado, Denver) for kindly porting CheSPI to python3, and making it accept NMRSTAR 3.x format. the updated CheSPI code is available at https://github.com/protein-nmr.
Funding
Danish Center for Ultrahigh-Field NMR Spectroscopy (Ministry of Higher Education and Science grant AU- 2010-612-181). Regione Lazio ‘Contributo premiale per i ricercatori 2022’, DE G05411. NSF Physics Frontier Center at JILA (PHY 1734006).
Open access funding provided by Johannes Kepler University Linz.
Author information
Authors and Affiliations
Contributions
R.J and F.A.A.M. conceived and supervised the project. R.J. was responsible for molecular biology and isotopically enriched NMR sample generation. F.A.A.M. recorded the NMR data. L.A.J. and M.S. processed NMR spectra and performed resonance assignments. M.S. prepared the BMRB submission. M.S. and F.A.A.M wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
N/A
Consent for publication
N/A
Competing interests
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sette, M., Johnson, L.A., Jimenez, R. et al. Backbone 1H, 15N and 13C resonance assignments of the 27kDa fluorescent protein mCherry. Biomol NMR Assign 17, 243–247 (2023). https://doi.org/10.1007/s12104-023-10149-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-023-10149-z