Abstract
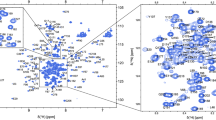
During the maturation of the HIV-1 particle, the Gag polyprotein is cleaved by the viral protease into several proteins: matrix (MA), capsid (CA), spacer peptide 1 (SP1), nucleocapsid (NC), spacer peptide 2 (SP2) and p6. After cleavage, these proteins rearrange to form infectious viral particles. The final cleavage by the protease occurs between CA and SP1 and is the limiting step for the maturation of the particle. The CA–SP1 junction is the target of HIV-1 maturation inhibitors. CA is responsible for the formation of the viral capsid which protects the viral RNA inside. The SP1 domain is essential for viral assembly and infectivity, it is flexible and in helix-coil equilibrium. The presence of NC allows the SP1 domain to be less dynamic. The perturbation of the natural coil-helix equilibrium to helix interferes with protease cleavage and leads to non-completion of viral maturation. In this work, two mutations, W316A and M317A, that abolish the oligomerization of CA were introduced into the protein. The HIV-1 CACTDW316A, M317A-SP1-NC which contains the C-terminal monomeric mutant of CA, SP1 and NC was produced to study the mechanism of action of HIV-1 maturation inhibitors. Here we report the backbone assignment of the protein CACTDW316A, M317A-SP1-NC. These results will be useful to study the interaction between HIV-1 Gag and HIV-1 maturation inhibitors.



Similar content being viewed by others
Data availability
The data generated and analyzed during the current study are available on the BMRB website (http://www.bmrb.wisc.edu) under accession number Entry 50,712.
References
Chen X, Coric P, Larue V et al (2020) The HIV-1 maturation inhibitor, EP39, interferes with the dynamic helix-coil equilibrium of the CA-SP1 junction of Gag. Eur J Med Chem 204:112634. https://doi.org/10.1016/j.ejmech.2020.112634
Coren LV, Thomas JA, Chertova E et al (2007) Mutational analysis of the C-terminal gag cleavage sites in human immunodeficiency virus type 1. J Virol 81:10047–10054. https://doi.org/10.1128/JVI.02496-06
Coric P, Turcaud S, Souquet F et al (2013) Synthesis and biological evaluation of a new derivative of bevirimat that targets the Gag CA-SP1 cleavage site. Eur J Med Chem 62:453–465. https://doi.org/10.1016/j.ejmech.2013.01.013
Deshmukh L, Ghirlando R, Clore GM (2014) Investigation of the structure and dynamics of the capsid-spacer peptide 1–nucleocapsid fragment of the HIV-1 Gag polyprotein by solution NMR spectroscopy. Angew Chem Int Ed 53:1025–1028. https://doi.org/10.1002/anie.201309127
Deshmukh L, Schwieters CD, Grishaev A et al (2013) Structure and dynamics of full length HIV-1 capsid protein in solution. J Am Chem Soc 135:16133–16147. https://doi.org/10.1021/ja406246z
Guzman RND, Wu ZR, Stalling CC et al (1998) Structure of the HIV-1 nucleocapsid protein bound to the SL3 Ψ-RNA recognition element. Science 279:384–388. https://doi.org/10.1126/science.279.5349.384
Lee S-K, Potempa M, Swanstrom R (2012) The choreography of HIV-1 proteolytic processing and virion assembly. J Biol Chem 287:40867–40874. https://doi.org/10.1074/jbc.R112.399444
Lescop E, Schanda P, Brutscher B (2007) A set of BEST triple-resonance experiments for time-optimized protein resonance assignment. J Magn Reson 187:163–169. https://doi.org/10.1016/j.jmr.2007.04.002
Li F, Goila-Gaur R, Salzwedel K et al (2003) PA-457: a potent HIV inhibitor that disrupts core condensation by targeting a late step in Gag processing. PNAS 100:13555–13560. https://doi.org/10.1073/pnas.2234683100
Morellet N, Druillennec S, Lenoir C et al (2005) Helical structure determined by NMR of the HIV-1 (345–392)Gag sequence, surrounding p2: implications for particle assembly and RNA packaging. Protein Sci 14:375–386. https://doi.org/10.1110/ps.041087605
Newman JL, Butcher EW, Patel DT et al (2004) Flexibility in the P2 domain of the HIV-1 Gag polyprotein. Protein Sci 13:2101–2107. https://doi.org/10.1110/ps.04614804
Schanda P, Kupce E, Brutscher B (2005) SOFAST-HMQC experiments for recording two-dimensional heteronuclear correlation spectra of proteins within a few seconds. J Biomol NMR 33:199–211. https://doi.org/10.1007/s10858-005-4425-x
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241. https://doi.org/10.1007/s10858-013-9741-y
Vranken WF, Boucher W, Stevens TJ et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins Struct Funct Bioinf 59:687–696. https://doi.org/10.1002/prot.20449
Wagner JM, Zadrozny KK, Chrustowicz J et al (2016) Crystal structure of an HIV assembly and maturation switch. Elife 5:e17063. https://doi.org/10.7554/eLife.17063
Wang M, Quinn CM, Perilla JR et al (2017) Quenching protein dynamics interferes with HIV capsid maturation. Nat Commun. https://doi.org/10.1038/s41467-017-01856-y
Wiegers K, Rutter G, Kottler H et al (1998) Sequential steps in human immunodeficiency virus particle maturation revealed by alterations of individual Gag polyprotein cleavage sites. J Virol 72:2846–2854
Zargarian L, Tisné C, Barraud P et al (2014) Dynamics of linker residues modulate the nucleic acid binding properties of the HIV-1 nucleocapsid protein zinc fingers. PLoS ONE 9:e102150. https://doi.org/10.1371/journal.pone.0102150
Acknowledgements
We gratefully acknowledge Michael F. Summers for kindly providing us the plasmid of HIV-1 CACTDW316A, M317A-SP1-NC.
Funding
This work was supported by CNRS and the University of Paris. Xiaowei Chen was granted by China Scholarship Council [No 201603250053].
Author information
Authors and Affiliations
Contributions
XC expressed and purified the protein and wrote the manuscript, XC, PC and SB performed the NMR experiments and manuscript revision. XC and PC analyzed the NMR data.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, X., Coric, P. & Bouaziz, S. 1H, 13C and 15N backbone resonance assignment of HIV-1 Gag (276–432) encompassing the C-terminal domain of the capsid protein, the spacer peptide 1 and the nucleocapsid protein. Biomol NMR Assign 15, 267–271 (2021). https://doi.org/10.1007/s12104-021-10016-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-021-10016-9