Abstract
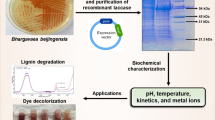
Among lignocellulolytic enzymes, laccases are the most versatile, broadly specific, and largely studied enzyme with a wide range of biotechnological potential. Putative laccase (CotA) from Bacillus pumilus MK001 was cloned and expressed in E. coli. In addition to soluble bioactive fraction, inactive inclusion body fraction was also harvested and refolded under optimized conditions resulting in 64 % of refolding efficiency. The enzyme was found to be thermostable exhibiting a half-life of 60 min at 80 °C. UV thermal CD spectra also supported the observation as about 9 % increase in β-sheets was recorded after thermal induction. The 3D CotA structure was constructed through homology modeling and the best selected model was verified through PROCHECK, ERRAT, Verify 3D, and PROSA servers. Final 3D model showed potential binding affinities with ferulic acid, caffeic acid, and vanillin. Results of the docking studies were further validated by HPLC analysis which signified the efficient bioconversion ability of CotA.






Similar content being viewed by others
References
Sharma, K. K., & Kuhad, R. C. (2008). Laccase: enzyme revisited and function redefined. Indian Journal of Microbiology, 48(3), 309–316. doi:10.1007/s12088-008-0028-z.
Margot, J., Bennati-Granier, C., Maillard, J., Blánquez, P., Barry, D. A., & Holliger, C. (2013). Bacterial versus fungal laccase: Potential for micropollutant degradation. AMB Express, 3(1), 1.
Ferraroni, M., Myasoedova, N. M., Schmatchenko, V., Leontievsky, A. A., Golovleva, L. A., Scozzafava, A., et al. (2007). Crystal structure of a blue laccase from Lentinus tigrinus: Evidences for intermediates in the molecular oxygen reductive splitting by multicopper oxidases. BMC Structural Biology, 7(1), 60. doi:10.1186/1472-6807-7-60.
Reiss, R., Ihssen, J., & Thöny-Meyer, L. (2011). Bacillus pumilus laccase: A heat stable enzyme with a wide substrate spectrum. BMC Biotechnology, 11(1), 1.
Mate, D. M., & Alcalde, M. (2015). Laccase engineering: From rational design to directed evolution. Biotechnology Advances, 33(1), 25–40. doi:10.1016/j.biotechadv.2014.12.007.
Sharma, P., Goel, R., & Capalash, N. (2007). Bacterial laccases. World Journal of Microbiology & Biotechnology, 23(6), 823–832. doi:10.1007/s11274-006-9305-3.
Enguita, F. J., Martins, L. O., Henriques, A. O., & Carrondo, M. A. (2003). Crystal structure of a bacterial endospore coat component: A laccase with enhanced thermostability properties. Journal of Biological Chemistry, 278(21), 19416–19425. doi:10.1074/jbc.M301251200.
Durão, P., Chen, Z., Fernandes, A. T., Hildebrandt, P., Murgida, D. H., Todorovic, S., et al. (2008). Copper incorporation into recombinant CotA laccase from Bacillus subtilis: Characterization of fully copper loaded enzymes. JBIC Journal of Biological Inorganic Chemistry, 13(2), 183–193. doi:10.1007/s00775-007-0312-0.
Kudanga, T., Nyanhongo, G. S., Guebitz, G. M., & Burton, S. (2011). Potential applications of laccase-mediated coupling and grafting reactions: A review. Enzyme and Microbial Technology, 48(3), 195–208. doi:10.1016/j.enzmictec.2010.11.007.
Mathew, S., & Abraham, T. E. (2004). Ferulic acid: An antioxidant found naturally in plant cell walls and feruloyl esterases involved in its release and their applications. Critical Reviews in Biotechnology, 24(2–3), 59–83. doi:10.1080/07388550490491467.
Christopher, L. P., Yao, B., & Ji, Y. (2014). Lignin biodegradation with laccase-mediator systems. Frontiers in Energy Research, 2, 12. doi:10.3389/fenrg.2014.00012.
Ausec, L., van Elsas, J. D., & Mandic-Mulec, I. (2011). Two- and three-domain bacterial laccase-like genes are present in drained peat soils. Soil Biology & Biochemistry, 43(5), 975–983. doi:10.1016/j.soilbio.2011.01.013.
Upadhyay, A. K., Murmu, A., Singh, A., & Panda, A. K. (2012). Kinetics of inclusion body formation and its correlation with the characteristics of protein aggregates in Escherichia coli. PLoS One, 7(3), e33951. doi:10.1371/journal.pone.0033951.
Singh, S. M., & Panda, A. K. (2005). Solubilization and refolding of bacterial inclusion body proteins. Journal of Bioscience and Bioengineering, 99(4), 303–310. doi:10.1263/jbb.99.303.
Kumar, S., Jain, K. K., Bhardwaj, K. N., Chakraborty, S., & Kuhad, R. C. (2015). Multiple Genes in a single host: Cost-effective production of bacterial laccase (cotA), pectate lyase (pel), and endoxylanase (xyl) by simultaneous expression and cloning in single vector in E. coli. PLoS One, 10(12), e0144379. doi:10.1371/journal.pone.0144379.
Kumar, S., Jain, K. K., Singh, A., Panda, A. K., & Kuhad, R. C. (2015). Characterization of recombinant pectate lyase refolded from inclusion bodies generated in E. coli BL21(DE3). Protein Expression and Purification, 110, 43–51. doi:10.1016/j.pep.2014.12.003.
Eswar, N., Webb, B., Marti-Renom, M. A., Madhusudhan, M. S., Eramian, D., Shen, M., et al. (2001). Comparative protein structure modeling using modeller. Current protocols in protein science. New York: Wiley.
Seeliger, D., & Groot, B. L. (2010). Ligand docking and binding site analysis with PyMOL and Autodock/Vina. Journal of Computer-Aided Molecular Design, 24(5), 417–422. doi:10.1007/s10822-010-9352-6.
Irwin, J. J., & Shoichet, B. K. (2005). ZINC—A free database of commercially available compounds for virtual screening. Journal of Chemical Information and Modeling, 45(1), 177–182. doi:10.1021/ci049714+.
Morris, G. M., Huey, R., Lindstrom, W., Sanner, M. F., Belew, R. K., Goodsell, D. S., et al. (2009). AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. Journal of Computational Chemistry, 30(16), 2785–2791. doi:10.1002/jcc.21256.
Dwivedi, U. N., Singh, P., Pandey, V. P., & Kumar, A. (2011). Structure–function relationship among bacterial, fungal and plant laccases. Journal of Molecular Catalysis. B, Enzymatic, 68(2), 117–128. doi:10.1016/j.molcatb.2010.11.002.
Sokalingam, S., Raghunathan, G., Soundrarajan, N., & Lee, S.-G. (2012). A study on the effect of surface lysine to arginine mutagenesis on protein stability and structure using green fluorescent protein. PLoS One, 7(7), e40410. doi:10.1371/journal.pone.0040410.
Guruprasad, K., Reddy, B. V., & Pandit, M. W. (1990). Correlation between stability of a protein and its dipeptide composition: A novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Engineering, 4(2), 155–161.
Nasoohi, N., Khajeh, K., Mohammadian, M., & Ranjbar, B. (2013). Enhancement of catalysis and functional expression of a bacterial laccase by single amino acid replacement. International Journal of Biological Macromolecules, 60, 56–61. doi:10.1016/j.ijbiomac.2013.05.011.
Trubitsina, L. I., Tishchenko, S. V., Gabdulkhakov, A. G., Lisov, A. V., Zakharova, M. V., & Leontievsky, A. A. (2015). Structural and functional characterization of two-domain laccase from Streptomyces viridochromogenes. Biochimie, 112, 151–159. doi:10.1016/j.biochi.2015.03.005.
Mollania, N., Khajeh, K., Ranjbar, B., Rashno, F., Akbari, N., & Fathi-Roudsari, M. (2013). An efficient in vitro refolding of recombinant bacterial laccase in Escherichia coli. Enzyme and Microbial Technology, 52(6–7), 325–330. doi:10.1016/j.enzmictec.2013.03.006.
Arakawa, T., & Tsumoto, K. (2003). The effects of arginine on refolding of aggregated proteins: Not facilitate refolding, but suppress aggregation. Biochemical and Biophysical Research Communications, 304(1), 148–152. doi:10.1016/S0006-291X(03)00578-3.
Yang, J. T., Wu, C. S., & Martinez, H. M. (1986). Calculation of protein conformation from circular dichroism. Methods in Enzymology, 130, 208–269.
Kumar, S., & Nussinov, R. (2001). How do thermophilic proteins deal with heat? Cellular and Molecular Life Sciences CMLS, 58(9), 1216–1233. doi:10.1007/PL00000935.
Mukhopadhyay, A., Dasgupta, A. K., & Chakrabarti, K. (2013). Thermostability, pH stability and dye degrading activity of a bacterial laccase are enhanced in the presence of Cu2O nanoparticles. Bioresource Technology, 127, 25–36. doi:10.1016/j.biortech.2012.09.087.
Zhu, G. P., Xu, C., Teng, M. K., Tao, L. M., Zhu, X. Y., Wu, C. J., et al. (1999). Increasing the thermostability of D-xylose isomerase by introduction of a proline into the turn of a random coil. Protein Engineering, 12(8), 635–638. doi:10.1093/protein/12.8.635.
Ragusa, S., Cambria, M. T., Pierfederici, F., Scirè, A., Bertoli, E., Tanfani, F., et al. (2002). Structure–activity relationship on fungal laccase from Rigidoporus lignosus: A Fourier-transform infrared spectroscopic study. Biochimica et Biophysica Acta (BBA)-Proteins and Proteomics, 1601(2), 155–162.
Mongkolthanaruk, Wiyada. (2012). Independent behavior of bacterial laccases to inducers and metal ions during production and activity. African Journal of Biotechnology, 11(39), 9391–9398. doi:10.5897/AJB11.3042.
Errami, M., Geourjon, C., & Deléage, G. (2003). Detection of unrelated proteins in sequences multiple alignments by using predicted secondary structures. Bioinformatics, 19(4), 506–512. doi:10.1093/bioinformatics/btg016.
Enguita, F. J., Marcal, D., Martins, L. O., Grenha, R., Henriques, A. O., Lindley, P. F., et al. (2004). Substrate and dioxygen binding to the endospore coat laccase from Bacillus subtilis. Journal of Biological Chemistry, 279(22), 23472–23476. doi:10.1074/jbc.M314000200.
Carunchio, F., Crescenzi, C., Girelli, A. M., Messina, A., & Tarola, A. M. (2001). Oxidation of ferulic acid by laccase: Identification of the products and inhibitory effects of some dipeptides. Talanta, 55(1), 189–200.
Adelakun, O. E. (2012). Biocatalytic production of new antioxidant compounds and the characterization of their antioxidant effects. Cape Town: Cape Peninsula University of Technology.
Gutteridge, A., & Thornton, J. M. (2005). Understanding nature’s catalytic toolkit. Trends in Biochemical Sciences, 30(11), 622–629. doi:10.1016/j.tibs.2005.09.006.
Dodson, G., & Wlodawer, A. (1998). Catalytic triads and their relatives. Trends in Biochemical Sciences, 23(9), 347–352. doi:10.1016/S0968-0004(98)01254-7.
Acknowledgments
The financial assistance from the University of Delhi South Campus and DU/DST PURSE Grant to RCK is highly acknowledged. Also the fellowship grants from Indian Council of Medical Research (ICMR), New Delhi to SK and from Council of Scientific and industrial research (CSIR) to KKJ is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, S., Jain, K.K., Rani, S. et al. In-Vitro Refolding and Characterization of Recombinant Laccase (CotA) From Bacillus pumilus MK001 and Its Potential for Phenolics Degradation. Mol Biotechnol 58, 789–800 (2016). https://doi.org/10.1007/s12033-016-9978-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-016-9978-2