Abstract
As the leading central immune organ, the thymus is where T cells differentiate and mature, and plays an essential regulatory role in the adaptive immune response. Tuft cells, as chemosensory cells, were first found in rat tracheal epithelial, later gradually confirmed to exist in various mucosal and non-mucosal tissues. Although tuft cells are epithelial-derived, because of their wide heterogeneity, they show functions similar to cholinergic and immune cells in addition to chemosensory ability. As newly discovered non-mucosal tuft cells, thymic tuft cells have been demonstrated to be involved in and play vital roles in immune responses such as antigen presentation, immune tolerance, and type 2 immunity. In addition to their unique functions in the thymus, thymic tuft cells have the characteristics of peripheral tuft cells, so they may also participate in the process of tumorigenesis and virus infection. Here, we review tuft cells’ characteristics, distribution, and potential functions. More importantly, the potential role of thymic tuft cells in immune response, tumorigenesis, and severe acute respiratory syndrome coronavirus 2(SARS-CoV-2) infection was summarized and discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The thymic microenvironment determines T cells' proliferation, differentiation, and selective development, mainly composed of thymic stromal cells, extracellular matrix, and local active factors. Various cells interact with thymocytes through corresponding mechanisms to form central tolerance in the thymic microenvironment. Tuft cells are one of the newly discovered cell types, widely distributed in the intestines, gallbladder, thymus, and other organs, and play a vital role in immune response, inflammatory response, tumorigenesis, and other diseases [1, 2]. Thymic tuft cells simultaneously have the characteristics of medullary thymic epithelial cells (mTECs) and peripheral tuft cells and are a subset of terminally differentiated mTECs [3]. Thymic tuft cells can promote the development of thymocytes and induce immune tolerance, which makes the thymus without tuft cells unable to form a complete immune tolerance, and some T cells begin to attack their tissues, resulting in autoimmune reactions [4,5,6]. Here, we mainly review the distribution differences and functional heterogeneity of tuft cells discovered in recent years and discuss in detail the regulation of development and the potential functions of tuft cells in the thymus.
Tuft cells
Discovery of tuft cells
As chemosensory cells with unique morphology, tuft cells are widely distributed in various organs and play various important roles [1, 2, 7]. Tuft cells were first found in the epithelium of rats’ tracheal mucosa, and they gained the name because of their microvilli on the top surface of the cells, which were similar to the tuft [8]. With further study of tuft cell distribution, they were found in the epithelium of the salivary gland, gallbladder, and gastrointestinal tract and distributed in a preferential manner [1, 7]. At first, tuft cells were considered to act as chemical signal sensors because they could secrete a variety of physical mediums [1]. People have gradually paid more attention to the immunoregulation and acetylcholine (Ach) synthesis of tuft cells in recent years [2, 9, 10]. Nevertheless, although tuft cells have been discovered for a long time, their distribution and function remain unknown and need to be explored further.
Distribution and markers of tuft cells
Tuft cells express various markers in different microenvironments. Although tuft cells have been confirmed to be widely distributed in many organs and tissues [7], there are significant differences in the distribution frequency of tuft cells in different microenvironments as well as in their markers (Table 1) [2]. In the rat airway where they were first found, the tuft cells accounted for 5–10% of all epithelial cells and expressed markers such as ChAT, TRPM5, and DCLK1 [2, 11, 12], thereby regulating respiratory reflex and airway remodeling [13, 14]. Moreover, Wattel et al. discovered many years ago that tuft cells frequently appeared in the forestomach ridge of rats, accounting for about 20% of the epithelial cells there [7, 15, 16]. Under the conditions of gastritis and mucus cell proliferation, the mRNA of tuft cells DCLK1 was significantly increased, which meant that the number of tuft cells in epithelial tissue increased with inflammation, proliferation, and metaplasia [17]. Huang et al. reported that when functional dyspepsia occurred, the number of tuft cells in duodenal epithelial tissue increased, further providing evidence for this view [18]. As one of the earliest tuft cell markers, villus protein was expressed in tuft cells and intestinal cells, making it not a specific marker for intestinal tuft cells [19]. Ruppert et al. further found that the expression of villin-related protein AVIL was limited to tuft cells, which enabled the intestinal tuft cells to be effectively identified and selected, although their expression accounted for only 0.4% of the local cell population [19,20,21]. Tuft cells can also be present in the urethral epithelium and induce bladder detrusor contraction by expressing Trpm5, expelling harmful substrates [12, 22, 23].
Besides the various distribution in various tissue microenvironments, organism species also cause the different distribution of tuft cells (Table 1). Previous studies have shown that tuft cells were present in the entire respiratory tract in rats, while they were rarely seen in the respiratory tract below the bronchial bifurcation in mice [2]. In addition, the tuft cells in the mice’s stomachs were mainly concentrated in the limiting ridge, while the human gastric tuft cells were widely distributed in the whole stomach, further proving the species difference in the distribution of the tuft cells [9]. Even in the same tissue and organ of the same species, tuft cells were not uniformly distributed. In the study of Chang et al., tuft cells were found in rat airways, accounting for approximately 3% of tracheal epithelium and 1.4% of terminal bronchi, while few tuft cells were seen in lobar bronchi, which meant that there were frequency differences in the distribution of tuft cells along the airways of rats [24].
For the molecular markers, tuft cells also showed differences between humans and mice, most notably DCLK1, whose expression has not been confirmed in human intestinal tuft cells [25, 26]. Miller et al. found that approximately 10% of mTECs in the thymus of adult mice were DCLK1-positive, while only 3.5% of mTECs were DCLK1-positive cells in the human thymus [5]. However, the effect of the difference in tuft cells between these species on their functions is still unknown and deserves to be explored.
Functions of tuft cells
Due to the unique morphology and distribution differences, the tuft cells show diverse effects in different tissues (Table 1). Tuft cells were scattered throughout the gastrointestinal tract and expressed a variety of taste receptors and signaling proteins; therefore, they were thought to be chemoreceptors similar to type II taste cells [1, 7, 26, 46]. Through the expression of taste signaling molecules such as bitter and sweet taste receptors and gustducin, gastrointestinal tuft cells were involved in the perception of sweet, bitter, and umami taste [26, 46]. In addition to sensing taste, the tuft cells can sense harmful stimuli, inflammation, and stimulation of harmful bacteria and parasites and express various signaling molecules to regulate the altered microenvironment [47]. Previous studies have found that tuft cells can respond to the destruction of intestinal microbiota caused by the influenza virus, which proved the sensory effect of tuft cells on harmful stimuli [32]. At the same time, tuft cells were necessary for the increase of type 2 innate lymphocytes (ILC2s) in the small intestine induced by the influenza virus, which suggested the regulator role of tuft cells in the immune response [32]. Xiong et al. also reported that intestinal tuft cells could sense bacterial metabolites and exert antibacterial immunity, further proving that the tuft cells had not only chemosensory effects but also mediated immune effects [48]. Moreover, Ach biosynthesis by tuft cells was another important effect in addition to immunologic and chemosensory functions. Ach derived from the tuft cells played vital roles in regulating airway remodeling, promoting muscle contraction, maintaining epithelial homeostasis, and promoting the occurrence of cancer [2].
Even in different parts of the same tissue, the function of tuft cells is varied (Table 1). In every anatomical position, tuft cells showed a high degree of specificity and adaptability to adjust their response to each microenvironment change. Haber et al. found two subtypes when observing intestinal tuft cells [34, 49]. Type 1 tuft cells signal contained genes related to neural development, while type 2 tuft cells were rich in immune-related genes. However, little is known about the number of regulated tuft cells in the process of different microenvironment alterations and how these sub-populations of tuft cells work, which still need to be further explored.
Tuft cells in the thymus
Thymus is a central immune organ responsible for producing various but self-tolerant T lymphocyte banks, which are essential for the development and maturity of T cells [50]. mTECs help shape the thymic microenvironment to promote the establishment of central self-tolerance by expressing multiple peripheral tissue-restricted antigens (TRAs) that bind to self-reactive T cells and induce negative selection of highly self-reactive T cells [51, 52]. To increase the diversity of TRAs, a portion of mTECs can “terminally differentiate” into atypical cells similar to peripheral epithelial cells [51]. Thymic tuft cells are one of the subsets of mTECs after terminal differentiation [5, 43]. Miller et al. confirmed the existence of tuft cells in the thymus through flow cytometry analysis [5].
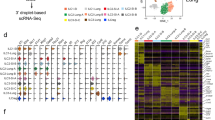
Moreover, thymic tuft cells were found to share a large number of regulatory factors and specific genes with intestinal tuft cells [43]. This reflected that thymic tuft cells might have similar characteristics to peripheral tuft cells. However, thymic tuft cells also have their specific gene expression. When Nevo et al. explored the differences between thymic tuft cells and peripheral tuft cells, they found that thymic tuft cells specifically expressed L1CAM and MHC II (Fig. 1a) [1]. In addition to the differences in gene expression, thymic tuft cells exist in tissue epithelium rather than mucosal tissue. Their morphological characteristics do not have clear top and bottom like intestinal tuft cells, but show lateral microvilli [33]. Therefore, understanding the development, heterogeneity, and potential functions of tuft cells in the thymus will help us to better understand the role of the thymus.
a The difference between thymic tuft cells and other tuft cells. b Insufficient sox4 level cannot maintain the normal development of thymic tuft cells. c Co-expression of Pou2f3 and OCA-T1 activates Trpm5, thus promoting the development of thymic tuft cells. d The deletion of Sirt6 reduces the accessibility of the Pou2f3 chromosome, thus hindering the development of thymic tuft cells. e The expression of IL-25 in thymic tuft cells promotes the expression of IL-4 in NKT2 cells and causes DC to express chemokines, thus affecting the development of thymocytes
Development and expression regulation of thymic tuft cells
Sox4 regulates the differentiation of stem cells in various tissues and is also significant for the development of various lymphocytes [53,54,55,56]. Thymic tuft cells are a subset of recently discovered terminally differentiated mTECs for which SRY-related high-mobility-group box 4 (Sox4) is essential for their development. Previous studies have proved that a certain abundance of Sox4 can maintain the normal development of thymic tuft cells. By gene expression analysis, Mino et al. assessed that in mTECs of mice lacking of Sox4, the mRNA expression level of Dclk1 and Pou2f3 decreased, which were the tuft cell-related genes and the tuft cell markers [51]. When the level of Sox4 in thymic epithelial cells decreased, the thymus only showed the normal development of conventional mTECs, while the number of thymic tuft cells decreased (Fig. 1b) [51]. This indicated that Sox4 might influence the terminal differentiation of mTECs into thymic tuft cells, thus affecting the development of thymic tuft cells.
In addition, taste signal molecules Pou2f3 and Trpm5 were also considered to be necessary for the development and function of thymic tuft cells. In the study of Miller et al., Pou2f3−/− mice and Trpm5−/− mice showed specific loss of thymic tuft cells [5, 43]. In normal thymic tuft cells, the co-expression of Pou2f3 and uncharacterized proteins C11orf53 (OCA-T1) activated the tuft cell-specific gene Trpm5, which further promoted the development of thymic tuft cells (Fig. 1c) [57]. The development and differentiation of thymic tuft cells also depended on the epigenetic regulatory factor Sirtuin 6 (Sirt6). The deletion of Sirt6 reduced the expression of the Pou2f3-regulated gene and chromatin accessibility in thymic tuft cells (Fig. 1d), thereby inhibiting Pou2f3-mediated differentiation of thymic tuft cells and finally significantly reducing the presence of thymic tuft cells [45]. Therefore, it suggested that nuclear transcription factors and epigenetic regulatory factors are crucial for the development and differentiation of thymic tuft cells, and the taste signal transduction pathway plays a significant role in maintaining the function of thymic tuft cells.
Effect on thymus development of tuft cells
Thymic tuft cells played an essential role in shaping T lymphocytes development [5, 42]. Similar to mucosal tuft cells, thymic tuft cells expressed high levels of IL-25, and IL-25 expression was limited to thymic tuft cells in the thymus [3, 42]. This meant that NKT2 cells expressing IL-25 receptor (IL-25 R) in the thymus might be regulated by thymic tuft cells. Lacus et al. discovered that the number of NKT2 cells was selectively decreased in the thymus of tuft-cell-deficient mice, and NKT2 cells were recovered after in vivo administration of IL-25, further proving that thymic tuft cells were a critical selective regulator of NKT2 [42]. NKT2 cells were confirmed to be the primary source of IL-4 in the thymus [6]. Through the stable production of IL-4, selectively activated NKT2 cells can regulate the expression of chemokines by thymic dendritic cells (DCs) and regulate CD8 + cells to become memory phenotypes, which strongly affects the development of thymocytes (Fig. 1e) [6, 42]. However, the stable production of IL-4 depends not only on the production of IL-25 by thymic tuft cells but also on the continuous production of T cell receptors (TCRs) in the thymic medulla [42]. The production of IL-25 by the thymic tuft cells affected the development of NKT2 cells, which were then stimulated by the TCR signal in the thymus medulla to differentiate, produce and maintain NKT2 function [6].
Previous studies have displayed that tuft cells in mice thymus medulla possessed the characteristics of cholinergic epithelial cells and expressed ChAT, which meant that thymic tuft cells owned the possibility of synthesizing Ach [33]. Ach is expressed by thymic tuft cells bound to acetylcholine receptors in the thymus, which made DCs polarize towards T helper 2 (Th2) promoting mode [58, 59]. Moreover, Ach may also play roles in regulating the differentiation, maturation, and selection of T cells, affecting the development of T cells and playing a pivotal role in maintaining the normal function of the thymus [1, 60].
Thymic tuft cells participating in immune response
Using the typical taste chemical sensory pathway, tuft cells can detect the potentially harmful compounds on the mucosal surface and further prevent the invasion of bacteria or harmful compounds by starting a protective reaction [47]. Unlike mucosal tuft cells, tuft cells located in the thymic medulla were not often directly exposed to foreign bacteria and harmful substances [33]. However, previous studies have indicated that during systemic infection, viruses, bacteria, and bacterial products can still reach the thymus, and the thymus can also sense and respond to these harmful stimuli [1, 33, 61], which means that thymic tuft cells might rely on their taste signal transduction pathway to feel the harmful stimuli reaching the thymus and initiate the corresponding protective reactions.
Although thymic tuft cells and mucosal tuft cells share a large number of regulatory factors and tuft-specific genes and possess many similarities, thymic tuft cells also express a large number of different genes and display unique functions [3, 43, 62]. Unlike intestinal tuft cells, which expressed MHC I and β2-microglobulin, thymic tuft cells were revealed to express Tas2r site at a relatively high level, as well as several members of MHC II and CD74 genes related to antigen presentation (Fig. 2) [3, 5, 10]. This specificity of thymic tuft cells increased their potential to present antigens to developing thymocytes.
The immune function of tuft cells was well reflected in the intestine. Similar to small intestinal tuft cells, thymic tuft cells also expressed IL-25 [42]. As IL-25 seems to be the key factor connecting tuft cell chemoperception with type 2 immunity [63], it can be speculated that thymic tuft cells may also participate in type 2 immunity. However, although a large number of thymic tuft cells expressed IL-25 to activate NKT2 (Fig. 2), and the number of NKT2 cells and EOMES + CD8 + single positive thymocytes decreased in thymus lacking tuft cells, which could provide evidence for thymic tuft cells to participate in type 2 immunity [5], Nevo et al. showed that the frequency and number of thymic innate lymphocyte cells type 2 (ILC2) in mice with tuft cells deficiency increased due to feedback loop or intestinal-thymus crosstalk [1]. Therefore, whether thymic tuft cells are involved in the activation of type 2 immune response needs further exploration.
In addition to possibly being associated with the immune response process, thymic tuft cells also played a certain role in inducing immune tolerance. It was found that the athymic mice transplanted with Pou2f3−/− thymus (with peripheral tuft cells but lacking thymic tuft cell antigen) produced a specific antibody response to IL-25 immune response, while the wild-type mice were tolerant [5, 63]. Therefore, we can speculate that thymic tuft cells seem to be involved in enhancing the tolerance of unique tuft-restricted antigens including IL-25.
Thymic tuft cells and thymic tumors
Tuft cells are thought to promote the occurrence of cancer because of their ability to regulate the microenvironment of the nerve around the tumor and their ability to act as progenitor cells leading to cancer [2, 17]. Recently, Yamada et al. discovered by immunohistochemistry that KIT and other known genes of thymic tuft cells were co-expressed in normal human thymus tuft cells and most thymic carcinoma, and even genes specific to thymic tuft cells, such as L1CAM, were expressed in thymic squamous cell carcinoma (Fig. 3) [64]. This suggests that thymic tuft cells may be involved in the occurrence and development of thymic tumors.
SRY-box transcription factor 9 (Sox9) plays a vital role in the occurrence and development of cancer because it is involved in the occurrence, transfer, and change of the tumor microenvironment [65]. When Yuan et al. explored the diagnostic value of Sox9 in thymic epithelial tumors (TETs), they found that Sox9 was highly expressed in the nucleus of TETs, and its high expression meant a lousy outcome for thymic tumors [66]. This reflects the possible role of Sox9 as a diagnostic and prognostic indicator for TETs. Notably, Sox9 could be used as a marker of tuft cells in some tissues, and its expression was also positively correlated with the expression of Pou2f3, an essential regulator of tuft cells [1, 66]. This further suggests that thymic tumors display a tuft cell-related phenotype.
However, as a thymic tumor, thymomas exhibit a much less tuft cell-like phenotype than thymic carcinoma [67]. This may be because many mTEC terminal differentiation genes are not expressed in thymomas, resulting in the final deletion of the developmental pathway of thymic tuft cells due to the non-expression of Pou2f3 in thymomas [64, 68]. However, thymic carcinoma expresses thymic tuft cells’ specific genes, which means that tuft cell-like thymic squamous cell carcinoma may be produced by mutated mTEC precursors [64]. We realize that tuft cell-like carcinoma has the characteristics of tuft cell gene expression, which may provide vital information for a more accurate diagnosis of thymic carcinoma. However, the relationship between thymic tuft cells and thymic tumors remains unclear and needs further exploration.
The potential association between thymic tuft cells and SARS-CoV-2 infection
Previous studies have found that tuft cells could directly cause diseases as the target cells of virus infection; on the other hand, they can indirectly affect the occurrence of diseases through the immune response driven by co-infection [25]. It has been demonstrated that murine rotavirus can directly infect and replicate within tuft cells [69]. Coincidentally, Wilen et al. revealed that intestinal tuft cells expressed MNV virus receptor CD300lf, and it was the main physiological target of the MNVCR6 strain [70]. Therefore, the proliferation of intestinal tuft cells can promote the infection of MNVCR6, which also reflects that the tuft cells have the potential to promote viral infection. In addition, the virus and tuft cell interacted with each other. While tuft cells promoted viral infection, viruses can also facilitate the development of nascent tuft cells [25].
During the SARS-CoV-2 global pandemic, the number of tuft cells in the upper respiratory tract of coronavirus-19 disease (COVID-19) patients increased by three folds, and ectopic development of the tuft cells occurred in the lung, suggesting that the tuft cells may also be involved in the SARS-CoV-2 infection process [71]. As a part of the central immune organs which participates in the immune process, it is not clear whether thymic tuft cells are involved in the pathogenesis of SARS-CoV-2. However, the existing studies have confirmed that the CD74 gene was significantly upregulated, and the antigen presentation function was enhanced in asymptomatic SARS-CoV-2 infected patients [72]. Given the specific expression of MHC II and CD74 in thymic tuft cells, compared to peripheral tuft cells, it is reasonable to suspect that thymic tuft cells may be involved in the process of infection with SARS-CoV-2. Previous studies have confirmed that SARS-CoV-2 required not only the combination of the S1 subunit of virus spike (S) protein and angiotensin-converting enzyme 2 (ACE2) of the host cell to locate the virus, but also the transmembrane serine protease 2 (TMPRSS2) of the host cell to cut the virus S protein and activate S2 subunit, thus promoting the virus to enter the host cell [73, 74]. In Bruchez et al.’s research, the MHC class II trans-activator (CIITA) activated the expression of the P41 subtype of CD74, and CD74 P41 inhibited the processing of S protein by TMPRSS2 (Fig. 4) [75]. This means that the specific high expression of the MHC II factor and CD74 gene in thymic tuft cells can not only play the role of antigen presentation but also prevent SARS-CoV-2 from infecting host cells. Considering the decrease in lymphocytes in patients infected with SARS-CoV-2 [76], we speculate that, unlike peripheral tuft cells that promote the onset of the virus, thymic tuft cells may play a role in the anti-SARS-CoV-2 response and SARS-CoV-2 may also damage thymic tuft cells in turn during this process to cause immunodeficiency. However, there have been no reports of SARS-CoV-2 infection in thymic tuft cells, which needs further exploration.
Conclusion and prospect
Although tuft cells can be distributed in different organs or tissues to produce a variety of effector molecules and play a variety of different effects, it is due to the differential distribution of tuft cells in organisms between diseases and health states, and among species, the known function of tuft cells varied. Collectively, thymic tuft cells express taste receptors, ChAT, IL-25, CD74, and MHC II, suggesting that they may have the ability of chemosensory, cholinergic, and immune response. However, thymic tuft cells’ role in thymic tumorigenesis, virus infection, and other diseases is still complex, and whether the thymic tuft cells own other thymic-specific effects needs further exploration. The heterogeneous expression of bitter receptors in thymic tuft cells makes it a significant research direction to assess whether the pool of chemosensory receptors is a feature that drives the functional heterogeneity of thymic tuft cells. Not only that, but the discovery of thymic tuft cells also provides a potential target for medical intervention strategies for immune aging and other related health problems and will provide the possibility for the diagnosis and treatment of autoimmune diseases and some tumors.
Data availability
Data and material are available upon request.
Code availability
Not applicable.
References
Nevo S, Kadouri N, Abramson J. Tuft cells: from the mucosa to the thymus. Immunol Lett. 2019;210:1–9. https://doi.org/10.1016/j.imlet.2019.02.003.
Pan J, Zhang L, Shao X, Huang J. Acetylcholine from tuft cells: the updated insights beyond its immune and chemosensory functions. Front Cell Develop Biol. 2020;8:606. https://doi.org/10.3389/fcell.2020.00606.
Kadouri N, Nevo S, Goldfarb Y, Abramson J. Thymic epithelial cell heterogeneity: TEC by TEC. Nat Rev Immunol. 2020;20:239–53. https://doi.org/10.1038/s41577-019-0238-0.
Marx A, Yamada Y, Simon-Keller K, Schalke B, Willcox N, Ströbel P, Weis C. Thymus and autoimmunity. Semin Immunopathol. 2021;43:45–64. https://doi.org/10.1007/s00281-021-00842-3.
Miller C, Proekt I, von Moltke J, Wells K, Rajpurkar A, Wang H, Rattay K, Khan I, Metzger T, Pollack J, Fries A, Lwin W, Wigton E, Parent A, Kyewski B, Erle D, Hogquist K, Steinmetz L, Locksley R, Anderson M. Thymic tuft cells promote an IL-4-enriched medulla and shape thymocyte development. Nature. 2018;559:627–31. https://doi.org/10.1038/s41586-018-0345-2.
Wang H, Breed E, Lee Y, Qian L, Jameson S, Hogquist K. Myeloid cells activate iNKT cells to produce IL-4 in the thymic medulla. Proc Natl Acad Sci U S A. 2019;116:22262–8. https://doi.org/10.1073/pnas.1910412116.
Sato A. Tuft cells. Anatomical Sci Int. 2007;82:187–99. https://doi.org/10.1111/j.1447-073X.2007.00188.x.
Rhodin J, Dalhamn T. Electron microscopy of the tracheal ciliated mucosa in rat. Z Zellforsch Mikrosk Anat. 1956;44:345–412. https://doi.org/10.1007/bf00345847.
Schneider C, O’Leary C, Locksley R. Regulation of immune responses by tuft cells. Nat Rev Immunol. 2019;19:584–93. https://doi.org/10.1038/s41577-019-0176-x.
Ting H, von Moltke J. The immune function of tuft cells at gut mucosal surfaces and beyond. J Immunol (Baltimore, Md:1950). 2019;202:1321–9. https://doi.org/10.4049/jimmunol.1801069.
Meyrick B, Reid L. The alveolar brush cell in rat lung–a third pneumonocyte. J Ultrastruct Res. 1968;23:71–80. https://doi.org/10.1016/s0022-5320(68)80032-2.
O’Leary C, Schneider C, Locksley R. Tuft cells-systemically dispersed sensory epithelia integrating immune and neural circuitry. Annu Rev Immunol. 2019;37:47–72. https://doi.org/10.1146/annurev-immunol-042718-041505.
Rane C, Jackson S, Pastore C, Zhao G, Weiner A, Patel N, Herbert D, Cohen N, Vaughan A. Development of solitary chemosensory cells in the distal lung after severe influenza injury. American journal of physiology. Lung Cell Mole Physiol. 2019;316:L1141–9. https://doi.org/10.1152/ajplung.00032.2019.
Krasteva G, Canning B, Hartmann P, Veres T, Papadakis T, Mühlfeld C, Schliecker K, Tallini Y, Braun A, Hackstein H, Baal N, Weihe E, Schütz B, Kotlikoff M, Ibanez-Tallon I, Kummer W. Cholinergic chemosensory cells in the trachea regulate breathing. Proc Natl Acad Sci U S A. 2011;108:9478–83. https://doi.org/10.1073/pnas.1019418108.
Luciano L, Reale E. The “limiting ridge” of the rat stomach. Arch Histol Cytol. 1992;131-8https://doi.org/10.1679/aohc.55.suppl_131
Wattel W, Geuze J. The cells of the rat gastric groove and cardia. An ultrastructural and carbohydrate histochemical study, with special reference to the fibrillovesicular cells. Cell Tissue Res. 1978;186:375–91. https://doi.org/10.1007/bf00224928.
Saqui-Salces M, Keeley T, Grosse A, Qiao X, El-Zaatari M, Gumucio D, Samuelson L, Merchant J. Gastric tuft cells express DCLK1 and are expanded in hyperplasia. Histochem Cell Biol. 2011;136:191–204. https://doi.org/10.1007/s00418-011-0831-1.
Huang X, Oshima T, Akiba Y, Yoshimoto T, Chen J, Taki M, Tomita T, Fukui H, Kaunitz J, Miwa H. Duodenal cholinergic tuft cell number is increased in functional dyspepsia. Neurogastroenterol Motility: Off J Eur Gastrointestinal Motility Soc. 2022;e14378. https://doi.org/10.1111/nmo.14378
Ruppert A, Keshavarz M, Winterberg S, Oberwinkler J, Kummer W, Schütz B. Advillin is a tuft cell marker in the mouse alimentary tract. J Mol Histol. 2020;51:421–35. https://doi.org/10.1007/s10735-020-09893-6.
Gerbe F, van Es J, Makrini L, Brulin B, Mellitzer G, Robine S, Romagnolo B, Shroyer N, Bourgaux J, Pignodel C, Clevers H, Jay P. Distinct ATOH1 and Neurog3 requirements define tuft cells as a new secretory cell type in the intestinal epithelium. J Cell Biol. 2011;192:767–80. https://doi.org/10.1083/jcb.201010127.
Schütz B, Ruppert A, Strobel O, Lazarus M, Urade Y, Büchler M, Weihe E. Distribution pattern and molecular signature of cholinergic tuft cells in human gastro-intestinal and pancreatic-biliary tract. Sci Rep. 2019;9:17466. https://doi.org/10.1038/s41598-019-53997-3.
Yamashita J, Ohmoto M, Yamaguchi T, Matsumoto I, Hirota J. Skn-1a/Pou2f3 functions as a master regulator to generate Trpm5-expressing chemosensory cells in mice. PLoS One. 2017;12:e0189340. https://doi.org/10.1371/journal.pone.0189340.
Deckmann K, Kummer W. Chemosensory epithelial cells in the urethra: sentinels of the urinary tract. Histochem Cell Biol. 2016;146:673–83. https://doi.org/10.1007/s00418-016-1504-x.
Chang L, Mercer R, Crapo J. Differential distribution of brush cells in the rat lung. Anat Rec. 1986;216:49–54. https://doi.org/10.1002/ar.1092160109.
Strine M, Wilen C. Tuft cells are key mediators of interkingdom interactions at mucosal barrier surfaces. PLoS Pathog. 2022;18:e1010318. https://doi.org/10.1371/journal.ppat.1010318.
Burman A, Kaji I. Luminal chemosensory cells in the small intestine. Nutrients. 2021;13. https://doi.org/10.3390/nu13113712
Howitt M, Lavoie S, Michaud M, Blum A, Tran S, Weinstock J, Gallini C, Redding K, Margolskee R, Osborne L, Artis D, Garrett W. Tuft cells, taste-chemosensory cells, orchestrate parasite type 2 immunity in the gut. Science (New York NY). 2016;351:1329–33. https://doi.org/10.1126/science.aaf1648.
Kjærgaard S, Jensen T, Feddersen U, Bindslev N, Grunddal K, Poulsen S, Rasmussen H, Budtz-Jørgensen E, Berner-Hansen M. Decreased number of colonic tuft cells in quiescent ulcerative colitis patients. Eur J Gastroenterol Hepatol. 2021;33:817–24. https://doi.org/10.1097/meg.0000000000001959.
Hendel S, Kellermann L, Hausmann A, Bindslev N, Jensen K, Nielsen O. Tuft cells and their role in intestinal diseases. Front Immunol. 2022;13:822867. https://doi.org/10.3389/fimmu.2022.822867.
Gerbe F, Jay P. Intestinal tuft cells: epithelial sentinels linking luminal cues to the immune system. Mucosal Immunol. 2016;9:1353–9. https://doi.org/10.1038/mi.2016.68.
McGinty J, Ting H, Billipp T, Nadjsombati M, Khan D, Barrett N, Liang H, Matsumoto I, von Moltke J. Tuft-cell-derived leukotrienes drive rapid anti-helminth immunity in the small intestine but are dispensable for anti-protist immunity. Immunity. 2020;52:528-541.e7. https://doi.org/10.1016/j.immuni.2020.02.005.
Roach S, Fiege J, Shepherd F, Wiggen T, Hunter R, Langlois R. Respiratory influenza virus infection causes dynamic tuft cell and innate lymphoid cell changes in the small intestine. J Virol. 2022;96:e0035222. https://doi.org/10.1128/jvi.00352-22.
Panneck A, Rafiq A, Schütz B, Soultanova A, Deckmann K, Chubanov V, Gudermann T, Weihe E, Krasteva-Christ G, Grau V, del Rey A, Kummer W. Cholinergic epithelial cell with chemosensory traits in murine thymic medulla. Cell Tissue Res. 2014;358:737–48. https://doi.org/10.1007/s00441-014-2002-x.
Haber A, Biton M, Rogel N, Herbst R, Shekhar K, Smillie C, Burgin G, Delorey T, Howitt M, Katz Y, Tirosh I, Beyaz S, Dionne D, Zhang M, Raychowdhury R, Garrett W, Rozenblatt-Rosen O, Shi H, Yilmaz O, Xavier R, Regev A. A single-cell survey of the small intestinal epithelium. Nature. 2017;551:333–9. https://doi.org/10.1038/nature24489.
Manco R, Averbukh I, Porat Z, Bahar Halpern K, Amit I, Itzkovitz S. Clump sequencing exposes the spatial expression programs of intestinal secretory cells. Nat Commun. 2021;12:3074. https://doi.org/10.1038/s41467-021-23245-2.
Ualiyeva S, Yoshimoto E, Barrett N, Bankova L. Isolation and quantitative evaluation of brush cells from mouse tracheas. Journal of visualized experiments: JoVE. 2019. https://doi.org/10.3791/59496.
Kotas M, Moore C, Gurrola J, Pletcher S, Goldberg A, Alvarez R, Yamato S, Bratcher P, Shaughnessy C, Zeitlin P, Zhang I, Li Y, Montgomery M, Lee K, Cope E, Locksley R, Seibold M, Gordon E. IL-13-programmed airway tuft cells produce PGE2, which promotes CFTR-dependent mucociliary function. JCI insight. 2022;7. https://doi.org/10.1172/jci.insight.159832
Ualiyeva S, Hallen N, Kanaoka Y, Ledderose C, Matsumoto I, Junger W, Barrett N, Bankova L. Airway brush cells generate cysteinyl leukotrienes through the ATP sensor P2Y2. Science immunology. 2020;5. https://doi.org/10.1126/sciimmunol.aax7224
Bankova L, Dwyer D, Yoshimoto E, Ualiyeva S, McGinty J, Raff H, von Moltke J, Kanaoka Y, Frank Austen K, Barrett N. The cysteinyl leukotriene 3 receptor regulates expansion of IL-25-producing airway brush cells leading to type 2 inflammation. Science immunology. 2018;3. https://doi.org/10.1126/sciimmunol.aat9453
Ualiyeva S, Lemire E, Aviles E, Wong C, Boyd A, Lai J, Liu T, Matsumoto I, Barrett N, Boyce J, Haber A, Bankova L. Tuft cell-produced cysteinyl leukotrienes and IL-25 synergistically initiate lung type 2 inflammation. Science immunology. 2021;6:eabj0474. https://doi.org/10.1126/sciimmunol.abj0474.
Montoro D, Haber A, Biton M, Vinarsky V, Lin B, Birket S, Yuan F, Chen S, Leung H, Villoria J, Rogel N, Burgin G, Tsankov A, Waghray A, Slyper M, Waldman J, Nguyen L, Dionne D, Rozenblatt-Rosen O, Tata P, Mou H, Shivaraju M, Bihler H, Mense M, Tearney G, Rowe S, Engelhardt J, Regev A, Rajagopal J. A revised airway epithelial hierarchy includes CFTR-expressing ionocytes. Nature. 2018;560:319–24. https://doi.org/10.1038/s41586-018-0393-7.
Lucas B, White A, Cosway E, Parnell S, James K, Jones N, Ohigashi I, Takahama Y, Jenkinson W, Anderson G. Diversity in medullary thymic epithelial cells controls the activity and availability of iNKT cells. Nat Commun. 2020;11:2198. https://doi.org/10.1038/s41467-020-16041-x.
Bornstein C, Nevo S, Giladi A, Kadouri N, Pouzolles M, Gerbe F, David E, Machado A, Chuprin A, Tóth B, Goldberg O, Itzkovitz S, Taylor N, Jay P, Zimmermann V, Abramson J, Amit I. Single-cell mapping of the thymic stroma identifies IL-25-producing tuft epithelial cells. Nature. 2018;559:622–6. https://doi.org/10.1038/s41586-018-0346-1.
Liu S, Lu S, Xu R, Atzberger A, Günther S, Wettschureck N, Offermanns S. Members of bitter taste receptor cluster Tas2r143/Tas2r135/Tas2r126 Are expressed in the epithelium of murine airways and other non-gustatory tissues. Front Physiol. 2017;8:849. https://doi.org/10.3389/fphys.2017.00849.
Zhang Q, Zhang J, Lei T, Liang Z, Dong X, Sun L, Zhao Y. Sirt6-mediated epigenetic modification of DNA accessibility is essential for Pou2f3-induced thymic tuft cell development. Communications biology. 2022;5:544. https://doi.org/10.1038/s42003-022-03484-9.
Akiba Y, Hashimoto S, Kaunitz J. Duodenal chemosensory system: enterocytes, enteroendocrine cells, and tuft cells. Curr Opin Gastroenterol. 2020;36:501–8. https://doi.org/10.1097/mog.0000000000000685.
O’Leary C, Ma Z, Culpepper T, Novak S, DelGiorno K. New insights into tuft cell formation: Implications for structure-function relationships. Curr Opin Cell Biol. 2022;76:102082. https://doi.org/10.1016/j.ceb.2022.102082.
Xiong Z, Zhu X, Geng J, Xu Y, Wu R, Li C, Fan D, Qin X, Du Y, Tian Y, Fan Z. Intestinal Tuft-2 cells exert antimicrobial immunity via sensing bacterial metabolite N-undecanoylglycine. Immunity. 2022;55:686-700.e7. https://doi.org/10.1016/j.immuni.2022.03.001.
Cheng X, Voss U, Ekblad E. Tuft cells: Distribution and connections with nerves and endocrine cells in mouse intestine. Exp Cell Res. 2018;369:105–11. https://doi.org/10.1016/j.yexcr.2018.05.011.
Thapa P, Farber D. The role of the thymus in the immune response. Thorac Surg Clin. 2019;29:123–31. https://doi.org/10.1016/j.thorsurg.2018.12.001.
Mino N, Muro R, Ota A, Nitta S, Lefebvre V, Nitta T, Fujio K, Takayanagi H. The transcription factor Sox4 is required for thymic tuft cell development. Int Immunol. 2022;34:45–52. https://doi.org/10.1093/intimm/dxab098.
Wang H, Pan W, Zheng L, Zhong X, Tan L, Liang Z, He J, Feng P, Zhao Y, Qiu Y. Thymic epithelial cells contribute to thymopoiesis and t cell development. Front Immunol. 2019;10:3099. https://doi.org/10.3389/fimmu.2019.03099.
Foronda M, Martínez P, Schoeftner S, Gómez-López G, Schneider R, Flores J, Pisano D, Blasco M. Sox4 links tumor suppression to accelerated aging in mice by modulating stem cell activation. Cell Rep. 2014;8:487–500. https://doi.org/10.1016/j.celrep.2014.06.031.
van de Wetering M, Oosterwegel M, van Norren K, Clevers H. Sox-4, an Sry-like HMG box protein, is a transcriptional activator in lymphocytes. EMBO J. 1993;12:3847–54. https://doi.org/10.1002/j.1460-2075.1993.tb06063.x.
Malhotra N, Qi Y, Spidale N, Frascoli M, Miu B, Cho O, Sylvia K, Kang J. SOX4 controls invariant NKT cell differentiation by tuning TCR signaling. J Exp Med. 2018;215:2887–900. https://doi.org/10.1084/jem.20172021.
Schilham M, Moerer P, Cumano A, Clevers H. Sox-4 facilitates thymocyte differentiation. Eur J Immunol. 1997;27:1292–5. https://doi.org/10.1002/eji.1830270534.
Wu XS, He XY, Ipsaro JJ, Huang YH, Preall JB, Ng D, Shue YT, Sage J, Egeblad M, Joshua-Tor L, Vakoc CR. OCA-T1 and OCA-T2 are coactivators of POU2F3 in the tuft cell lineage. Nature. 2022;607:169–75. https://doi.org/10.1038/s41586-022-04842-7.
Gori S, Vermeulen M, Remes-Lenicov F, Jancic C, Scordo W, Ceballos A, Towstyka N, Bestach Y, Belli C, Sabbione F, Geffner J, Salamone G. Acetylcholine polarizes dendritic cells toward a Th2-promoting profile. Allergy. 2017;72:221–31. https://doi.org/10.1111/all.12926.
Raimond F, Morel E, Bach J. Evidence for the presence of immunoreactive acetylcholine receptors on human thymus cells. J Neuroimmunol. 1984;6:31–40. https://doi.org/10.1016/0165-5728(84)90040-7.
Kawashima K, Fujii T. Expression of non-neuronal acetylcholine in lymphocytes and its contribution to the regulation of immune function. Front Biosci. 2004;9:2063–85. https://doi.org/10.2741/1390.
Dooley J, Liston A. Molecular control over thymic involution: from cytokines and microRNA to aging and adipose tissue. Eur J Immunol. 2012;42:1073–9. https://doi.org/10.1002/eji.201142305.
Nadjsombati M, McGinty J, Lyons-Cohen M, Jaffe J, DiPeso L, Schneider C, Miller C, Pollack J, Nagana Gowda G, Fontana M, Erle D, Anderson M, Locksley R, Raftery D, von Moltke J. Detection of succinate by intestinal tuft cells triggers a type 2 innate immune circuit. Immunity. 2018;49:33-41.e7. https://doi.org/10.1016/j.immuni.2018.06.016.
Borowczyk J, Shutova M, Brembilla N, Boehncke W. IL-25 (IL-17E) in epithelial immunology and pathophysiology. J Allergy Clin Immunol. 2021;148:40–52. https://doi.org/10.1016/j.jaci.2020.12.628.
Yamada Y, Simon-Keller K, Belharazem-Vitacolonnna D, Bohnenberger H, Kriegsmann M, Kriegsmann K, Hamilton G, Graeter T, Preissler G, Ott G, Roessner E, Dahmen I, Thomas R, Ströbel P, Marx A. A Tuft cell-like signature is highly prevalent in thymic squamous cell carcinoma and delineates new molecular subsets among the major lung cancer histotypes. J Thoracic Oncol: Off Publ Int Assoc Study Lung Cancer. 2021;16:1003–16. https://doi.org/10.1016/j.jtho.2021.02.008.
Grimm D, Bauer J, Wise P, Krüger M, Simonsen U, Wehland M, Infanger M, Corydon TJ. The role of SOX family members in solid tumours and metastasis. Semin Cancer Biol. 2020;67:122–53. https://doi.org/10.1016/j.semcancer.2019.03.004.
Yuan X, Huang L, Luo W, Zhao Y, Nashan B, Yu F, Liu Y. Diagnostic and prognostic significances of SOX9 in thymic epithelial tumor. Front Oncol. 2021;11:708735. https://doi.org/10.3389/fonc.2021.708735.
Yamada Y, Sugimoto A, Hoki M, Yoshizawa A, Hamaji M, Date H, Haga H, Marx A. POU2F3 beyond thymic carcinomas: expression across the spectrum of thymomas hints to medullary differentiation in type A thymoma. Virchows Archiv: Int J Pathol. 2022;480:843–51. https://doi.org/10.1007/s00428-021-03229-9.
Ströbel P, Hartmann E, Rosenwald A, Kalla J, Ott G, Friedel G, Schalke B, Kasahara M, Tomaru U, Marx A. Corticomedullary differentiation and maturational arrest in thymomas. Histopathology. 2014;64:557–66. https://doi.org/10.1111/his.12279.
Bomidi C, Robertson M, Coarfa C, Estes M, Blutt S. Single-cell sequencing of rotavirus-infected intestinal epithelium reveals cell-type specific epithelial repair and tuft cell infection. Proc Natl Acad Sci U S A. 2021;118. https://doi.org/10.1073/pnas.2112814118
Wilen C, Lee S, Hsieh L, Orchard R, Desai C, Hykes B, McAllaster M, Balce D, Feehley T, Brestoff J, Hickey C, Yokoyama C, Wang Y, MacDuff D, Kreamalmayer D, Howitt M, Neil J, Cadwell K, Allen P, Handley S, van Lookeren Campagne M, Baldridge M, Virgin H. Tropism for tuft cells determines immune promotion of norovirus pathogenesis. Science (New York, NY). 2016;360:204–8. https://doi.org/10.1126/science.aar3799.
Melms J, Biermann J, Huang H, Wang Y, Nair A, Tagore S, Katsyv I, Rendeiro A, Amin A, Schapiro D, Frangieh C, Luoma A, Filliol A, Fang Y, Ravichandran H, Clausi M, Alba G, Rogava M, Chen S, Ho P, Montoro D, Kornberg A, Han A, Bakhoum M, Anandasabapathy N, Suárez-Fariñas M, Bakhoum S, Bram Y, Borczuk A, Guo X, Lefkowitch J, Marboe C, Lagana S, Del Portillo A, Tsai E, Zorn E, Markowitz G, Schwabe R, Schwartz R, Elemento O, Saqi A, Hibshoosh H, Que J, Izar B. A molecular single-cell lung atlas of lethal COVID-19. Nature. 2021;595:114–9. https://doi.org/10.1038/s41586-021-03569-1.
Ma D, Liu S, Hu L, He Q, Shi W, Yan D, Cao Y, Zhang G, Wang Z, Wu J, Jiang C. Single-cell RNA sequencing identify SDCBP in ACE2-positive bronchial epithelial cells negatively correlates with COVID-19 severity. J Cell Mol Med. 2021;25:7001–12. https://doi.org/10.1111/jcmm.16714.
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu NH, Nitsche A, Müller MA, Drosten C, Pöhlmann S. SARS-CoV-2 Cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181:271-280.e8. https://doi.org/10.1016/j.cell.2020.02.052.
Seyedpour S, Khodaei B, Loghman AH, Seyedpour N, Kisomi MF, Balibegloo M, Nezamabadi SS, Gholami B, Saghazadeh A, Rezaei N. Targeted therapy strategies against SARS-CoV-2 cell entry mechanisms: a systematic review of in vitro and in vivo studies. J Cell Physiol. 2021;236:2364–92. https://doi.org/10.1002/jcp.30032.
Bruchez A, Sha K, Johnson J, Chen L, Stefani C, McConnell H, Gaucherand L, Prins R, Matreyek K, Hume A, Mühlberger E, Schmidt E, Olinger G, Stuart L, Lacy-Hulbert A. MHC class II transactivator CIITA induces cell resistance to Ebola virus and SARS-like coronaviruses. Science (New York NY). 2020;370:241–7. https://doi.org/10.1126/science.abb3753.
Lins M, Smaniotto S. Potential impact of SARS-CoV-2 infection on the thymus. Can J Microbiol. 2021;67:23–8. https://doi.org/10.1139/cjm-2020-0170.
Acknowledgements
We would like to give our sincere gratitude to the reviewers for their constructive comments.
Author information
Authors and Affiliations
Contributions
The manuscript was written by JS and YRC. MXL and YRZ took part in writing the manuscript. YMX made all the figures of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent to publication
Not applicable.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, J., Li, Mx., Xie, Ym. et al. Thymic tuft cells: potential “regulators” of non-mucosal tissue development and immune response. Immunol Res 71, 554–564 (2023). https://doi.org/10.1007/s12026-023-09372-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12026-023-09372-6