Abstract
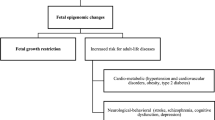
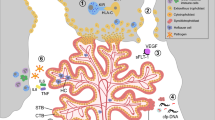
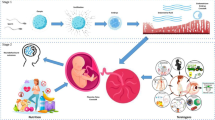
The placenta stands out as a unique, transitory, and multifaceted organ, essential to the optimal growth and maturation of the fetus. Functioning as a vital nexus between the maternal and fetal circulatory systems, it oversees the critical exchange of nutrients and waste. This exchange is facilitated by placental cells, known as trophoblasts, which adeptly invade and remodel uterine blood vessels. Deviations in placental development underpin a slew of pregnancy complications, notably fetal growth restriction (FGR), preeclampsia (PE), recurrent spontaneous abortions (RSA), and preterm birth. Central to placental function and development is epigenetic regulation. Despite its importance, the intricate mechanisms by which epigenetics influence the placenta are not entirely elucidated. Recently, the scientific community has turned its focus to parsing out the epigenetic alterations during placental development, such as variations in promoter DNA methylation, genomic imprints, and shifts in non-coding RNA expression. By establishing correlations between epigenetic shifts in the placenta and pregnancy complications, researchers are unearthing invaluable insights into the biology and pathophysiology of these conditions. This review seeks to synthesize the latest findings on placental epigenetic regulation, spotlighting its crucial role in shaping fetal growth trajectories and development. Through this lens, we underscore the overarching significance of the placenta in the larger narrative of gestational health.
Graphical Abstract




Similar content being viewed by others
Data Availability
Data availability is not applicable to this article as no new data were created or analyzed in this study.
Abbreviations
- ACE :
-
Angiotensin-converting enzyme
- AV :
-
Arteriovenous
- CTBs :
-
Cytotrophoblasts
- CuZn–SOD :
-
Antioxidant copper-zinc superoxide dismutase
- C14MC :
-
Chromosome 14 miRNA cluster
- C19MC :
-
Chromosome 19 miRNA cluster
- DMRs :
-
Differentially methylated regions
- dNK :
-
Decidual natural killer
- ECM :
-
Extracellular matrix
- EG-VEGF :
-
Endocrine gland-derived vascular endothelial growth factor
- EIcircRNA :
-
Exonic circRNA
- ESCd :
-
BAP treated hESC
- EVTs :
-
Extravillous trophoblasts
- EZH2 :
-
Enhancer of Zeste Homolog 2
- FASLG :
-
FAS ligand
- FGR :
-
Fetal growth restriction
- GDM :
-
Gestational diabetes mellitus
- GLUTs :
-
Glucose transporter proteins
- GNG7 :
-
G protein γ 7
- hCG :
-
Human chorionic gonadotropin
- HDACs :
-
Histone deacetylases
- hESCs :
-
Human embryonic stem cells
- HLA :
-
Human leukocyte antigen
- HMTs :
-
Histone methyltransferases
- HNE :
-
4-Hydroxynonenal
- HO-1 :
-
Heme oxygenase-1
- hTSCs :
-
Human pluripotent stem cells
- MHC :
-
Major histocompatibility complex
- MMPs :
-
Matrix metalloproteinases
- Nrf2 :
-
Nuclear factor erythroid 2-like protein 2
- PE :
-
Preeclampsia
- PlGF :
-
Placental Growth Factor
- RSA :
-
Recurrent spontaneous abortions
- SCNT :
-
Somatic cell nuclear transfer
- sEng :
-
Soluble endoglin
- sFlt-1 :
-
Soluble fms-like tyrosine kinase-1
- Slc38a2 :
-
Sodium-coupled neutral amino acid transporter 2
- STBs :
-
Syncytiotrophoblasts
- TET2/TET3 :
-
DNA demethylases
- TIMPs :
-
Matrix metalloproteinases tissue inhibitors
- TGF :
-
Transforming growth factor
- TRAF6 :
-
Tumor necrosis factor receptor-associated factor 6
- TWEAK :
-
Tumor necrosis factor-like weak inducer of apoptosis
- uNK :
-
Uterine natural killer cells
References
Maltepe, E., Bakardjiev, A. I., & Fisher, S. J. (2010). The placenta: Transcriptional, epigenetic, and physiological integration during development. The Journal of Clinical Investigation, 120, 1016–1025.
Reik, W., & Walter, J. (2001). Genomic imprinting: Parental influence on the genome. Nature Reviews Genetics, 2, 21–32.
Tobi, E. W., van den Heuvel, J., Zwaan, B. J., Lumey, L. H., Heijmans, B. T., & Uller, T. (2018). Selective Survival of Embryos Can Explain DNA Methylation Signatures of Adverse Prenatal Environments. Cell Reports, 25(2660–7), e4.
Nugent, B. M., & Bale, T. L. (2015). The omniscient placenta: Metabolic and epigenetic regulation of fetal programming. Frontiers in Neuroendocrinology, 39, 28–37.
Hemberger, M., Hanna, C. W., & Dean, W. (2020). Mechanisms of early placental development in mouse and humans. Nature Reviews Genetics, 21, 27–43.
Gude, N. M., Roberts, C. T., Kalionis, B., & King, R. G. (2004). Growth and function of the normal human placenta. Thrombosis Research, 114, 397–407.
Gundling, W. E., Jr., & Wildman, D. E. (2015). A review of inter- and intraspecific variation in the eutherian placenta. Philosophical Transactions of the Royal Society of London. Series B, Biological sciences, 370, 20140072.
Turco, M., Y., & Moffett, A. (2019) Development of the human placenta. Development. 146(22), dev163428. https://doi.org/10.1242/dev.163428
Carter, A. M. (1997). When is the maternal placental circulation established in man? 1941. Placenta, 18, 83–87.
Demir, R., Kaufmann, P., Castellucci, M., Erbengi, T., & Kotowski, A. (1989). Fetal vasculogenesis and angiogenesis in human placental villi. Acta Anatomica (Basel), 136, 190–203.
Aiko, Y., Askew, D. J., Aramaki, S., Myoga, M., Tomonaga, C., Hachisuga, T., et al. (2014). Differential levels of amino acid transporters System L and ASCT2, and the mTOR protein in placenta of preeclampsia and IUGR. BMC Pregnancy and Childbirth, 14, 181.
Burton, G. J., Watson, A. L., Hempstock, J., Skepper, J. N., & Jauniaux, E. (2002). Uterine glands provide histiotrophic nutrition for the human fetus during the first trimester of pregnancy. Journal of Clinical Endocrinology and Metabolism, 87, 2954–2959.
Illsley, N. P. (2000). Glucose transporters in the human placenta. Placenta, 21, 14–22.
Cariappa, R., Heath-Monnig, E., & Smith, C. H. (2003). Isoforms of amino acid transporters in placental syncytiotrophoblast: Plasma membrane localization and potential role in maternal/fetal transport. Placenta, 24, 713–726.
Stulc, J. (1997). Placental transfer of inorganic ions and water. Physiological Reviews, 77, 805–836.
Shennan, D. B., & Boyd, C. A. (1987). Ion transport by the placenta: A review of membrane transport systems. Biochimica et Biophysica Acta, 906, 437–457.
Costa, M. A. (2016). The endocrine function of human placenta: An overview. Reproductive Biomedicine Online, 32, 14–43.
Liu, Z., Wang, C., Pei, J., Li, M., & Gu, W. (2022). SIRT1: A Novel Protective Molecule in Pre-eclampsia. International Journal of Medical Sciences, 19, 993–1002.
Yu, Y., He, J. H., Hu, L. L., Jiang, L. L., Fang, L., Yao, G. D., et al. (2020). Placensin is a glucogenic hormone secreted by human placenta. EMBO Reports, 21, e49530.
Brady, P. C., Farland, L. V., Racowsky, C., & Ginsburg, E. S. (2020). Hyperglycosylated human chorionic gonadotropin as a predictor of ongoing pregnancy. American Journal of Obstetrics and Gynecology, 222(68), e1–e12.
Manaster, I., Mizrahi, S., Goldman-Wohl, D., Sela, H. Y., Stern-Ginossar, N., Lankry, D., et al. (2008). Endometrial NK cells are special immature cells that await pregnancy. The Journal of Immunology, 181, 1869–1876.
Huhn, O., Zhao, X., Esposito, L., Moffett, A., Colucci, F., & Sharkey, A. M. (2021). How Do Uterine Natural Killer and Innate Lymphoid Cells Contribute to Successful Pregnancy? Frontiers in Immunology, 12, 607669.
Lachapelle, M. H., Miron, P., Hemmings, R., & Roy, D. C. (1996). Endometrial T, B, and NK cells in patients with recurrent spontaneous abortion Altered profile and pregnancy outcome. The Journal of Immunology, 156, 4027–34.
Rutkowski, K., Sowa, P., Rutkowska-Talipska, J., Kuryliszyn-Moskal, A., & Rutkowski, R. (2014). Dehydroepiandrosterone (DHEA): Hypes and hopes. Drugs, 74, 1195–1207.
Coulam, C. B., & Roussev, R. G. (2003). Correlation of NK cell activation and inhibition markers with NK cytoxicity among women experiencing immunologic implantation failure after in vitro fertilization and embryo transfer. Journal of Assisted Reproduction and Genetics, 20, 58–62.
Parham, P. (2004). NK cells and trophoblasts: Partners in pregnancy. Journal of Experimental Medicine, 200, 951–955.
Kovats, S., Main, E. K., Librach, C., Stubblebine, M., Fisher, S. J., & DeMars, R. (1990). A class I antigen, HLA-G, expressed in human trophoblasts. Science, 248, 220–223.
Smith, G. C. (2010). First-trimester determination of complications of late pregnancy. JAMA, 303, 561–562.
Burton, G. J., Fowden, A. L., & Thornburg, K. L. (2016). Placental Origins of Chronic Disease. Physiological Reviews, 96, 1509–1565.
Goel, A., Maski, M. R., Bajracharya, S., Wenger, J. B., Zhang, D., Salahuddin, S., et al. (2015). Epidemiology and Mechanisms of De Novo and Persistent Hypertension in the Postpartum Period. Circulation, 132, 1726–1733.
Rana, S., Lemoine, E., Granger, J. P., & Karumanchi, S. A. (2019). Preeclampsia: Pathophysiology, Challenges, and Perspectives. Circulation Research, 124, 1094–1112.
Jim, B., & Karumanchi, S. A. (2017). Preeclampsia: Pathogenesis, Prevention, and Long-Term Complications. Seminars in Nephrology, 37, 386–397.
Phipps, E., Prasanna, D., Brima, W., & Jim, B. (2016). Preeclampsia: Updates in Pathogenesis, Definitions, and Guidelines. Clinical Journal of the American Society of Nephrology, 11, 1102–1113.
Karumanchi, S. A. (2016). Angiogenic Factors in Preeclampsia: From Diagnosis to Therapy. Hypertension, 67, 1072–1079.
Young, B. C., Levine, R. J., & Karumanchi, S. A. (2010). Pathogenesis of preeclampsia. Annual Review of Pathology: Mechanisms of Disease, 5, 173–192.
El-Sayed, A. A. F. (2017). Preeclampsia: A review of the pathogenesis and possible management strategies based on its pathophysiological derangements. Taiwanese Journal of Obstetrics & Gynecology, 56, 593–598.
Wang, A., Rana, S., & Karumanchi, S. A. (2009). Preeclampsia: The role of angiogenic factors in its pathogenesis. Physiology (Bethesda, Md.), 24, 147–158.
Mustafa, R., Ahmed, S., Gupta, A., & Venuto, R. C. (2012). A comprehensive review of hypertension in pregnancy. Journal of Pregnancy, 2012, 105918.
Spradley, F. T. (2019). Sympathetic nervous system control of vascular function and blood pressure during pregnancy and preeclampsia. Journal of Hypertension, 37, 476–487.
Kweider, N., Wruck, C. J., & Rath, W. (2013). New Insights into the Pathogenesis of Preeclampsia - The Role of Nrf2 Activators and their Potential Therapeutic Impact. Geburtshilfe und Frauenheilkunde, 73, 1236–1240.
Lai, W. S., & Ding, Y. L. (2019). GNG7 silencing promotes the proliferation and differentiation of placental cytotrophoblasts in preeclampsia rats through activation of the mTOR signaling pathway. International Journal of Molecular Medicine, 43, 1939–1950.
Parchem, J. G., Kanasaki, K., Kanasaki, M., Sugimoto, H., Xie, L., Hamano, Y., et al. (2018). Loss of placental growth factor ameliorates maternal hypertension and preeclampsia in mice. The Journal of Clinical Investigation, 128, 5008–5017.
Resnik, R. (2002). Intrauterine growth restriction. Obstetrics and Gynecology, 99, 490–496.
Kaufmann, P., Black, S., & Huppertz, B. (2003). Endovascular trophoblast invasion: Implications for the pathogenesis of intrauterine growth retardation and preeclampsia. Biology of Reproduction, 69, 1–7.
Hafner, E., Metzenbauer, M., Hofinger, D., Munkel, M., Gassner, R., Schuchter, K., et al. (2003). Placental growth from the first to the second trimester of pregnancy in SGA-foetuses and pre-eclamptic pregnancies compared to normal foetuses. Placenta, 24, 336–342.
Proctor, L. K., Toal, M., Keating, S., Chitayat, D., Okun, N., Windrim, R. C., et al. (2009). Placental size and the prediction of severe early-onset intrauterine growth restriction in women with low pregnancy-associated plasma protein-A. Ultrasound in Obstetrics and Gynecology, 34, 274–282.
Jones, S., Bischof, H., Lang, I., Desoye, G., Greenwood, S. L., Johnstone, E. D., et al. (2015). Dysregulated flow-mediated vasodilatation in the human placenta in fetal growth restriction. Journal of Physiology, 593, 3077–3092.
Hayward, C. E., Lean, S., Sibley, C. P., Jones, R. L., Wareing, M., Greenwood, S. L., et al. (2016). Placental Adaptation: What Can We Learn from Birthweight: Placental Weight Ratio? Frontiers in Physiology, 7, 28.
Vaughan, O. R., Maksym, K., Silva, E., Barentsen, K., Anthony, R. V., Brown, T. L., et al. (2021). Placenta-specific Slc38a2/SNAT2 knockdown causes fetal growth restriction in mice. Clinical Science (London, England), 135, 2049–2066.
Damodaram, M., Story, L., Eixarch, E., Patel, A., McGuinness, A., Allsop, J., et al. (2010). Placental MRI in intrauterine fetal growth restriction. Placenta, 31, 491–498.
Krebs, C., Macara, L. M., Leiser, R., Bowman, A. W., Greer, I. A., & Kingdom, J. C. (1996). Intrauterine growth restriction with absent end-diastolic flow velocity in the umbilical artery is associated with maldevelopment of the placental terminal villous tree. American Journal of Obstetrics and Gynecology, 175, 1534–1542.
Chen, C. P., Bajoria, R., & Aplin, J. D. (2002). Decreased vascularization and cell proliferation in placentas of intrauterine growth-restricted fetuses with abnormal umbilical artery flow velocity waveforms. American Journal of Obstetrics and Gynecology, 187, 764–769.
Junaid, T. O., Brownbill, P., Chalmers, N., Johnstone, E. D., & Aplin, J. D. (2014). Fetoplacental vascular alterations associated with fetal growth restriction. Placenta, 35, 808–815.
El Hachem, H., Crepaux, V., May-Panloup, P., Descamps, P., Legendre, G., & Bouet, P. E. (2017). Recurrent pregnancy loss: Current perspectives. Int J Womens Health., 9, 331–345.
Hustin, J., Jauniaux, E., & Schaaps, J. P. (1990). Histological study of the materno-embryonic interface in spontaneous abortion. Placenta, 11, 477–486.
Gupta, S. K., Malhotra, S. S., Malik, A., Verma, S., & Chaudhary, P. (2016). Cell Signaling Pathways Involved During Invasion and Syncytialization of Trophoblast Cells. American Journal of Reproductive Immunology, 75, 361–371.
Abrahams, V. M., Visintin, I., Aldo, P. B., Guller, S., Romero, R., & Mor, G. (2005). A role for TLRs in the regulation of immune cell migration by first trimester trophoblast cells. The Journal of Immunology, 175, 8096–8104.
Harris, L. K., Smith, S. D., Keogh, R. J., Jones, R. L., Baker, P. N., Knofler, M., et al. (2010). Trophoblast- and vascular smooth muscle cell-derived MMP-12 mediates elastolysis during uterine spiral artery remodeling. American Journal of Pathology, 177, 2103–2115.
Zhu, J. Y., Pang, Z. J., & Yu, Y. H. (2012). Regulation of trophoblast invasion: The role of matrix metalloproteinases. Reviews in Obstetrics & Gynecology, 5, e137–e143.
Pollheimer, J., Vondra, S., Baltayeva, J., Beristain, A. G., & Knofler, M. (2018). Regulation of Placental Extravillous Trophoblasts by the Maternal Uterine Environment. Frontiers in Immunology, 9, 2597.
Deng, W., Cha, J., Yuan, J., Haraguchi, H., Bartos, A., Leishman, E., et al. (2016). p53 coordinates decidual sestrin 2/AMPK/mTORC1 signaling to govern parturition timing. The Journal of Clinical Investigation, 126, 2941–2954.
Du, L., Deng, W., Zeng, S., Xu, P., Huang, L., Liang, Y., et al. (2021). Single-cell transcriptome analysis reveals defective decidua stromal niche attributes to recurrent spontaneous abortion. Cell Proliferation, 54, e13125.
Hocher, B., & Hocher, C. F. (2018). Epigenetics of recurrent pregnancy loss. EBioMedicine., 35, 18–19.
Cross, J. C. (2003). The genetics of pre-eclampsia: A feto-placental or maternal problem? Clinical Genetics, 64, 96–103.
Menon, R., Taylor, R. N., & Fortunato, S. J. (2010). Chorioamnionitis–a complex pathophysiologic syndrome. Placenta, 31, 113–120.
Xue, W. C., Chan, K. Y., Feng, H. C., Chiu, P. M., Ngan, H. Y., Tsao, S. W., et al. (2004). Promoter hypermethylation of multiple genes in hydatidiform mole and choriocarcinoma. The Journal of Molecular Diagnostics, 6, 326–334.
Pliushch, G., Schneider, E., Weise, D., El Hajj, N., Tresch, A., Seidmann, L., et al. (2010). Extreme methylation values of imprinted genes in human abortions and stillbirths. American Journal of Pathology, 176, 1084–1090.
Vasconcelos, S., Ramalho, C., Marques, C. J., & Doria, S. (2019). Altered expression of epigenetic regulators and imprinted genes in human placenta and fetal tissues from second trimester spontaneous pregnancy losses. Epigenetics, 14, 1234–1244.
Yuen, R. K., & Robinson, W. P. (2011). Review: A high capacity of the human placenta for genetic and epigenetic variation: Implications for assessing pregnancy outcome. Placenta, 32(Suppl 2), S136–S141.
Lim, Y. C., Li, J., Ni, Y., Liang, Q., Zhang, J., Yeo, G. S. H., et al. (2017). A complex association between DNA methylation and gene expression in human placenta at first and third trimesters. PLoS ONE, 12, e0181155.
Ohinata, Y., Payer, B., O’Carroll, D., Ancelin, K., Ono, Y., Sano, M., et al. (2005). Blimp1 is a critical determinant of the germ cell lineage in mice. Nature, 436, 207–213.
Du, G., Yu, M., Xu, Q., Huang, Z., Huang, X., Han, L., et al. (2020). Hypomethylation of PRDM1 is associated with recurrent pregnancy loss. Journal of Cellular and Molecular Medicine, 24, 7072–7077.
Wu, A. H., Guo, L. Y., Lu, S., Chen, X. L., Wang, A. A., Wang, X. Y., et al. (2020). Aberrant methylation of IGF2-AS promoter in early pregnancy loss. Taiwanese Journal of Obstetrics & Gynecology, 59, 109–114.
Yu, M., Du, G., Xu, Q., Huang, Z., Huang, X., Qin, Y., et al. (2018). Integrated analysis of DNA methylome and transcriptome identified CREB5 as a novel risk gene contributing to recurrent pregnancy loss. eBioMedicine, 35, 334–344.
Hanna, C. W., McFadden, D. E., & Robinson, W. P. (2013). DNA methylation profiling of placental villi from karyotypically normal miscarriage and recurrent miscarriage. American Journal of Pathology, 182, 2276–2284.
Yin, L. J., Zhang, Y., Lv, P. P., He, W. H., Wu, Y. T., Liu, A. X., et al. (2012). Insufficient maintenance DNA methylation is associated with abnormal embryonic development. BMC Medicine, 10, 26.
Voss, A. K., & Thomas, T. (2018). Histone Lysine and Genomic Targets of Histone Acetyltransferases in Mammals. BioEssays, 40, e1800078.
Gebremedhin, K. G., & Rademacher, D. J. (2016). Histone H3 acetylation in the postmortem Parkinson’s disease primary motor cortex. Neuroscience Letters, 627, 121–125.
Tsaprouni, L. G., Ito, K., Powell, J. J., Adcock, I. M., & Punchard, N. (2011). Differential patterns of histone acetylation in inflammatory bowel diseases. Journal of Inflammation (London), 8, 1.
Eddy, A. C., Chapman, H., & George, E. M. (2019). Acute Hypoxia and Chronic Ischemia Induce Differential Total Changes in Placental Epigenetic Modifications. Reproductive Sciences, 26, 766–773.
Seto, E., & Yoshida, M. (2014). Erasers of histone acetylation: The histone deacetylase enzymes. Cold Spring Harbor Perspectives in Biology, 6, a018713.
Togher, K. L., Kenny, L. C., & O’Keeffe, G. W. (2017). Class-Specific Histone Deacetylase Inhibitors Promote 11-Beta Hydroxysteroid Dehydrogenase Type 2 Expression in JEG-3 Cells. International Journal of Cell Biology, 2017, 6169310.
Wang, Y., Gu, Y., Alexander, J. S., & Lewis, D. F. (2019). Histone deacetylase inhibition disturbs the balance between ACE and chymase expression in endothelial cells: A potential mechanism of chymase activation in preeclampsia. Hypertension Research, 42, 155–164.
Cruz-Munoz, W., Sanchez, O. H., Di Grappa, M., English, J. L., Hill, R. P., & Khokha, R. (2006). Enhanced metastatic dissemination to multiple organs by melanoma and lymphoma cells in timp-3-/- mice. Oncogene, 25, 6489–6496.
Xie, D., Zhu, J., Liu, Q., Li, J., Song, M., Wang, K., et al. (2019). Dysregulation of HDAC9 Represses Trophoblast Cell Migration and Invasion Through TIMP3 Activation in Preeclampsia. American Journal of Hypertension, 32, 515–523.
Tasta, O., Swiader, A., Grazide, M. H., Rouahi, M., Parant, O., Vayssiere, C., et al. (2021). A role for 4-hydroxy-2-nonenal in premature placental senescence in preeclampsia and intrauterine growth restriction. Free Radical Biology & Medicine, 164, 303–314.
Tran, T. Q., Lowman, X. H., & Kong, M. (2017). Molecular Pathways: Metabolic Control of Histone Methylation and Gene Expression in Cancer. Clinical Cancer Research, 23, 4004–4009.
Black, J. C., Van Rechem, C., & Whetstine, J. R. (2012). Histone lysine methylation dynamics: Establishment, regulation, and biological impact. Molecular Cell, 48, 491–507.
Sheng, W., Gu, Y., Chu, X., Morgan, J. A., Cooper, D. B., Lewis, D. F., et al. (2021). Upregulation of histone H3K9 methylation in fetal endothelial cells from preeclamptic pregnancies. Journal of Cellular Physiology, 236, 1866–1874.
Padeken, J., Methot, S. P., & Gasser, S. M. (2022). Establishment of H3K9-methylated heterochromatin and its functions in tissue differentiation and maintenance. Nature Reviews Molecular Cell Biology, 23, 623–640.
Matsui, H., Iriyama, T., Sayama, S., Inaoka, N., Suzuki, K., Yoshikawa, M., et al. (2021). Elevated placental histone H3K4 methylation via upregulated histone methyltransferases SETD1A and SMYD3 in preeclampsia and its possible involvement in hypoxia-induced pathophysiological process. Placenta, 115, 60–69.
Sirohi, V. K., Medrano, T. I., Kannan, A., Bagchi, I. C., & Cooke, P. S. (2023). Uterine-specific Ezh2 deletion enhances stromal cell senescence and impairs placentation, resulting in pregnancy loss. Science, 26, 107028.
Mattick, J. S., & Makunin, I. V. (2006). Non-coding RNA. Human molecular genetics, 15(suppl_1), R17–R29. https://doi.org/10.1093/hmg/ddl046
Chen, B., & Huang, S. (2018). Circular RNA: An emerging non-coding RNA as a regulator and biomarker in cancer. Cancer Letters, 418, 41–50.
Bartel, D. P. (2004). MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell, 116, 281–297.
Bushati, N., & Cohen, S. M. (2007). microRNA functions. Annual Review of Cell and Developmental Biology, 23, 175–205.
Morales-Prieto, D. M., Ospina-Prieto, S., Chaiwangyen, W., Schoenleben, M., & Markert, U. R. (2013). Pregnancy-associated miRNA-clusters. Journal of Reproductive Immunology, 97, 51–61.
Liang, Y., Ridzon, D., Wong, L., & Chen, C. (2007). Characterization of microRNA expression profiles in normal human tissues. BMC Genomics, 8, 166.
Kshitiz, Afzal, J., Maziarz, J. D., Hamidzadeh, A., Liang, C., Erkenbrack, E. M., ... & Wagner, G. P. (2019). Evolution of placental invasion and cancer metastasis are causally linked. Nature Ecology & Evolution, 3(12), 1743–1753. https://doi.org/10.1038/s41559-019-1046-4
Gonzalez, T. L., Eisman, L. E., Joshi, N. V., Flowers, A. E., Wu, D., Wang, Y., et al. (2021). High-throughput miRNA sequencing of the human placenta: Expression throughout gestation. Epigenomics, 13, 995–1012.
Zhang, H., He, Y., Wang, J. X., Chen, M. H., Xu, J. J., Jiang, M. H., et al. (2020). miR-30-5p-mediated ferroptosis of trophoblasts is implicated in the pathogenesis of preeclampsia. Redox Biology, 29, 101402.
Ding, J., Zhang, Y., Cai, X., Zhang, Y., Yan, S., Wang, J., et al. (2021). Extracellular vesicles derived from M1 macrophages deliver miR-146a-5p and miR-146b-5p to suppress trophoblast migration and invasion by targeting TRAF6 in recurrent spontaneous abortion. Theranostics., 11, 5813–5830.
Su, M. T., Tsai, P. Y., Tsai, H. L., Chen, Y. C., & Kuo, P. L. (2017). miR-346 and miR-582-3p-regulated EG-VEGF expression and trophoblast invasion via matrix metalloproteinases 2 and 9. BioFactors, 43, 210–219.
Herman, A. B., Tsitsipatis, D., & Gorospe, M. (2022). Integrated lncRNA function upon genomic and epigenomic regulation. Molecular Cell, 82, 2252–2266.
Yu, J., Hong, J. F., Kang, J., Liao, L. H., & Li, C. D. (2017). Promotion of LncRNA HOXA11-AS on the proliferation of hepatocellular carcinoma by regulating the expression of LATS1. European Review for Medical and Pharmacological Sciences, 21, 3402–3411.
Zhang, Q., Wang, Z., Cheng, X., & Wu, H. (2021). lncRNA DANCR promotes the migration an invasion and of trophoblast cells through microRNA-214-5p in preeclampsia. Bioengineered, 12, 9424–9434.
Ogoyama, M., Ohkuchi, A., Takahashi, H., Zhao, D., Matsubara, S., Takizawa, T. (2021). LncRNA H19-derived miR-675-5p accelerates the invasion of extravillous trophoblast cells by inhibiting GATA2 and Subsequently activating matrix metalloproteinases. International Journal of Molecular Sciences, 22(3), 1237. https://doi.org/10.3390/ijms22031237
Zhang, L., Deng, X., Shi, X., & Dong, X. (2019). Silencing H19 regulated proliferation, invasion, and autophagy in the placenta by targeting miR-18a-5p. Journal of Cellular Biochemistry, 120, 9006–9015.
Xu, J., Xia, Y., Zhang, H., Guo, H., Feng, K., & Zhang, C. (2018). Overexpression of long non-coding RNA H19 promotes invasion and autophagy via the PI3K/AKT/mTOR pathways in trophoblast cells. Biomedicine & Pharmacotherapy, 101, 691–697.
Wu, L., Liu, Q., Fan, C., Yi, X., & Cheng, B. (2021). MALAT1 recruited the E3 ubiquitin ligase FBXW7 to induce CRY2 ubiquitin-mediated degradation and participated in trophoblast migration and invasion. Journal of Cellular Physiology, 236, 2169–2177.
Chen, H., Meng, T., Liu, X., Sun, M., Tong, C., Liu, J., et al. (2015). Long non-coding RNA MALAT-1 is downregulated in preeclampsia and regulates proliferation, apoptosis, migration and invasion of JEG-3 trophoblast cells. International Journal of Clinical and Experimental Pathology, 8, 12718–12727.
Wu, H. Y., Wang, X. H., Liu, K., & Zhang, J. L. (2020). LncRNA MALAT1 regulates trophoblast cells migration and invasion via miR-206/IGF-1 axis. Cell Cycle, 19, 39–52.
Wang, R., & Zou, L. (2020). Downregulation of LncRNA-MEG3 promotes HTR8/SVneo cells apoptosis and attenuates its migration by repressing Notch1 signal in preeclampsia. Reproduction, 160, 21–29.
Yu, L., Kuang, L. Y., He, F., Du, L. L., Li, Q. L., Sun, W., et al. (2018). The Role and Molecular Mechanism of Long Nocoding RNA-MEG3 in the Pathogenesis of Preeclampsia. Reproductive Sciences, 25, 1619–1628.
Zhang, J., Liu, X., & Gao, Y. (2021). The long noncoding RNA MEG3 regulates Ras-MAPK pathway through RASA1 in trophoblast and is associated with unexplained recurrent spontaneous abortion. Molecular Medicine, 27, 70.
Hansen, T. B., Jensen, T. I., Clausen, B. H., Bramsen, J. B., Finsen, B., Damgaard, C. K., et al. (2013). Natural RNA circles function as efficient microRNA sponges. Nature, 495, 384–388.
Floris, G., Zhang, L., Follesa, P., & Sun, T. (2017). Regulatory Role of Circular RNAs and Neurological Disorders. Molecular Neurobiology, 54, 5156–5165.
Cheng, J., Huang, J., Yuan, S., Zhou, S., Yan, W., Shen, W., et al. (2017). Circular RNA expression profiling of human granulosa cells during maternal aging reveals novel transcripts associated with assisted reproductive technology outcomes. PLoS ONE, 12, e0177888.
Qian, Y., Lu, Y., Rui, C., Qian, Y., Cai, M., & Jia, R. (2016). Potential Significance of Circular RNA in Human Placental Tissue for Patients with Preeclampsia. Cellular Physiology and Biochemistry, 39, 1380–1390.
Zhang, Y. G., Yang, H. L., Long, Y., & Li, W. L. (2016). Circular RNA in blood corpuscles combined with plasma protein factor for early prediction of pre-eclampsia. BJOG, 123, 2113–2118.
Hu, X., Ao, J., Li, X., Zhang, H., Wu, J., & Cheng, W. (2018). Competing endogenous RNA expression profiling in pre-eclampsia identifies hsa_circ_0036877 as a potential novel blood biomarker for early pre-eclampsia. Clinical Epigenetics, 10, 48.
Maass, P. G., Glazar, P., Memczak, S., Dittmar, G., Hollfinger, I., Schreyer, L., et al. (2017). A map of human circular RNAs in clinically relevant tissues. Journal of Molecular Medicine (Berlin, Germany), 95, 1179–1189.
Yan, L., Feng, J., Cheng, F., Cui, X., Gao, L., Chen, Y., et al. (2018). Circular RNA expression profiles in placental villi from women with gestational diabetes mellitus. Biochemical and Biophysical Research Communications, 498, 743–750.
Wang, H., She, G., Zhou, W., Liu, K., Miao, J., & Yu, B. (2019). Expression profile of circular RNAs in placentas of women with gestational diabetes mellitus. Endocrine Journal, 66, 431–441.
Zhang, Y., Yang, H., Zhang, Y., Shi, J., Chen, R., & Xiao, X. (2020). CircSFXN1 regulates the behaviour of trophoblasts and likely mediates preeclampsia. Placenta, 101, 115–123.
Gai, S., Sun, L., Wang, H., & Yang, P. (2020). Circular RNA hsa_circ_0007121 regulates proliferation, migration, invasion, and epithelial-mesenchymal transition of trophoblast cells by miR-182-5p/PGF axis in preeclampsia. Open Med (Wars)., 15, 1061–1071.
Tang, R., Zhang, Z., & Han, W. (2021). CircLRRK1 targets miR-223-3p to inhibit the proliferation, migration and invasion of trophoblast cells by regulating the PI3K/AKT signaling pathway. Placenta, 104, 110–118.
Shen, X. Y., Zheng, L. L., Huang, J., Kong, H. F., Chang, Y. J., Wang, F., et al. (2019). CircTRNC18 inhibits trophoblast cell migration and epithelial-mesenchymal transition by regulating miR-762/Grhl2 pathway in pre-eclampsia. RNA Biology, 16, 1565–1573.
Zhang, S., & Guo, G. (2022). Circ_FURIN promotes trophoblast cell proliferation, migration and invasion in preeclampsia by regulating miR-34a-5p and TFAP2A. Hypertension Research, 45, 1334–1344.
Jing, M. Y., Xie, L. D., Chen, X., Zhou, Y., Jin, M. M., He, W. H., ... & Liu, A. X. (2022). Circ-CCNB1 modulates trophoblast proliferation and invasion in spontaneous abortion by regulating miR-223/SIAH1 axis. Endocrinology, 163(8), bqac093. https://doi.org/10.1210/endocr/bqac093
Zhang, Y., Yang, H., Zhang, Y., Shi, J., & Chen, R. (2020). circCRAMP1L is a novel biomarker of preeclampsia risk and may play a role in preeclampsia pathogenesis via regulation of the MSP/RON axis in trophoblasts. BMC Pregnancy and Childbirth, 20, 652.
Wang, H., Luo, C., Wu, X., Zhang, J., Xu, Z., Liu, Y., et al. (2021). Circular RNA hsa_circ_0081343 promotes trophoblast cell migration and invasion and inhibits trophoblast apoptosis by regulating miR-210-5p/DLX3 axis. Reproductive Biology and Endocrinology, 19, 123.
Wang, D., Na, Q., Song, G., Wang, Y., & Wang, Y. (2020). The Role of circRNA-SETD2/miR-519a/PTEN Axis in Fetal Birth Weight through Regulating Trophoblast Proliferation. BioMed Research International, 2020, 9809632.
Li, Z., Zhou, G., Tao, F., Cao, Y., Han, W., & Li, Q. (2020). circ-ZUFSP regulates trophoblasts migration and invasion through sponging miR-203 to regulate STOX1 expression. Biochemical and Biophysical Research Communications, 531, 472–479.
Surani, M. A., & Barton, S. C. (1983). Development of gynogenetic eggs in the mouse: Implications for parthenogenetic embryos. Science, 222, 1034–1036.
Haycock, P. C., & Ramsay, M. (2009). Exposure of mouse embryos to ethanol during preimplantation development: Effect on DNA methylation in the h19 imprinting control region. Biology of Reproduction, 81, 618–627.
Constancia, M., Pickard, B., Kelsey, G., & Reik, W. (1998). Imprinting mechanisms. Genome Research, 8, 881–900.
Wood, A. J., & Oakey, R. J. (2006). Genomic imprinting in mammals: Emerging themes and established theories. PLoS Genetics, 2, e147.
Constância, M., Hemberger, M., Hughes, J., et al. (2002). Placental-specific IGF-II is a major modulator of placental and fetal growth. Nature, 417(6892), 945–948. https://doi.org/10.1038/nature00819
Tucci, V., Isles, A. R., Kelsey, G., Ferguson-Smith, A. C., & Erice, I. G. (2019). Genomic Imprinting and Physiological Processes in Mammals. Cell, 176, 952–965.
Ogata T, Kagami M. Kagami-Ogata syndrome: a clinically recognizable upd(14)pat and related disorder affecting the chromosome 14q32.2 imprinted region. J Hum Genet. 2016; 61: 87–94.
Kagami, M., Sekita, Y., Nishimura, G., Irie, M., Kato, F., Okada, M., ... & Ogata, T. (2008). Deletions and epimutations affecting the human 14q32. 2 imprinted region in individuals with paternal and maternal upd (14)-like phenotypes. Nature Genetics, 40(2), 237–242. https://doi.org/10.1038/ng.2007.56
Sekita, Y., Wagatsuma, H., Nakamura, K., Ono, R., Kagami, M., Wakisaka, N., et al. (2008). Role of retrotransposon-derived imprinted gene, Rtl1, in the feto-maternal interface of mouse placenta. Nature Genetics, 40, 243–248.
Li, E., Beard, C., & Jaenisch, R. (1993). Role for DNA methylation in genomic imprinting. Nature, 366, 362–365.
Inoue, A., Jiang, L., Lu, F., Suzuki, T., & Zhang, Y. (2017). Maternal H3K27me3 controls DNA methylation-independent imprinting. Nature, 547, 419–424.
Chen, Z., Yin, Q., Inoue, A., Zhang, C., & Zhang, Y. (2019). Allelic H3K27me3 to allelic DNA methylation switch maintains noncanonical imprinting in extraembryonic cells. Science Advances, 5(12), eaay7246. https://doi.org/10.1126/sciadv.aay7246
Hanna, C. W., Perez-Palacios, R., Gahurova, L., Schubert, M., Krueger, F., Biggins, L., et al. (2019). Endogenous retroviral insertions drive non-canonical imprinting in extra-embryonic tissues. Genome Biology, 20, 225.
Mira-Bontenbal, H., & Gribnau, J. (2016). New Xist-Interacting Proteins in X-Chromosome Inactivation. Current Biology, 26, R338–R342.
Inoue, A., Jiang, L., Lu, F., & Zhang, Y. (2017). Genomic imprinting of Xist by maternal H3K27me3. Genes & Development, 31, 1927–1932.
Okamoto, I., Otte, A. P., Allis, C. D., Reinberg, D., & Heard, E. (2004). Epigenetic dynamics of imprinted X inactivation during early mouse development. Science, 303, 644–649.
Inoue, A., Chen, Z., Yin, Q., & Zhang, Y. (2018). Maternal Eed knockout causes loss of H3K27me3 imprinting and random X inactivation in the extraembryonic cells. Genes & Development, 32, 1525–1536.
Wilmut, I., Schnieke, A. E., McWhir, J., Kind, A. J., & Campbell, K. H. (1997). Viable offspring derived from fetal and adult mammalian cells. Nature, 385, 810–813.
Wang, X., Qu, J., Li, J., He, H., Liu, Z., & Huan, Y. (2020). Epigenetic Reprogramming During Somatic Cell Nuclear Transfer: Recent Progress and Future Directions. Frontiers in Genetics, 11, 205.
Inoue, K., Ogonuki, N., Kamimura, S., Inoue, H., Matoba, S., Hirose, M., et al. (2020). Loss of H3K27me3 imprinting in the Sfmbt2 miRNA cluster causes enlargement of cloned mouse placentas. Nature Communications, 11, 2150.
Romero, R., Kusanovic, J. P., Chaiworapongsa, T., & Hassan, S. S. (2011). Placental bed disorders in preterm labor, preterm PROM, spontaneous abortion and abruptio placentae. Best Practice & Research. Clinical Obstetrics & Gynaecology, 25, 313–327.
Windsperger, K., Dekan, S., Pils, S., Golletz, C., Kunihs, V., Fiala, C., et al. (2017). Extravillous trophoblast invasion of venous as well as lymphatic vessels is altered in idiopathic, recurrent, spontaneous abortions. Human Reproduction, 32, 1208–1217.
Zhou, Y., Damsky, C. H., & Fisher, S. J. (1997). Preeclampsia is associated with failure of human cytotrophoblasts to mimic a vascular adhesion phenotype. One cause of defective endovascular invasion in this syndrome?. The Journal of Clinical Investigation, 99(9), 2152–2164. https://doi.org/10.1172/JCI119388
Schmidt, A., Morales-Prieto, D. M., Pastuschek, J., Frohlich, K., & Markert, U. R. (2015). Only humans have human placentas: Molecular differences between mice and humans. Journal of Reproductive Immunology, 108, 65–71.
Bentwich, I., Avniel, A., Karov, Y., Aharonov, R., Gilad, S., Barad, O., et al. (2005). Identification of hundreds of conserved and nonconserved human microRNAs. Nature Genetics, 37, 766–770.
Morales-Prieto, D. M., Ospina-Prieto, S., Schmidt, A., Chaiwangyen, W., & Markert, U. R. (2014). Elsevier Trophoblast Research Award Lecture: Origin, evolution and future of placenta miRNAs. Placenta, 35(Suppl), S39-45.
Rielland, M., Hue, I., Renard, J. P., & Alice, J. (2008). Trophoblast stem cell derivation, cross-species comparison and use of nuclear transfer: New tools to study trophoblast growth and differentiation. Developmental Biology, 322, 1–10.
Wilkinson, A. L., Zorzan, I., & Rugg-Gunn, P. J. (2023). Epigenetic regulation of early human embryo development. Cell Stem Cell, 30, 1569–1584.
Cinkornpumin, J. K., Kwon, S. Y., Guo, Y., Hossain, I., Sirois, J., Russett, C. S., et al. (2020). Naive Human Embryonic Stem Cells Can Give Rise to Cells with a Trophoblast-like Transcriptome and Methylome. Stem Cell Reports., 15, 198–213.
Pastor, W. A., Chen, D., Liu, W., Kim, R., Sahakyan, A., Lukianchikov, A., et al. (2016). Naive Human Pluripotent Cells Feature a Methylation Landscape Devoid of Blastocyst or Germline Memory. Cell Stem Cell, 18, 323–329.
Takahashi, S., Okae, H., Kobayashi, N., Kitamura, A., Kumada, K., Yaegashi, N., et al. (2019). Loss of p57(KIP2) expression confers resistance to contact inhibition in human androgenetic trophoblast stem cells. Proc Natl Acad Sci U S A., 116, 26606–26613.
Sheridan, M. A., Zhao, X., Fernando, R. C., Gardner, L., Perez-Garcia, V., Li, Q., & Turco, M. Y. (2021). Characterization of primary models of human trophoblast. Development, 148(21), dev199749. https://doi.org/10.1242/dev.199749
Turco, M. Y., Gardner, L., Kay, R. G., Hamilton, R. S., Prater, M., Hollinshead, M. S., et al. (2018). Trophoblast organoids as a model for maternal-fetal interactions during human placentation. Nature, 564, 263–267.
Goustin, A. S., Betsholtz, C., Pfeifer-Ohlsson, S., Persson, H., Rydnert, J., Bywater, M., et al. (1985). Coexpression of the sis and myc proto-oncogenes in developing human placenta suggests autocrine control of trophoblast growth. Cell, 41, 301–312.
Lewis, M. P., Clements, M., Takeda, S., Kirby, P. L., Seki, H., Lonsdale, L. B., et al. (1996). Partial characterization of an immortalized human trophoblast cell-line, TCL-1, which possesses a CSF-1 autocrine loop. Placenta, 17, 137–146.
Rong-Hao, L., Luo, S., & Zhuang, L. Z. (1996). Establishment and characterization of a cytotrophoblast cell line from normal placenta of human origin. Human Reproduction, 11, 1328–1333.
Msheik, H., El Hayek, S., Bari, M. F., Azar, J., Abou-Kheir, W., Kobeissy, F., et al. (2019). Transcriptomic profiling of trophoblast fusion using BeWo and JEG-3 cell lines. Molecular Human Reproduction, 25, 811–824.
Burres, N. S., & Cass, C. E. (1986). Density-dependent inhibition of expression of syncytiotrophoblastic markers by cultured human choriocarcinoma (BeWo) cells. Journal of Cellular Physiology, 128, 375–382.
Burleigh, D. W., Kendziorski, C. M., Choi, Y. J., Grindle, K. M., Grendell, R. L., Magness, R. R., et al. (2007). Microarray analysis of BeWo and JEG3 trophoblast cell lines: Identification of differentially expressed transcripts. Placenta, 28, 383–389.
Tuan, R. S., Moore, C. J., Brittingham, J. W., Kirwin, J. J., Akins, R. E., & Wong, M. (1991). In vitro study of placental trophoblast calcium uptake using JEG-3 human choriocarcinoma cells. Journal of Cell Science, 98(Pt 3), 333–342.
Msheik, H., Azar, J., El Sabeh, M., Abou-Kheir, W., & Daoud, G. (2020). HTR-8/SVneo: A model for epithelial to mesenchymal transition in the human placenta. Placenta, 90, 90–97.
Graham, C. H., Hawley, T. S., Hawley, R. G., MacDougall, J. R., Kerbel, R. S., Khoo, N., et al. (1993). Establishment and characterization of first trimester human trophoblast cells with extended lifespan. Experimental Cell Research, 206, 204–211.
Abou-Kheir, W., Barrak, J., Hadadeh, O., & Daoud, G. (2017). HTR-8/SVneo cell line contains a mixed population of cells. Placenta, 50, 1–7.
Harun, R., Ruban, L., Matin, M., Draper, J., Jenkins, N. M., Liew, G. C., et al. (2006). Cytotrophoblast stem cell lines derived from human embryonic stem cells and their capacity to mimic invasive implantation events. Human Reproduction, 21, 1349–1358.
Genbacev, O., Donne, M., Kapidzic, M., Gormley, M., Lamb, J., Gilmore, J., et al. (2011). Establishment of human trophoblast progenitor cell lines from the chorion. Stem Cells., 29, 1427–1436.
Xu, R. H., Chen, X., Li, D. S., Li, R., Addicks, G. C., Glennon, C., et al. (2002). BMP4 initiates human embryonic stem cell differentiation to trophoblast. Nature Biotechnology, 20, 1261–1264.
Amita, M., Adachi, K., Alexenko, A. P., Sinha, S., Schust, D. J., Schulz, L. C., et al. (2013). Complete and unidirectional conversion of human embryonic stem cells to trophoblast by BMP4. Proc Natl Acad Sci U S A., 110, E1212–E1221.
Zdravkovic, T., Nazor, K. L., Larocque, N., Gormley, M., Donne, M., Hunkapillar, N., et al. (2015). Human stem cells from single blastomeres reveal pathways of embryonic or trophoblast fate specification. Development, 142, 4010–4025.
Funding
This study was supported by the Key Project on the Integration of Industry, Education and Research Collaborative Innovation of Fujian Province (No. 2021YZ034011), the Key Project on Science and Technology Program of Fujian Health Commission (No. 2021ZD01002),the Joint Funds for the Innovation of Science and Technology, Fujian Province (No: 2021Y9185),the Province-level special subsidy funds for health in Fujian Province (No: Fujian Finance Index [2019] 827),the Fujian Province health scientific research personnel training project(No: Fujian Finance Index [2021] 55),the Fujian Province health scientific research personnel training project(No: Fujian Finance Index [2021] 500).
Author information
Authors and Affiliations
Contributions
YC and ZY write of the manuscript, ML and LZ made diagrams of the manuscript, XW and LX made revisions to the manuscript.
Corresponding authors
Ethics declarations
Ethical Approval
Ethical Procedure The research meets all applicable standards with regard to the ethics of experimentation and research integrity, and the following is being certified/declared true. As an expert scientist and along with co-authors of concerned field, the paper has been submitted with full responsibility, following due ethical procedure, and there is no duplicate publication, fraud, plagiarism, or concerns about animal or human experimentation.
Consent to Participate
All authors involved in the study obtained informed consent.
Consent to Publish
Paper is containing original research and has not been submitted / published earlier in any journal and is not being considered for publication elsewhere. All authors have seen and approved the manuscript and have contributed significantly for the paper.
Conflict of Interest
None of the authors of this paper has a financial or personal relationship with other people or organizations that could inappropriately influence or bias the content of the paper. It is to specifically state that “No Competing interests are at stake and there is No Conflict of Interest” with other people or organizations that could inappropriately influence or bias the content of the paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yujia Chen and Zhoujie Ye are co-first authors.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Y., Ye, Z., Lin, M. et al. Deciphering the Epigenetic Landscape: Placental Development and Its Role in Pregnancy Outcomes. Stem Cell Rev and Rep (2024). https://doi.org/10.1007/s12015-024-10699-2
Accepted:
Published:
DOI: https://doi.org/10.1007/s12015-024-10699-2