Abstract
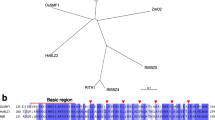
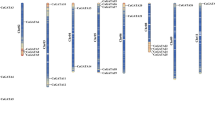
Heme activator protein (HAP) is a CCAAT box-binding factor family conserved in all higher eukaryotes. HAP transcription factors (TFs) consist of three subunits: HAP2, HAP3, and HAP5. Using recent rice genome information, we identified 11 HAP2 genes, 11 HAP3 genes, and 12 HAP5 genes. A HAP2 and five HAP5 genes were acknowledged more in this study than in the previous studies. Due to the functional association of HAP subunits, we carried out clustering analysis for expression patterns of all HAP members using anatomical meta-expression data of 983 rice Affymetrix arrays collected from the public database. Subsequently, we identified HAP members with endosperm and seed-preferred expression and focused on nine HAP family members associated with this expression pattern. We confirmed their expression patterns by RT-PCR analyses. The nine members were OsHAP2G, OsHAP2I, OsHAP3D, OsHAP3K, and five HAP5 genes (LOC_Os05g23910, LOC_Os10g11580, LOC_Os01g24460, LOC_Os01g01290, and LOC_Os01g39850). Promoter analyses using GUS activity again demonstrated that OsHAP3K is preferentially expressed in the aleurone layer of the seed and the LOC_Os05g23910 gene-encoding HAP5 showed endosperm-preferred expression. Network analysis using RiceNet suggests a functional gene network mediated by four rice HAP genes, further helping in the prediction of the biological function of OsHAP2G and OsHAP5 (LOC_Os10g11580). Our data provide new insight into HAP family-mediated regulatory pathways for endosperm or seed development in rice.





Similar content being viewed by others
References
Abe N, Asai H, Yago H et al (2014) Relationships between starch synthase I and branching enzyme isozymes determined using double mutant rice lines. BMC Plant Biol 14:1–12
Alam MM, Tanaka T, Nakamura H et al (2015) Overexpression of a rice heme activator protein gene (OsHAP2E) confers resistance to pathogens, salinity and drought, and increases photosynthesis and tiller number. Plant Biotechnol J 13:85–96
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME suite. Nucleic Acids Res 43:W39–W49
Cai X, Ballif J, Endo S et al (2007) A putative CCAAT-binding transcription factor is a regulator of flowering timing in Arabidopsis. Plant Physiol 145:98–105
Chandran AKN, Yoo YH, Cao P et al (2016) Updated Rice Kinase Database RKD 2.0: enabling transcriptome and functional analysis of rice kinase genes. Rice 9:40
Chow CN, Zheng HQ, Wu NY et al (2016) PlantPAN 2.0: an update of plant promoter analysis navigator for reconstructing transcriptional regulatory networks in plants. Nucleic Acids Res 44:D1154–D1164
Doroshenk KA, Crofts AJ, Washida H et al (2010) Characterization of the rice glup4 mutant suggests a role for the small GTPase Rab5 in the biosynthesis of carbon and nitrogen storage reserves in developing endosperm. Breed Sci 60:556–567
Dreyfuss G, Swanson MS, Piñol-Roma S (1988) Heterogeneous nuclear ribonucleoprotein particles and the pathway of mRNA formation. Trends Biochem Sci 13:86–91
Ezcurra I, Ellerström M, Wycliffe P et al (1999) Interaction between composite elements in the napA promoter: both the B-box ABA-responsive complex and the RY/G complex are necessary for seed-specific expression. Plant Mol Biol 40:699–709
Gusmaroli G, Tonelli C, Mantovani R (2001) Regulation of the CCAAT-binding NF-Y subunits in Arabidopsis thaliana. Gene 264:173–185
Higo K, Ugawa Y, Iwamoto M, Higo H (1998) PLACE: a database of plant cis-acting regulatory DNA elements. Nucleic Acids Res 26:358–359
Hong WJ, Yoo YH, Park SA et al (2017) Genome-wide identification and extensive analysis of rice-endosperm preferred genes using reference expression database. J Plant Biol 60:249–258
Karppinen SM, Heikkinen K, Rapakko K, Winqvist R (2004) Mutation screening of the BARD1 gene: evidence for involvement of the Cys557Ser allele in hereditary susceptibility to breast cancer. J Med Genet 41:1–5
Kim SR, Lee DY, Il Yang J et al (2009) Cloning vectors for rice. J Plant Biol 52:73–78
Krishnan S, Dayanandan P (2003) Structural and histochemical studies on grain-filling in the caryopsis of rice (Oryza sativa L.). J Biosci 28:455–469
Laloum T, De Mita S, Gamas P et al (2013) CCAAT-box binding transcription factors in plants: y so many? Trends Plant Sci 18:157–166
Lee SK, Hwang SK, Han M et al (2007) Identification of the ADP-glucose pyrophosphorylase isoforms essential for starch synthesis in the leaf and seed endosperm of rice (Oryza sativa L.). Plant Mol Biol 65:531–546
Lee I, Seo Y-S, Coltrane D et al (2011) Genetic dissection of the biotic stress response using a genome-scale gene network for rice. Proc Natl Acad Sci 108:18548–18553
Li G, Li Q, Xing Y et al (2016) Duplication of OsHAP family genes and their association with heading date in rice. J Exp Bot 67:1759–1768
Lotan T, Ohto M, Yee KM et al (1998) Arabidopsis LEAFY COTYLEDON1 is sufficient to induce embryo development in vegetative cells. Cell 93:1195–1205
Maity SN, De Crombrugghe B (1998) Role of the CCAAT-binding protein CBF/NF-Y in transcription. Trends Biochem Sci 23:174–178
Miyoshi K, Ito Y, Serizawa A, Kurata N (2003) OsHAP3 genes regulate chloroplast biogenesis in rice. Plant J 36:532–540
Nguyen VNT, Moon S, Jung KH (2014) Genome-wide expression analysis of rice ABC transporter family across spatio-temporal samples and in response to abiotic stresses. J Plant Physiol 171:1276–1288
Ogawa M (2003) Gibberellin biosynthesis and response during arabidopsis seed germination. Plant Cell Online 15:1591–1604
Pavletich NP (1999) Mechanisms of cyclin-dependent kinase regulation: structures of Cdks, their cyclin activators, and Cip and INK4 inhibitors. J Mol Biol 287:821–828
Qu LQ, Wei XL, Satoh H et al (2003) Biochemical and molecular characterization of a rice glutelin allele for the GluA-1 gene. Theor Appl Genet 107:20–25
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Sutoh K, Yamauchi D (2003) Two cis-acting elements necessary and sufficient for gibberellin-upregulated proteinase expression in rice seeds. Plant J 34:635–645
Thirumurugan T, Ito Y, Kubo T et al (2008) Identification, characterization and interaction of HAP family genes in rice. Mol Genet Genomics 279:279–289
Wakasa Y, Takaiwa F (2013) The use of rice seeds to produce human pharmaceuticals for oral therapy. Biotechnol J 8:1133–1143
Warpeha KM, Upadhyay S, Yeh J et al (2007) The GCR31, GPA1, PRN1, NF-Y signal chain mediates both blue light and abscisic acid responses in arabidopsis. Plant Physiol 143:1590–1600
Washida H, Wu CY, Suzuki A et al (1999) Identification of cis-regulatory elements required for endosperm expression of the rice storage protein glutelin gene GluB-1. Plant Mol Biol 40:1–12
Wu CY, Washida H, Onodera Y et al (2000) Quantitative nature of the prolamin-box, ACGT and AACA motifs in a rice glutelin gene promoter: minimal cis-element requirements for endosperm-specific gene expression. Plant J 23:415–421
Xing Q, Zheng Z, Zhou X et al (2015) Ds9 was isolated encoding as OsHAP3H and its C-terminus was required for interaction with HAP2 and HAP5. J Plant Biol 58:26–37
Xu J-J, Zhang X-F, Xue H-W (2016) Rice aleurone layer specific OsNF-YB1 regulates grain filling and endosperm development by interacting with an ERF transcription factor. J Exp Bot 67:6399–6411
Yang W, Lu Z, Xiong Y, Yao J (2017) Genome-wide identification and co-expression network analysis of the OsNF-Y gene family in rice. Crop J 5:21–31
Acknowledgements
This work was supported by a grant from Kyung Hee University (20151928).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11816_2019_545_MOESM1_ESM.tif
Supplemental material 1: Figure S1. The magnified image of aleurone layer with GUS expression under HAP3K promoter. (TIFF 7725 kb)
Rights and permissions
About this article
Cite this article
Nguyen, V.N.T., Gho, YS., An, G. et al. Identification of a module of HAP transcription factors for seed development in rice. Plant Biotechnol Rep 13, 389–397 (2019). https://doi.org/10.1007/s11816-019-00545-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-019-00545-0