Abstract
The Proteaceae, a diverse family of woody flowering plants in the Southern Hemisphere, contains many species known to be susceptible to Phytophthora cinnamomi, both in the natural environment and in cut-flower orchards. Very little is known about the prevalence of P. cinnamomi and other oomycetes across these landscapes. To address this knowledge gap, we used a double ITS1 and RPS10 gene metabarcoding approach and traditional isolation protocols to investigate oomycetes in orchards and natural stands of Proteaceae across South Africa, South Africa (eastern and western), Australia, and Europe. The RPS10 primers amplified more samples, including various Pythium species, while the ITS primers detected more Phytophthora phylotypes. Both datasets showed that geographic regions influenced oomycete species richness and community composition, while they did not show any variation between orchards and natural vegetation. RPS10 metabarcoding detected the largest number of species and provided greater statistical confidence than ITS1 when considering oomycete species composition. Metabarcoding also showed that orchards had a higher abundance of P. cinnamomi compared to native stands, although this was not found when isolating through baiting of roots and rhizosphere soil. Direct isolation and metabarcoding are complementary, with metabarcoding serving as an early detection tool. However, it cannot distinguish living viable propagules from residual DNA of dead propagules, limiting its use for diagnostic purposes related to Phytophthora management and control. These results, along with those of other recent studies, show that metabarcoding offers an effective tool to describe the dynamics of soil oomycetes in different ecosystems.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Proteaceae is a large family of woody flowering plants native to the Southern Hemisphere (Weston and Barker 2006). These plants are remarkably diverse in regions with soils poor in nutrients and Mediterranean climates (Lambers et al. 2011) with approximately 1000 and 370 native species in Western Australia and South Africa, respectively (https://florabase.dpaw.wa.gov.au/science/key/proteaceae/, Crous et al. 2003; Sauquet et al. 2009). Additionally, numerous varieties and species hybrids are cultivated for cut flower production in many Mediterranean regions of Europe, Oceania, and Africa (https://www.proteaatlas.org.za/).
Despite their ecological and commercial importance, studies on the impact of oomycetes on Proteaceae are scarce and limited to the genus Phytophthora. South African literature is limited. It focuses on Phytophthora distribution in natural ecosystems and host response in productive plantations (Von Broembsen 1984a, b; Von Broembsen and Kruger 1985; Von Broembsen and Deacon 1997), and, with the exception of a 2023 survey on the critically endangered Fynbos Proteaceae species Sorocephalus imbricatus (Paap et al. 2023), it is not recent. In contrast, a greater number of studies have been conducted in Australia, including reports on cultivated Proteaceae (Hardy and Sivasithamparam 1988; Boersma et al. 2000), endemic Proteaceae (Shearer and Dillon 1996; Wills 1993) and investigations on native hyper-diverse communities of local shrubland ecosystems dominated by Banksia and Hakea (Rea et al. 2011; Jung et al. 2011; Burgess et al. 2018, 2021a; Scott et al. 2009). Outside their native range, research reports limited to the diagnosis of Phytophthora have been conducted on Proteaceae imported for production purposes in Portugal (Madeira Island) and Hawaii (Rodrigues and Moura 2000; Nakao et al. 2002).
Studies on Phytophthora in soil samples have traditionally relied on baiting-based isolation techniques for samples from all environments and host associations. However, in the past few years, High-throughput sequencing of amplicons (metabarcoding) from environmental DNA (eDNA) has become the primary tool for studying Phytophthora community composition, ecology, and spread (Khaliq et al. 2019; Burgess et al. 2022). For example, large-scale surveys have explored Phytophthora biodiversity and distribution across Australia (Burgess et al. 2017b), a Quercus ilex forest in eastern Spain (Català et al. 2017), gardens and amenity woodland sites in Great Britain (Riddell et al. 2019; Green et al. 2021), as well as a rain forest in French Guiana (Legeay et al. 2020). Metabarcoding also has been successfully employed to investigate patterns of diversity and community assembly of Phytophthora in soil samples (Bose et al. 2018) and soil and root samples (Bose et al. 2021) of different vegetation types and across geo-climatic gradients (Redondo et al. 2018).
Most Phytophthora metabarcoding studies have used ITS1 Phytophthora-specific primers (Scibetta et al. 2012) as implemented for a metabarcoding protocol by Català et al. (2015). To extend the analysis to the whole soil-rhizosphere oomycete communities, alternative barcoding regions such as the coxI and coxII mitochondrial genes can be used (Sapp et al. 2019; Landa et al. 2021; Maciá-Vicente et al. 2020). In this regard, a new method was recently validated by Foster et al. (2022) using the mitochondrial RPS10 gene (Martin et al. 2014). The advantages and disadvantages of the different primers used for metabarcoding with oomycetes have recently been reviewed by Burgess et al. (2022).
While implementing accurate metabarcoding protocols, oomycetes research also makes extensive use of traditional isolation techniques such as baiting (La Spada et al. 2022). Isolates are required for the diagnosis and taxonomic classification of new taxa (Abad et al. 2023; Pérez-Sierra et al. 2022). However, certain limitations may arise when using both techniques to analyze the same samples, raising the problem of simultaneous use of the two approaches in the same investigation. Metabarcoding, focused on Phytophthora diversity, frequently reveals additional species compared to those detected with baiting (Sarker et al. 2023a). Furthermore, in some recent works, the two techniques have provided non-comparable, discordant results (Bose et al. 2018; Khaliq et al. 2018, 2021; Landa et al. 2021).
This study aimed to compare the oomycetes community composition in the roots and rhizosphere soil of Proteaceae across different vegetative and geographical regions. The study included orchards in South Africa, Australia, Europe (Portugal and Italy) and natural bush areas in South Africa and Australia. This comparison was achieved using both direct isolations and ITS1 and RPS10 metabarcoding methods. Specifically, we investigated i) whether oomycete species richness and composition differ between orchards and natural stands, ii) whether there are differences among the countries, and iii) whether soil baiting and metabarcoding produced comparable results for estimating P. cinnamomi distribution in Proteaceae orchards versus natural stands of these plants.
Materials and methods
Sites sampled
Ornamental Proteaceae orchards were selected in thirteen different locations across four countries. The sites were grouped in four different geographic areas, South Africa east (one in Gauteng, one in Kwa-Zulu Natal, two in Eastern Cape), South Africa west (four in Western Cape), Australia (three in Western Australia), and Europe (one in Italy and one in Portugal). Sampling was conducted between June 2016 and April 2017, during the season with local higher rain occurrence (summer in South Africa East, late spring in South Africa West, late autumn in Australia, mid-spring in Europe (Fig. 1A and Table S1).
Maps of the sampling locations in South Africa (east: 1 in Gauteng, 2 in Kwa-Zulu Natal, 3 and 4 in Eastern Cape; west: 5–8 in Western Cape), Australia (9–11) and Europe (12 in Portugal and 13 in Italy) and scheme of the sampling design in the left down corner of the image: four sampling plots per location consisting of a 50–60 m transect with five sampling units (the circles) equidistant along the transect for a total of twenty sampling units; two transects were realized in natural ecosystems vegetation out of the farm (A) and two inside the farm in planted Proteaceae (B); each sampling unit contained 5 plants
Four sampling plots per location consisted of a 50–60 m transect with five sampling units equidistant along the transect, each consisting of 5 plants (4 sampling plots × 5 sampling units, 20 sampling units per location; Fig. 1B). Two sampling plots were surveyed and sampled within Proteaceae orchards, one planted with the cultivar “pink ice” (Protea compacta × Protea susannae P Matthews), and one planted with the cultivar “safari sunset” (Leucadendron salignum × Leucadendron laureolum; J Stevens and I Bell). Two additional plots were surveyed and sampled outside the cultivated orchards in natural vegetation containing Proteaceae vegetation. However, in Europe, where natural Proteaceae do not occur, sampling outside the orchards was conducted on native vegetation close to the plantation borders in Portugal. In Italy, where Proteaceae orchards were not present, samples were collected from a nursery located in a large area cultivated for intensive agricultural and nursery production, far from natural ecosystems. As a result, samples from native vegetation were not included in the collection from Italy.
Processing of samples
From each sampling unit, the under-canopy surface litter of each of the five plants was scrapped away on a 20 × 20 cm spot. At a depth of 15 cm, a soil sample of approximately 100 g including roots from the target plant was taken from each hole and mixed in a plastic bag to make a 500-g bulk sample. The bags were kept open to prevent condensation. On the same day, approximately 5 g of cluster roots was picked out of this bulk sample and allowed to dry naturally in paper bags for 7 days before being transferred to 2-mL microcentrifuge tubes (Pirouet) and stored at – 80 °C. The soil bound with the roots was not actively removed, although some fell away during the drying process. Consequently, eDNA was extracted from roots and attached to rhizosphere soil. These samples were then subjected to metabarcoding to detect the presence of oomycete species, while the remaining bulk samples were processed by baiting to isolate Phytophthora species. Sampling tools were cleaned between each sampling unit with a brush using water and commercial dishwashing detergent to remove all soil residues and then sprayed with 70% ethanol.
DNA extraction
Approximately 200 mg of each root sample was ground with a Qiagen TissueLyser attached to a grinding jar (Qiagen-manufactured by Retch) for 4 min at 24 Hz using two sterile 0.5-mm iron balls. Approximately 50 µg of each ground root sample was transferred into the extraction kit tubes, and the total genomic DNA was extracted using the Mo Bio Laboratories, Power Plant® Pro DNA Isolation Kit (Cat# 13400–50) following the manufacturer’s protocol. A total of 273 samples were extracted, and the DNA was stored at – 20 °C.
Library preparation
Separate Illumina metabarcoding runs were conducted using two primer sets. The ITS1 gene region was amplified with Phytophthora-specific primers. A nested PCR approach was used with Phytophthora-specific primers 18h2f and 5.8RBis (Scibetta et al. 2012) in the first round, and nested primers ITS6 and 5.8S-IR (Cooke et al. 2000; Scibetta et al. 2012) in the second round, following the methods of Scibetta et al. (2012) and Català et al. (2015). The 40S ribosomal protein S10 (RPS10) gene region was amplified with oomycete-specific primers (Foster et al. 2022). A nested approach was employed with PRV9-F and PRV9-R primers in the first round (Martin and Coffey 2012) and oomycete-specific primers RPS10-F and RPS10-R in the second round (http://oomycetedb.cgrb.oregonstate.edu/). In both cases, the primers used in the second round had Illumina MiSeq adapter sequences attached to the 5′ end, following standard protocols for the MiSeq platform (Illumina Demonstrated Protocols: Metagenomic Sequencing Library Preparation).
Each 25 μL PCR reaction included 12.5 μL of PCR buffer KAPA HiFi HotStart ReadyMix (KAPA Biosystems), 8 μL of PCR grade water, 1 μM of each primer (1 µL), and 2.5 μL of genomic DNA (for the first round) or 1 μL of the PCR product (for the second round). No-template negative PCR controls were included each time a PCR reaction was set up and carried forward to the second round in the same manner as for the samples. PCR cycling conditions were 94 °C for 2 min, 30 cycles of 95 °C for 20 s, 60 °C for 25 s, and 72 °C for 1 min followed by a final step at 72 °C for 7 min and holding at 4 °C. The products were discarded if a band was visualized in the negative PCR controls.
First-round PCR was conducted in duplicate and combined after the second-round PCR products based on the intensity of bands on 2% agarose gel electrophoresis. Biological replicates were assigned the same barcode and combined before the Illumina run. The PCR products were then purified with Agencourt AMPure XP magnetic beads (Beckman Coulter) and uniquely barcoded. These samples were again purified with Agencourt AMPure XP magnetic beads and pooled (separately for each gene region) based on DNA concentrations as determined using Qbit kits (Invitrogen Qubit TM ds DNA HS Assay Kit). Following the manufacturer’s recommendations, the two uniquely indexed libraries were sequenced at the Murdoch University laboratory facilities on an Illumina MiSeq using 500-cycle V2 chemistry (250 bp paired-end reads).
Bioinformatic analysis
Paired-end reads were merged using USEARCH v10 (Edgar 2010) with a minimum overlap length of 50 bp and no gaps were allowed in the merged alignments. Sequence deconvolution, including quality control and clustering, was also done using USEARCH v10 (Edgar 2010). Specifically, sequences less than 200 bp and low mean quality (< 20) were removed. The remaining sequences that passed quality control were clustered into amplicon sequence variants (ASVs).
The subsequent analysis was conducted in Geneious ver. R10 (https://www.geneious.com/) to assign ASVs to a species based on phylogenetic inference. Consensus sequences were aligned using the MAFFT alignment within Geneious using the default parameters. Identities of the ASVs were initially assigned by conducting an internal BLAST search against a customized reference database. For the ITS gene region this database included sequences of type isolates as identified in IDPhy (Abad et al. 2023) and consisted of 300 Phytophthora species and undescribed (but designated) taxa and Phytophthora phylotypes recognized through metabarcoding from other previous studies (Burgess et al. 2017b, a; Bose et al. 2018). For the RPS10 gene region, the reference database was downloaded from OomyceteDB (Foster et al. 2022) and consisted of 351 sequences of oomycetes, predominantly Phytophthora (144 species) and Pythium (133 species).
For the next step, all ASVs were then separated into phylogenetic clades, and phylogenetic analyses were conducted using Geneious tree builder and verified sequences of all known Phytophthora species within each clade. Several ASVs often clustered together in strongly supported terminal clades. These final identities were considered to be phylotypes. A phylotype was considered to represent a putative new species if it did not match any known species in the phylogenetic analysis. For Phytophthora species, these phylotypes were assigned a number representing the phylogenetic clade and a letter to distinguish between putative new phylotypes in the same clade. New phylotypes from unknown clades were allocated an “X.”
Baiting and identification of isolates
Baiting Phytophthora spp. from soil samples was carried out at 20–24 °C using young leaves from Hedera helix, Quercus ilex, Quercus robur, cotyledons of Eucalyptus sieberi, and petals of ornamental roses following best practice protocols (Burgess et al. 2021b). After 3–10 days the portion of infected baits was first placed on NARPH medium (Hüberli et al. 2000) as described by Simamora et al. (2018) and then plated onto cornmeal agar (17 g/L water of corn meal agar) to examine for the presence of Phytophthora structures using a dissection microscope. Oomycete-like cultures were selected based on their morphology and transferred to carrot agar (5 g/L water of corn meal agar, 400 g/L water of fresh carrots). Isolates obtained were then grouped based on culture morphology.
DNA was extracted from the mycelium of 7-day-old aerial hyphae of one representative isolate for each morphological group using PrepManTM Ultra (Applied Biosystems by Life Technologies), following the manufacturer’s protocol. The nuclear ribosomal internal transcribed spacer (ITS) was amplified with primers ITS6 (Cooke et al. 2000) and ITS4 (White et al. 1990). Each 25 µL PCR reaction contained 12.5 μL 2 × GoTaq® Green Master Mix (Promega Corporation, Madison, WI, USA), 0.5 μL of each of forward and reverse primer, 1.5 μL of DNA, and 10 μL with PCR grade water. Amplification conditions were as follows: one cycle at 94 °C for 3 min., 35 cycles at 94 °C for 1 min, 55 °C for 1 min, 72 °C for 1 min, and a final step at 72 °C for 1 min. Five microliters of each PCR product was separated by electrophoresis in 1% agarose gels in 5% TAE buffer (40 mM Tris, 40 mM acetate, 2 mM EDTA, pH 8.0). The amplified PCR fragments were purified through a Sephadex spin column (Sigma, Steinheim, Germany) and sequenced with the BigDye terminator cycle sequencing kit (Perkin-Elmer Applied Biosystems, Foster City, CA, USA). Products were separated with an ABI 3730 48 capillary sequencer (Perkin-Elmer Applied Biosystems).
Identities were assigned to isolates in Geneious by conducting an internal BLAST search against the same customized reference database described above, which includes sequences of type isolates of all described Phytophthora species as identified in IDPhy (Abad et al. 2023). All obtained sequences were submitted to GenBank, and accession numbers are provided in Table S2.
Statistical analysis
Venn diagrams were used to visualize the number of oomycete taxa shared between (1) the vegetation types (orchards and natural stands) and (2) the different geographic areas. The “VennDiagram” package in the R software (R Core Team 2021) was used to construct the Venn diagrams. Separate Venn diagrams were constructed for data obtained from barcoding (ITS1 and RPS10) gene regions and baiting.
To analyze oomycete species richness, the number of taxa per sample was calculated. The effects of the vegetation type, the different geographic areas, and their interaction (V × G) on species richness were analyzed with a generalized linear model (GLM). The model was fitted assuming a Poisson error distribution and a log-link function. Additionally, the natural logarithm of the total number of reads was included as an offset to consider the variation in fungal richness triggered by sequencing depth. The “vegan” package in R software (R Core Team 2021) was employed for analyzing oomycete species richness using data obtained from ITS and RPS10 gene regions.
For analyzing the oomycete taxa community composition in different vegetation types and geographic areas, a principal coordinate analysis (PCoA) was performed using Jaccard distance. Differences in oomycetes community composition between vegetation types and/or geographic areas were assessed using permutational multivariate variance analysis (PERMANOVA). The explanatory variables used for PERMANOVA were the vegetation type, the geographic areas, and their interaction. The natural logarithm of Proteaceae sample sequencing depth was included as the first effect in the PERMANOVA to consider the variation in fungal composition triggered by sequencing depth. The “vegan” package in R software (R Core Team 2021) was utilized to analyze oomycete taxa community composition using data obtained from ITS and RPS10 gene regions.
Phytophthora cinnamomi was the only species routinely detected through baiting, and thus, the baiting and metabarcoding results for P. cinnamomi were compared. To determine if P. cinnamomi showed a different abundance between vegetation types, a GLM was fitted, assuming a Poisson error distribution and a log link function. The natural logarithm of the total number of positive identifications (baiting) or natural logarithm of sequencing depth (ITS and RPS10) was included as an offset to account for differences in sample size. The “vegan” package in R software (R Core Team 2021) was used to analyze P. cinnamomi abundance using data obtained from ITS, RPS10 gene regions and baiting to check for differences between the detection methods.
Results
Taxa identified through metabarcoding
ITS sequencing
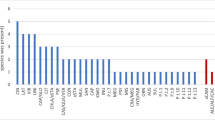
Of the 257 samples, 36% (92) produced an amplification product. The 26,597 reads that passed quality control were clustered into 131 ASVs, of which all were classified as oomycetes; 45% of the 59 ASVs (60% of the reads) were identified as Phytophthora. On average 633 reads were found for each sample. Phylogenetic identification revealed 17 phylotypes, 9 of which were Phytophthora taxa residing in five of the eleven clades found within the Phytophthora phylogeny (Table S3 and Fig. S1). The highest number of reads was observed for P. cinnamomi, followed by Phytophthora clade 12A, P. nicotianae, P. multivora, P. pachypleura, P. cactorum, P. pseudocryptogea, P. AUS XA, and P. cryptogea. The other 10 phylotypes accounted for 40% of total reads and were assigned to Plasmopara halstedii, Pythium aff. heterothallicum, Pythium rostratifingens, Bremia sp., Lagena sp., Myzocytiopsis subuliformis, Peronospora aff. trivialis, and Peronospora effusa.
Among the 17 phylotypes, five were shared between the two vegetation types and 10 were exclusively found in root samples collected in orchards (Fig. 2a). None of these 17 phylotypes were shared among all areas, 5 were shared between two areas, 7 were found only in Australia, 3 only in South Africa, and 2 only in Europe (Fig. 2b and Appendix S1).
RPS10 sequencing
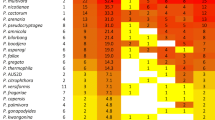
Out of the 257 samples, 64% (164) produced an amplification product. The 139,602 reads that passed quality control were clustered into 562 ASVs, of which 8% were Phytophthora (30% of the reads) and 92% were other oomycetes (70% of the reads). On average 3324 reads were found for each sample. Among the 65 phylotypes detected, 5 were matched with Phytophthora species; Phytophthora clade 12A, P. cactorum, P. cryptogea, P. sojae, and P. cinnamomi (Table S3). A total of 18 Pythium spp. were identified to the species level (28% of the reads) (Table S3 and Fig. S2); Pythium camurandrum, Py. cederbergense, Py. conidiophorum, Py. contiguanum, Py. cryptoirregulare, Py. dissotocum, Py. folliculosum, Py. irregulare, Py. mamillatum, Py. mastophorum, Py. myriotylum, Py. nunn, Py. oopapillium, Py. ornamentatum, Py. ornacarpon, Py. rostratifingens, Py. vanterpoolii, and Py. volutum. An additional 31 Pythium spp. were detected, of which 28 were placed into a clade, while one Bremia sp. and one Phytopythium sp. were identified to the genus level. The remaining 9 oomycete phylotypes could not be matched with a genus.
Of the 65 phylotypes, 28 were shared in the root samples of the two vegetation types, 34 were exclusively found in samples from orchards, and only 3 were exclusively in samples from native vegetation (Fig. 2c). Moreover, of the 65 phylotypes, the majority (29) were found exclusively in Australia, 5 were exclusively found in South Africa, and 5 in Europe (Fig. 2d and Appendix S1).
Taxa identified using baiting
Phytophthora spp. were recovered from 48 (19%) sampling units of which 33 were from South Africa (21 in native vegetation and 12 in orchards), 8 in Portugal (3 in native vegetation and 5 in orchards), 4 in Italy (orchards only), and three in Australia (orchards only) (Table S4). These sampling units matched five species, three of which were shared between the two vegetation types (Fig. 2e), and one was exclusively found in South African and Australian samples (Fig. 2f). Phytophthora cinnamomi was the most recovered species (isolated in 40 sampling units), followed by P. cryptogea and P. ornamentata (isolated in 3 sampling units), P. niederhauserii (isolated in 1 sampling unit), and P. multivora (isolated in 1 sampling unit) (Table S4). Genbank sequence number of Phytophthora isolates and sampling details are reported in Table S2.
Oomycete species richness and community composition
Oomycete species richness did not vary between orchards and natural vegetation. However, it was significantly influenced by geographic areas and the interaction between vegetation types and geographic areas (Table 1). Specifically, Proteaceae samples from South Africa showed the lowest species richness (mean ± SE: 3.6 ± 1.1 and 1.4 ± 0.2, RPS10 and ITS1, respectively, Fig. 3). While species richness did not vary between environment or geographic area, the PCoA plot illustrates a difference in oomycetes community composition between Proteaceae samples from Australia and South Africa (Fig. 3). PERMANOVA confirmed that the geographic area influenced oomycetes community composition (Table 2).
Comparison of P. cinnamomi detection through baiting and metabarcoding
Phytophthora cinnamomi was the only species consistently isolated with baiting: 31 positive sampling units in South Africa, of which 20 were from native vegetation and 11 in orchards; 7 in Europe of which one was native vegetation and 6 in orchards and 2 in Australian orchards (Tables S3 and S4). Data analysis did not show a significant difference in P. cinnamomi abundance between vegetation types using baiting (Table 3). However, P. cinnamomi abundance significantly differed between vegetation types according to datasets obtained with RPS10 and ITS1 metabarcoding approaches (Fig. 4 and Table 3). Specifically, P. cinnamomi abundance was significantly higher in orchards compared to natural stands.
Discussion
The metabarcoding approach used in this study revealed a significant interaction between the oomycetes richness of different vegetation types and on different continents. Most detected Phytophthora taxa were well-known pathogens with global distributions such as P. cinnamomi, P. cactorum, P. cryptogea, P. multivora, P. pseudocryptagea, and P. sojae. The oomycete-specific RPS10 primers amplified 164 out of 257 samples, while the Phytopthora-specific ITS1 primers amplified only 92 out of 257 samples. As expected, more phylotypes were detected using the RPS10 primers, including many Pythium species. However, more Phytophthora phylotypes were detected using the ITS primers. Despite these differences, both datasets demonstrated that the geographic region influenced oomycetes community composition.
Consistent with our results, other recent research has also tested different metabarcoding techniques on the same environmental samples and mock communities, demonstrating that multiple primer pairs for the same target gene (Legeay et al. 2020) or different genes (Landa et al. 2021; Burgess et al. 2022) provide partially divergent data. When taxonomic coverage is the principal aim, sequencing multiple regions may be advisable and provide complementary data. For example, Landa et al. (2021) found the COI region detected other oomycete taxa in addition to the Phytophthora and better differentiated several Phytophthora spp. from clades that were difficult to resolve using the ITS region. In our study, although the two gene regions amplified did not produce statistically significant differences in species richness between environments, the RPS10 region gave a greater coverage, amplifying other Peronosporaceae in addition to Phytophthora.
Oomycetes association with Proteaceae in orchards versus natural stands
The oomycete species richness and community composition found in this study were not significantly different in cultivated orchards and natural Proteaceae environments. Although there is little information on this topic, recent studies across various environments and comparing various levels of disturbance have shown that less disturbed environments have fewer Phytophthora species (Dale et al. 2022; Redondo et al. 2018). Bose et al. (2018) found that vegetation type significantly influenced the Phytophthora community composition in soils. However, their study is not directly comparable to ours because they sampled different plant species from each of the different environments. By focusing solely on Proteaceae, we were able to demonstrate that the host had a greater impact on oomycetes diversity than the environment.
Phytophthora cinnamomi was more abundant in the sampled orchards than in the natural environment. Globally, P. cinnamomi is known as the most important pathogen of Proteaceae and other tree crops (Burgess et al. 2017a) and is often the focus of control strategies in both forestry and agriculture (Hardham and Blackman 2018), where the constant human activity contributes to its spread. We believe that the weekly use of machinery for the chemical control of pests together with a regular movement of flower pickers in Proteaceae orchards may have been crucial for the higher presence of this pathogen in comparison to the less disturbed natural stands sampled in this work.
Oomycetes association with Proteaceae in different countries
The metabarcoding approach demonstrated that oomycete species richness and community composition differed in the samples collected across countries and on different continents. While numerous studies of oomycete communities have been conducted (summarized in Burgess et al. 2022), to the best of our knowledge, our work was the only one that focused on the difference of soil-borne pathogens across continents. These results represent a significant practical application for this molecular tool, which is currently underutilized in large-scale inter-country investigations of plant-associated microorganisms (Tedersoo et al. 2019). Of the few examples is the recent work by Franić et al. (2023) about the global distribution of twig fungal endophytes in trees from botanical gardens and arboreta, and two studies of fugal endophytes in twigs of ornamental plants (Migliorini et al. 2021) and on commercial seeds imported from overseas into Europe (Franić et al. 2019).
Therefore, our work advocates for the further development and implementation of the metabarcoding approach to study the global movement of plant pathogenic soil-borne oomycetes. These oomycetes are highly prevalent in the soil of potted plants traded from overseas into Europe (Eschen et al. 2015; Migliorini et al. 2015) and are responsible for causing significant ecological and economic damages (Santini et al. 2013).
Comparison of metabarcoding molecular detection methods and traditional baiting
Despite metabarcoding showing that P. cinnamomi was more common in orchards than in natural vegetation, isolation rates did not reflect this difference. The difference in several species detected through direct isolation (by soil/root baiting) versus metabarcoding has been a subject of long-standing debate (Sarker et al. 2023a). In soil/root baiting, the isolation outcomes depend on the presence of active Phytophthora propagules in the analyzed samples since the method relies on motile zoospores released from the sporangia being attracted to a live bait (Sarker et al. 2021). Baiting conditions favor fast-sporulating species, able to infect a range of bait species, achieve infection with a low number of zoospores, and grow rapidly on selective agar (Sarker et al. 2023a). For this reason, baiting success depends upon the species present and the viability of the propagules when the sample was collected (Sarker et al. 2021, 2023b). In our study, P. cinnamomi proved to be the most active species in our sampling areas and was therefore easily isolated.
The two methods, direct isolation and metabarcoding, when applied to the study of soil oomycetes, should be considered complementary, and results interpreted with care depending upon the study system and the aim of the investigation. Metabarcoding can be used as an early detection tool but cannot distinguish living viable propagules from residual DNA of dead propagules (false positive) (Sanzani et al. 2014; Kunadiya et al. 2021). The accurate overview of oomycetes taxonomy provided is fundamental for conducting diversity studies aimed at comparing different situations included in the sampling design, but it is unable to report the state of things at the exact moment of collection. This practical limit is acceptable for conducting diversity studies where the time range is generally not considered variable. However, metabarcoding or similar molecular methods should not be used for diagnostic purposes if the aim is the management and control of Phytophthora.
Conclusion
Our study provides a comprehensive analysis of the distribution and diversity of oomycete species in the soil and roots of cultivated and naturally growing Proteaceae across different continents. The prevalence of widely recorded Phytophthora species in various environments underscores the risks posed by the spread of this pathogen in both Proteaceae orchards and natural ecosystems.
The consistent presence of P. cinnamomi, especially in orchards, highlights its negative economic impact on the production sector and raises concerns about the considerable environmental risk to South African and Australian native ecosystems, in line with the recent report by Paap et al. (2023). The high isolation frequency of P. cinnamomi in our investigation using baiting demonstrates how propagules of this species are actively spreading in farms, emphasizing the importance for managers to establish good hygiene practices to prevent further transfer within the site and to the surrounding environment.
Furthermore, our work confirms the reliability of metabarcoding as a taxonomical diagnostic tool, marking the first-time utilization of this approach to assess oomycetes associated with Proteaceae. Additionally, the successful outcomes achieved using the RPS10 gene underscore its efficiency with soil environmental DNA samples, further strengthening the applicability of this approach for ecological studies of oomycete communities.
Data availability
The ITS gene dataset generated and analyzed during the current study is available in the GenBank repository, https://www.ncbi.nlm.nih.gov/genbank/. The metabarcoding dataset generated and analyzed during the current study are available from the corresponding author on reasonable request.
References
Abad ZG, Burgess TI, Redford AJ, Bienapfl JC, Srivastava S, Mathew R, Jennings K (2023) IDphy: An international online resource for molecular and morphological identification of Phytophthora. Plant Dis 107(4):987–998. https://doi.org/10.1094/PDIS-02-22-0448-FE
Boersma JG, Cooke DEL, Sivasithamparam K (2000) A survey of wildflower farms in the south-west of Western Australia for Phytophthora spp. associated with root rots. Aust J Exp Agric 40(7):1011–1019. https://doi.org/10.1071/EA00013
Bose T, Wingfield MJ, Roux J, Vivas M, Burgess TI (2018) Community composition and distribution of Phytophthora species across adjacent native and non-native forests of South Africa. Fungal Ecol 36:17–25. https://doi.org/10.1016/j.funeco.2018.09.001
Bose T, Wingfield MJ, Roux J, Vivas M, Burgess TI (2021) Phytophthora species associated with roots of native and non-native trees in natural and managed forests. Microb Ecol 81:122–133. https://doi.org/10.1007/s00248-020-01563-0
Burgess TI, Scott JK, Mcdougall KL, Stukely MJ, Crane C, Dunstan WA, Brigg F, Andjic V, White D, Rudman T, Arentz F, Hardy GESJ (2017a) Current and projected global distribution of Phytophthora cinnamomi, one of the world’s worst plant pathogens. Glob Change Biol 23(4):1661–1674. https://doi.org/10.1111/gcb.13492
Burgess TI, White D, McDougall KM, Garnas J, Dunstan WA, Català S, Carnegie AJ, Worboys S, Cahill D, Vettraino AM, Stukely MJ, Hardy GESJ (2017b) Distribution and diversity of Phytophthora across Australia. Pac Conserv Biol 23(2):150–162. https://doi.org/10.1071/PC16032
Burgess TI, Simamora AV, White D, Wiliams B, Schwager M, Stukely MJC, Hardy GESJ (2018) New species from Phytophthora Clade 6a: evidence for recent radiation. Persoonia 41(1):1–17. https://doi.org/10.3767/persoonia.2018.41.01
Burgess TI, Edwards J, Drenth A, Massenbauer T, Cunnington J, Mostowfizadeh-Ghalamfarsa R, Dinh Q, Liew ECY, White D, Scott P, Barber PA, Tan YP (2021a) Current status of Phytophthora in Australia. Persoonia 47(1):151–177. https://doi.org/10.3767/persoonia.2021.47.05
Burgess TI, López-Villamor A, Paap T, Williams B, Belhaj R, Crone M, Dunstan WA, Howard K, Hardy GESJ (2021b) Towards a best practice methodology for the detection of Phytophthora species in soils. Plant Pathol 70(3):604–614. https://doi.org/10.1111/ppa.13312
Burgess TI, White D, Sapsford SJ (2022) Comparison of primers for the detection of Phytophthora (and other oomycetes) from environmental samples. J Fungi 8(9):980. https://doi.org/10.3390/jof8090980
Català S, Perez-Sierra A, Abad-Campos P (2015) The use of genus-specific amplicon pyrosequencing to assess Phytophthora species diversity using eDNA from soil and water in northern Spain. PLoS ONE 10(3):e0119311. https://doi.org/10.1371/journal.pone.0119311
Català S, Berbegal M, Pérez-Sierra A, Abad-Campos P (2017) Metabarcoding and development of new real-time specific assays reveal Phytophthora species diversity in holm oak forests in eastern Spain. Plant Pathol 66(1):115–123. https://doi.org/10.1111/ppa.12541
Cooke DEL, Drenth A, Duncan JM, Wagels G, Brasier CM (2000) A molecular phylogeny of Phytophthora and related oomycetes. Fungal Genet Biol 30(1):17–32. https://doi.org/10.1006/fgbi.2000.1202
Crous PW, Denman S, Taylor JE, Swart LZ, Palm ME (2003) Cultivation and diseases of Proteaceae: Leucadendron, Leucospermum and Protea. Centraalbureau voor Schimmelcultures (CBS)
Dale AL, Feau N, Berube JA, Ponchart J, Bilodeau GJ, Hamelin RC (2022) Urban environments harbor greater oomycete and Phytophthora diversity, creating a bridgehead for potential new pathogens to natural ecosystems. Environmental DNA 4(5):1039–1051. https://doi.org/10.1002/edn3.300
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19):2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Eschen R, Britton K, Brockerhoff E, Burgess TI, Dalley V, Epanchin-Niell RS, Gupta K, Hardy GESJ, Huang Y, Kenis M, Kimani E, Li HM, Olsen S, Ormrod R, Otieno W, Sadof C, Tadeu E, Theyse M (2015) International variation in phytosanitary legislation and regulations governing importation of plants for planting. Environ Sci Policy 51:228–237. https://doi.org/10.1016/j.envsci.2015.04.021
Foster ZS, Albornoz FE, Fieland VJ, Larsen MM, Jones FA, Tyler BM, Nguyen HD, Burgess TI, Riddell C, Voglmayr H, Martin FN, Günwald NJ (2022) A new oomycete metabarcoding method using the RPS10 gene. Phytobiomes J 6(3):214–226. https://doi.org/10.1094/PBIOMES-02-22-0009-R
Franić I, Prospero S, Hartmann M, Allan E, Auger-Rozenberg MA, Grünwald NJ, Kenis M, Roques A, Schneider S, Sniezko R, Williams W, Eschen R (2019) Are traded forest tree seeds a potential source of nonnative pests? Ecol Appl 29(7):e01971. https://doi.org/10.1002/eap.1971
Franić I, Allan E, Prospero S, Adamson K, Attorre F, Auger-Rozenberg MA, Augustin S, Avtzis D, Baert W, Barta M, Bauters K, Bellahirech A, Boroń P, Bragança H, Brestovanská T, Brurberg MB, Burgess TI, Burokienė D, Cleary M, Corley J, Coyle D, Csóka G, Černý K, Davydenko K, de Groot M, Diez J, Doğmuş Lehtijärvi HT, Drenkhan R, Edwards J, Elsafy M, Eötvös C, Falko R, Fan J, Feddern N, Fürjes-Mikó Á, Gossner M, Grad B, Hartmann M, Havrdova L, Kádasi Horáková M, Hrabětová M, Justesen M, Kacprzyk M, Kenis M, Kirichenko N, Kovač M, Kramarets V, Lacković N, Lantschner M, Lazarević J, Leskiv M, Li H, Madsen CL, Malumphy C, Matošević D, Matsiakh I, May TW, Meffert J, Migliorini D, Nikolov C, O’Hanlon R, Oskay F, Paap T, Parpan T, Piškur B, Ravn HP, Richard J, Ronse A, Roques A, Ruffner B, Santini A, Sivickis K, Soliani C, Talgø V, Tomoshevich M, Uimari A, Ulyshen M, Vettraino AM, Villari C, Wang Y, Witzell J, Zlatković M, Eschen R (2023) Climate, host and geography shape insect and fungal communities of trees. Sci Rep 13(1):11570. https://doi.org/10.1038/s41598-023-36795-w
Green S, Cooke DE, Dunn M, Barwell L, Purse B, Chapman DS, Valatin G, Schlenzig A, Barbrook J, Pettitt T, Price C, Marzano M (2021) PHYTO-THREATS: addressing threats to UK forests and woodlands from Phytophthora; identifying risks of spread in trade and methods for mitigation. Forests 12(12):1617. https://doi.org/10.3390/f12121617
Hardham AR, Blackman LM (2018) Phytophthora cinnamomi. Mol Plant Pathol 19(2):260–285. https://doi.org/10.1111/mpp.12568
Hardy GESJ, Sivasithamparam K (1988) Phytophthora spp. associated with container-grown plants in nurseries in Western Australia. Plant Dis 72(5):435–437. https://doi.org/10.1094/PD-72-0435
Hüberli D, Tommerup IC, Hardy GESJ (2000) False-negative isolations or absence of lesions may cause mis-diagnosis of diseased plants infected with Phytophthora cinnamomi. Australas Plant Pathol 29:164–169. https://doi.org/10.1071/AP00029
Jung T, Stukely MJC, Hardy GESJ, White D, Paap T, Dunstan WA, Burgess TI (2011) Multiple new Phytophthora species from ITS Clade 6 associated with natural ecosystems in Australia: evolutionary and ecological implications. Persoonia 26(1):13–39. https://doi.org/10.3767/003158511X557577
Khaliq I, Hardy GESJ, White D, Burgess TI (2018) eDNA from roots: a robust tool for determining Phytophthora communities in natural ecosystems. FEMS Microbiol Ecol 94(5):fiy 048. https://doi.org/10.1093/femsec/fiy048
Khaliq I, Hardy GESJ, McDougall KL, Burgess TI (2019) Phytophthora species isolated from alpine and sub-alpine regions of Australia, including the description of two new species; Phytophthora cacuminis sp. nov and Phytophthora oreophila sp. nov. Fungal Biology 123(1):29–41. https://doi.org/10.1016/j.funbio.2018.10.006
Khaliq I, Burgess TI, Hardy GESJ, White D, McDougall KL (2021) Phytophthora and vascular plant species distributions along a steep elevation gradient. Biol Invasions 23:1443–1459. https://doi.org/10.1007/s10530-020-02450-y
Kunadiya MB, Burgess TI, Dunstan WA, White D, Hardy GESJ (2021) Persistence and degradation of Phytophthora cinnamomi DNA and RNA in different soil types. Environmental DNA 3(1):92–104. https://doi.org/10.1002/edn3.127
La Spada F, Cock PJ, Randall E, Pane A, Cooke DE, Cacciola SO (2022) DNA metabarcoding and isolation by baiting complement each other in revealing Phytophthora diversity in anthropized and natural ecosystems. J Fungi 8(4):330. https://doi.org/10.3390/jof8040330
Lambers H, Brundrett MC, Raven JA, Hopper SD (2011) Plant mineral nutrition in ancient landscapes: high plant species diversity on infertile soils is linked to functional diversity for nutritional strategies. Plant Soil 348:7–27. https://doi.org/10.1007/s11104-011-0977-6
Landa BB, Arias-Giraldo LF, Henricot B, Montes-Borrego M, Shuttleworth LA, Pérez-Sierra A (2021) Diversity of Phytophthora species detected in disturbed and undisturbed British soils using high-throughput sequencing targeting ITS rRNA and COI mtDNA regions. Forests 12(2):229. https://doi.org/10.3390/f12020229
Legeay J, Husson C, Boudier B, Louisanna E, Baraloto C, Schimann H, Marcais B, Buée M (2020) Surprising low diversity of the plant pathogen Phytophthora in Amazonian forests. Environ Microbiol 22(12):5019–5032. https://doi.org/10.1111/1462-2920.15099
Maciá-Vicente JG, Nam B, Thines M (2020) Root filtering, rather than host identity or age, determines the composition of root-associated fungi and oomycetes in three naturally co-occurring Brassicaceae. Soil Biol Biochem 146:107806. https://doi.org/10.1016/j.soilbio.2020.107806
Martin FN, Coffey MD (2012) Mitochondrial haplotype analysis for differentiation of isolates of Phytophthora cinnamomi. Phytopathology 102(2):229–239. https://doi.org/10.1094/PHYTO-04-11-0115
Martin FN, Blair JE, Coffey MD (2014) A combined mitochondrial and nuclear multilocus phylogeny of the genus Phytophthora. Fungal Genet Biol 66:19–32. https://doi.org/10.1016/j.fgb.2014.02.006
Migliorini D, Ghelardini L, Tondini E, Luchi N, Santini A (2015) The potential of symptomless potted plants for carrying invasive soilborne plant pathogens. Divers Distrib 21(10):1218–1229. https://doi.org/10.1111/ddi.12347
Migliorini D, Messal M, Santini A, Ramos AP, Talhinhas P, Wingfield MJ, Burgess TI (2021) Metabarcoding reveals southern hemisphere fungal endophytes within wood of cultivated Proteaceae in Portugal. Eur J Plant Pathol 160:173–184. https://doi.org/10.1007/s10658-021-02233-8
Nakao P, Leonhardt K, Nagata N, Shingaki P (2002, March) Screening Leucospermum hybrids for tolerance to Phytophthora cinnamomi. In: VI International Protea Research Symposium 602 :113–117. https://doi.org/10.17660/ActaHortic.2003.602.15
Paap T, Nndanduleni M, Wingfield MJ (2023) A critically endangered Proteaceae in the Cape Florisitic Region threatened by an invasive pathogen. Bothalia 53(1):1–7
Pérez-Sierra A, Jung MH, Jung T (2022) Survey and monitoring of Phytophthora species in natural ecosystems: methods for sampling, isolation, purification, storage, and pathogenicity tests. In: Plant Pathology: Method and Protocols :13–49. New York, NY: Springer US. https://doi.org/10.1007/978-1-0716-2517-0_2
R Core Team (2021) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
Rea AJ, Burgess TI, Hardy GESJ, Stukely MJC, Jung T (2011) Two novel and potentially endemic species of Phytophthora associated with episodic dieback of Kwongan vegetation in the south-west of Western Australia. Plant Pathol 60(6):1055–1068. https://doi.org/10.1111/j.1365-3059.2011.02463.x
Redondo MA, Boberg J, Stenlid J, Oliva J (2018) Functional traits associated with the establishment of introduced Phytophthora spp. Swedish forests. J Appl Ecol 55(3):1538–1552. https://doi.org/10.1111/1365-2664.13068
Riddell CE, Frederickson-Matika D, Armstrong AC, Elliot M, Forster J, Hedley PE, Morris J, Thorpe P, Cooke DE, Pritchard L, Sharp PM, Green S (2019) Metabarcoding reveals a high diversity of woody host-associated Phytophthora spp. in soils at public gardens and amenity woodlands in Britain. PeerJ 7:e6931. https://doi.org/10.7717/peerj.6931
Rodrigues PF, Moura MF (2000, April) Fungal diseases on proteas identified in Madeira Island. In: V International Protea Research Symposium 545 :265–268. https://doi.org/10.17660/ActaHortic.2001.545.34
Santini A, Ghelardini L, De Pace C, Desprez-Loustau ML, Capretti P, Chandelier A, Stenlid J (2013) Biogeographical patterns and determinants of invasion by forest pathogens in Europe. New Phytol 197(1):238–250. https://doi.org/10.1111/j.1469-8137.2012.04364.x
Sanzani SM, Li Destri Nicosia MG, Faedda R, Cacciola SO, Schena L (2014) Use of quantitative PCR detection methods to study biocontrol agents and phytopathogenic fungi and oomycetes in environmental samples. J Phytopathol 162(1):1–13. https://doi.org/10.1111/jph.12147
Sapp M, Tyborski N, Linstädter A, Lopez Sanchez A, Mansfeldt T, Waldhoff G, Bareth G, Bonkowski M, Rose LE (2019) Site-specific distribution of oak rhizosphere-associated oomycetes revealed by cytochrome c oxidase subunit II metabarcoding. Ecol Evol 9(18):10567–10581. https://doi.org/10.1002/ece3.5577
Sarker SR, McComb J, Burgess TI, Hardy GESJ (2021) Timing and abundance of sporangia production and zoospore release influences the recovery of different Phytophthora species by baiting. Fungal Biol 125(6):477–484. https://doi.org/10.1016/j.funbio.2021.01.009
Sarker SR, Burgess TI, Hardy GESJ, McComb J (2023a) Closing the gap between the number of Phytophthora species isolated through baiting a soil sample and the number revealed through metabarcoding. Mycol Prog 22(6):39. https://doi.org/10.1007/s11557-023-01892-7
Sarker SR, McComb J, Hardy GESJ, Burgess TI (2023b). Sample volume affects the number of Phytophthora and Phytopythium species detected by soil baiting. Eur J Plant Pathol 1–11. https://doi.org/10.1007/s10658-023-02661-8
Sauquet H, Weston PH, Barker NP, Anderson CL, Cantrill DJ, Savolainen V (2009) Using fossils and molecular data to reveal the origins of the Cape proteas (subfamily Proteoideae). Mol Phylogenet Evol 51(1):31–43. https://doi.org/10.1016/j.ympev.2008.12.013
Scibetta S, Schena L, Chimento A, Cacciola SO, Cooke DE (2012) A molecular method to assess Phytophthora diversity in environmental samples. J Microbiol Methods 88(3):356–368. https://doi.org/10.1016/j.mimet.2011.12.012
Scott PM, Burgess TI, Barber PA, Shearer BL, Stukely MJC, Hardy GESJ, Jung T (2009) Phytophthora multivora sp. nov., a new species recovered from declining Eucalyptus, Banksia, Agonis and other plant species in Western Australia. Persoonia 22(1):1–13. https://doi.org/10.3767/003158509X415450
Shearer BL, Dillon M (1996) Impact and disease centre characteristics of Phytophthora cinnamomi infestations of Banksia woodlands on the Swan Coastal Plain Western Australia. Aust J Bot 44(1):79–90. https://doi.org/10.1071/BT9960079
Simamora AV, Paap T, Howard K, Stukely MJ, Hardy GESJ, Burgess TI (2018) Phytophthora contamination in a nursery and its potential dispersal into the natural environment. Plant Dis 102(1):132–139. https://doi.org/10.1094/PDIS-05-17-0689-RE
Tedersoo L, Drenkhan R, Anslan S, Morales-Rodriguez C, Cleary M (2019) High-throughput identification and diagnostics of pathogens and pests: overview and practical recommendations. Mol Ecol Resour 19(1):47–76. https://doi.org/10.1111/1755-0998.12959
von Broembsen S (1984a) Distribution of Phytophthora cinnamomi in rivers of the south-western Cape Province. Phytophylactica 16(3):227–230
von Broembsen S (1984b) Occurrence of Phytophthora cinnamomi on indigenous and exotic hosts in South Africa, with special reference to the South-Western Cape Province. Phytophylactica 16(3):221–226
von Broembsen SL, Deacon JW (1997) Calcium interference with zoospore biology and infectivity of Phytophthora parasitica in nutrient irrigation solutions. Phytopathology 87(5):522–528. https://doi.org/10.1094/PHYTO.1997.87.5.522
von Broembsen SL, Kruger FJ (1985) Phytophthora cinnamomi associated with mortality of native vegetation in South Africa. Plant Dis 69(8):715–717. https://doi.org/10.1094/PD-69-715
Weston PH, Baker NP (2006) A new suprageneric classification of the Proteaceae, with an annotated checklist of genera. Telopea 11(3):314–344. https://doi.org/10.7751/telopea20065733
White TJ, Bruns T, Lee SJWT, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: a Guide to Methods and Applications 18(1):315–322
Wills RT (1993) The ecological impact of Phytophthora cinnamomi in the Stirling range National Park Western Australia. Austral J Ecol 18(2):145–159. https://doi.org/10.1111/j.1442-9993.1993.tb00439.x
Acknowledgements
We are grateful for the assistance provided by the European Cooperation in Science and Technology (Action FP1401: A global network of nurseries as early warning system against alien tree pests—Global Warning). We also thank Diane White for her contribution to the laboratory analysis.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions This work was supported by the European Cooperation in Science and Technology (Action FP1401: A global network of nurseries as early warning system against alien tree pests—Global Warning).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation, data collection, and analysis were performed by Duccio Migliorini, Christopher Shaw, and Maria Vivas. The first draft of the manuscript was written by Duccio Migliorini and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Section Editor: Tanay Bose
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Migliorini, D., Vivas, M., Wingfield, M.J. et al. Oomycete composition in Proteaceae orchards and natural stands on three continents. Mycol Progress 22, 75 (2023). https://doi.org/10.1007/s11557-023-01925-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-023-01925-1