Abstract
In forest ecosystems, decomposition is essential for carbon and nutrient cycling and therefore a key process for ecosystem functioning. During the decomposition process, litter chemistry, involved decomposer organisms, and enzymatic activity change interdependently. Chemical composition of the litter is the most complex and dynamic component in the decomposition process and therefore challenging to assess holistically. In this study, we aimed to characterize chemical shifts during decomposition and link them to changes in decomposer fungal activity. We characterized the chemical composition of freshly fallen autumn leaves of European beech (Fagus sylvatica) and the corresponding leaf litter after 1 year of decomposition by proton nuclear magnetic resonance spectroscopy. We further tested the applicability of spiking experiments for qualitative and quantitative characterization of leaves and litter chemistry. The composition and transcriptional activity of fungal communities was assessed by high-throughput Illumina sequencing in the same litter samples. We were able to distinguish freshly fallen leaves from 1-year-old litter based on their chemical composition. Chemical composition of leaves converged among regions with progressing decomposition. Fungal litter communities differed in composition among regions, but they were functionally redundant according to the expression of genes encoding litter degrading enzymes (CAZymes). Fungi of the saprotrophic genera Mycena and Chalara correlated with transcription of litter-degrading CAZymes in 1-year-old litter. Forestry measures influenced the diversity and transcription rate of the detected CAZymes transcripts in litter. Their expression was primarily predicted by composition of the soluble chemical fraction of the litter. Environmental NMR fingerprints thus proved valuable for inferring ecological contexts. We propose and discuss a holistic framework to link fungal activity, enzyme expression, and chemical composition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The decomposition of plant litter is essential for nutrient cycling and therefore a key process for ecosystem functioning. This is a dynamic process where litter chemistry, microbial communities, and fungal activity closely interact in a complex feedback network, and our current understanding regarding these feedback responses is still very limited (Peršoh 2015). During plant litter decomposition, CO2 is released, and organic matter is converted into decomposition products (Berg and McClaugherty 2014). Litter decomposition is a vital component of the nutrient cycle and therefore essential for soil fertility and long-term storage of carbon (Berg and McClaugherty 2014). Leaf litter requires several years to decades to fully decompose (Parton et al. 2007). Our current understanding of this complex process is that the decomposition rate of plant litter is mainly controlled by climate and litter quality, but also mechanistically linked with decomposers and extracellular enzyme activities at local scales (Parton et al. 2007; Bradford et al. 2016; Xiao et al. 2020).
Assessing and defining litter quality (i.e., chemical composition of litter) is challenging, because plant litter is composed of a plethora of organic compounds with different susceptibilities to break down during decomposition (Berg and McClaugherty 2014). The early stage of decomposition is particularly challenging to define and study, due to its complex composition, quick degradation and synthesis of decomposition products, and loss of soluble compounds by leaching (Berg and McClaugherty 2014). Traditional approaches characterize litter quality by sum parameters, such as total phenolics, carbohydrate content, and “Klason lignin” (Dence 1992) and do not allow for a holistic characterization of the litter chemistry. Understanding of the litter decomposition process would benefit from a more detailed characterization of the various molecules being degraded, their degradation rate and mechanisms.
Nuclear magnetic resonance (NMR) spectroscopy is a technique commonly applied in plant metabolomics (Fiehn 2002; Holmes et al. 2006) and able to fully characterize the chemical composition of organic matter at the molecular level (Krishnan et al. 2005; Wishart 2008). NMR is advantageous over techniques often used to analyze litter composition (e.g., chemical stoichiometry or mass spectrometry), because, while being highly reproducible, stable, rapid, and requiring minimal sample preparation, NMR allows quantification of biomolecules in liquid or solid phase (Dunn et al. 2005; Dunn and Ellis 2005; Pauli et al. 2005; Holmes et al. 2006; Bonanomi et al. 2019). This is a convenient and powerful tool for metabolic profiling and monitoring metabolic changes in natural processes (Pauli et al. 2005). By analyzing plant litter at different decomposition stages, it is possible to assess the litter chemical composition during the decomposition process (Preston et al. 2009; Bonanomi et al. 2019).
Plant-litter decomposition in forest ecosystems is accomplished primarily by fungi and secondarily by bacteria (Baldrian et al. 2012; Stursová et al. 2012; Žifčáková et al. 2016; Žifčáková et al. 2017; Lladó et al. 2017), while other saprotrophic organisms, such as animals and protists, play a minor role (Persson et al. 1980; Bonanomi et al. 2019). Fungi dominate the synthesis and secretion of extracellular enzymes which degrade the plant compounds (de Boer et al. 2005; Žifčáková et al. 2017), while bacteria comprise nitrogen fixation and some cellulolytic activity (Lladó et al. 2017). Key exoenzymes for litter decomposition are the so-called CAZymes (“Carbohydrate Active enZymes”) (Lombard et al. 2014). They have a major role during the early stage of decomposition and quickly degrade the easily accessible soluble compounds of the plant litter (Davidson and Janssens 2006; Berg and McClaugherty 2014). The genetic capabilities of individual fungi in general (Zhao et al. 2013; Osono 2020), and functional guilds in particular (Wolfe et al. 2012; Zhao et al. 2013; Sista Kameshwar and Qin 2017), differ strongly with regard to CAZyme synthesis. However, fungal decomposer communities are considered to be functionally redundant across forest ecosystems, even though they differ in taxonomic composition (Talbot et al. 2014). Activity of released enzymes changes the chemical composition of litter, and thus accessibility and availability of nutrients for fungi, entailing a succession of differently specialized decomposer communities (Baldrian et al. 2012; Voříšková and Baldrian 2013; Purahong et al. 2014b; Purahong et al. 2015; Purahong et al. 2016). While some studies already analyzed fungal community composition and their transcriptional activity together (Albright et al. 2020; Žifčáková et al. 2016; Schneider et al. 2021), so far, individual fungi remain to be linked to specific chemical changes in litter.
Fungal communities and their functions are commonly assessed by DNA metabarcoding and enzyme activities, respectively (Voříšková and Baldrian 2013; e.g. Eichlerová et al. 2015; Purahong et al. 2016; Žifčáková et al. 2016). Functions are linked to communities by transcription (mRNA) of active (rRNA) organisms, followed by post-transcriptional modifications and interaction of the synthesized enzymes with the surrounding environment. These linkages are mostly indirect since active and total organismic communities can differ in composition (Baldrian et al. 2012) and enzyme abundance may be weakly linked to enzyme activity in litter (Schneider et al. 2012). A more direct approach is required to provide insights into the fungal response to the surrounding substrate (i.e., litter).
In this study, we attempted to identify chemical shifts of compounds decomposed and synthesized during the first year of decomposition. We aimed at characterizing the chemical composition of falling and decomposing autumn leaves, while providing insights into the fungal decomposer community and respective CAZyme transcription. We hypothesized that the CAZyme transcription profiles are a response to the soluble fraction of the litter chemistry. Furthermore, given the expected enzymatic decomposition, we hypothesized that the soluble fraction (assessed by 1H-NMR) can clearly differentiate leaf and litter samples that might be unclear in the complete unfractionated litter chemistry (assessed by 13C solid state NMR). We expected the availability of energy-rich and easily accessible compounds, such as glucose, to decrease with decomposition. We hypothesized that chemistry of freshly fallen leaves shapes the total fungal community detected in litter 1-year-old litter, while the part of the community being active at that time is predominantly shaped by the litter chemistry at that time. Communities and substrate chemistry are expected to reflect adaptations to local factors. However, the response of the different communities is hypothesized to be functionally redundant, as shown before in different soil layers and forest types (Talbot et al. 2014; Žifčáková et al. 2017).
To test our hypotheses and to reduce and account for effects of local and regional factors, we selected 14 different beech (Fagus sylvatica) forest plots in three distant regions in Germany (Fig. 1A). We used proton nuclear magnetic resonance (1H-NMR) spectroscopy to assess the chemical composition of freshly fallen and 1-year-old decaying leaves. To identify NMR peaks corresponding to compounds of interest, we performed spiking experiments. The species composition of the fungal community in litter was assessed by metabarcoding and their transcriptional activity by metatranscriptome sequencing. We conducted model-based and correlative analyses to determine interactions between total and active fungal communities, litter-degrading enzyme transcription, and substrate chemistry (Fig. 1B).
Study design (A) and hypothesis (B). Falling autumn leaves and 1-year-old litter of beech (Fagus sylvatica) were collected in 14 plots in three distant regions in Germany (A). The chemical composition was assessed by 1H-NMR and fungal litter community composition and activity by metagenome barcoding and metatranscriptome sequencing. Hypothesized connections tested (black) and inferred (gray) in this study (B). Chemical composition of leaves shapes the composition of the fungal communities. Chemical composition of the litter influences the fraction of the total fungi being metabolically active at a certain point in time. The active community transcribes genes encoding litter-degrading CAZymes, which change the chemical composition of the leaf-litter. A continuous feedback loop is maintained by adaptation of fungal activity to changing litter chemistry
Materials and methods
Study area and sample collection
To avoid effects of host-specific microbiomes and chemistry, this study focused on a single tree species, i.e., European Beech (Fagus sylvatica). The study sites were beech-dominated forest plots, selected within the framework of the German Biodiversity Exploratory project for large-scale and long-term functional biodiversity research (Fischer et al. 2010). The studied forest plots were located in three widely distant (>300 km) regions in Germany (Hainich-Dün, Schorfheide-Chorin and Swabian Alb), which comprised natural forests as well as stands managed by age-class and selection-cut practices (Table S1). Selection-cut practice was exclusive to Hainich-Dün, while age-class practice was similar among the different regions (Fischer et al. 2010). The broad sampling design allowed us to test reproducibility of results among the regions, while assessing if community functions are generalizable on the different geographical scales and biological levels.
Senescent beech leaves were collected in autumn 2014 at five positions per plot, by shaking whole trees or major branches and collecting the falling leaves in nets (Table S2). Damaged leaves or visually colonized by fungi or insects were discarded. The leaf samples were stored in paper bags and transported to the laboratory at ambient temperature. A soil core (Ø 5.5 cm) was collected in the same positions, stored at 4 °C and transported to the laboratory where it was processed within 12 h. Processing the samples in the laboratory assured that the different datasets were obtained from the same source material. The 1-year-old litter layer was subsequently subsampled from the soil cores and pooled from two or three positions per plot (Table S2 and S3). One-year-old litter recognizable as mostly structurally intact but flattened leaves between the freshly fallen litter layer and the at least initially fragmented to largely decomposed litter. The similar macro-morphological decomposition states are likely to indicate the same year of leaf abscission, even if litter age may be arguably variable among samples.
Nucleic acid co-extraction and RNA processing
The nucleic acids from 0.2 g of litter were extracted under RNase-free conditions by adapting the protocol from Peršoh et al. (2008). To co-extract the nucleic acids, 700 μL of Tris-Buffer (0.1 M, pH 8) and 35 μL of aluminum sulfate (Al2(SO4)3; 4 M) were added to each 0.2 g of litter sample. The sample was homogenized in a FastPrep®-24 Instrument (MP Biomedicals, Eschwege, Germany) at 4.0 m s−1 for 20 s. The pH was then adjusted to pH 8 by adding NaOH (4 M), and re-adjusted to 5.5 by adding 3.0 μl of acetic acid. A mixture of glass beads (Ø 0.1–0.25 mm: 0.15 g; Ø 0.25–0.5 mm: 0.12 g; Ø 1.25–1.55 mm: 0.12 g) and steel beads (Ø 3 mm: 3 beads), 325 μL of extraction buffer (0.4 M LiCl, 200 mM Tris-HCl (pH 8), 120 mM EDTA (pH 8), 10% SDS), and 500 μl of phenol were added to the sample. Cells were disrupted using a FastPrep®-24 Instrument (MP Biomedicals) at 4.0 m s−1 for 30 s, followed by two steps of 6.5 m s−1 for 40 s, with an intermediate incubation on ice of 5 min. After centrifugation (14,165 g for 1 min at room temperature), 750 μL phenol:chloroform:isoamyl alcohol (25:24:1 v/v) was added to 750 μL of the supernatant. The sample was mixed and incubated on ice for 5 min, while being mixed every 90 s. After centrifugation (17,650 g for 15 min at 4 °C), three consecutive centrifugation (17,650 g for 15 min at 4 °C) steps, with addition of an equal volume of chloroform:isoamyl alcohol (24:1) to the supernatant, with intermediate incubations on ice of 5 min were performed. Nucleic acids were precipitated by adding 0.1 volumes of NaCl (5 M) and 0.7 volumes of isopropanol to 500 μl of the supernatant to the supernatant. After overnight incubation at 20 °C, the sample was centrifuged (17,650 g for 60 min at 15 °C), and the resulting pellet was air-dried for 3 min and resuspended in 20 μL of RNase-free water. Concentration and quality of the nucleic acids were assessed using a NanoDrop 1000 spectrophotometer (Thermo Fisher Scientific Inc., Darmstadt, Germany). The nucleic acids were stored at 80 °C until further processing.
To purify the total RNA, 16 μl of nucleic acids was incubated with 2 μl of DNase I (1 unit/μl) (Sigma-Aldrich, Munich, Germany), in 2 μl of the correspondent buffer (200 mM Tris-HCl, pH 8.3, with 20 mM MgCl2), for 15 min at room temperature. The reaction was stopped by adding 2 μl of stop solution (50 mM EDTA), followed by an incubation for 10 min at 70 °C. The DNA digestion was tested by attempting to amplify the ITS region by PCR according to Guerreiro et al. (2018). The quality of the total RNA was assessed on a Eukaryotic Total RNA Nano chip with a 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA).
mRNA library preparation and sequencing
Metatranscriptomic libraries (Table S3) were prepared and barcoded using NEBNext® Ultra™ Directional RNA Library Prep Kit for Illumina® (New England Biolabs GmbH, Frankfurt am Main, Germany), according to the Poly(A) mRNA Magnetic Isolation Module protocol and to the manufacturer’s recommendations. To assure similar coverage among samples, the libraries were sequenced twice on an Illumina MiSeq® sequencer (Illumina, Inc., San Diego, CA, USA), with 2 × 300 bp paired end sequencing (MiSeq Reagent Kit v3 Chemistry, Illumina, Inc., San Diego, CA, USA). The raw sequence data were deposited in the EBI Metagenomics database (https://www.ebi.ac.uk/metagenomics; PRJEB7861 and PRJEB8419).
Processing of metatranscriptomic data
Metatranscriptomic raw reads from both runs were assembled de novo as paired-end reads by CLC Genomics Workbench (CLCbio, Qiagen, Valencia, CA, USA), using default settings for trimming, automatic word and bubble size, and a minimum contig length of 110 bp. The reads were afterwards mapped against the obtained contigs.
Assembled metatranscriptomic data were annotated using the online tool KAAS (KEGG Automatic Annotation Server) Ver. 2.1 (Moriya et al. 2007) and the dbCAN2 release 9.0 (Zhang et al. 2018). Coverages of contigs were normalized per sample by dividing the sum of coverages for contigs encoding for the ribosome pathway (03010 Ribosome [PATH:ko03010]), as identified by KAAS. The normalized coverage was treated as abundance of each contig (Table S4). Litter-decomposing CAZymes were selected according to the literature (Sinsabaugh et al. 1991; Kubicek et al. 1993; Schneider et al. 2010; Burns et al. 2013; Eichlerová et al. 2015; Talbot et al. 2015; Kameshwar et al. 2019). Only CAZyme families detected in 5 or more samples were considered for further analyses (Table S4). The read coverage for each contig was considered the contig-level transcription rate. The sum of the transcription rates of all contigs encoding the same CAZyme family was considered the CAZyme-level transcription rate.
Amplicon library preparation and sequencing
To assess composition of the fungal communities, the complete fungal barcode ITS rRNA gene region (Schoch et al. 2012; Stielow et al. 2015) was amplified and sequenced as detailed by Guerreiro et al. (2018). Briefly, the preparation of the amplicon libraries comprised two sequential amplification steps. The first amplification step was performed with the fungal specific primer combination ITS1F (Gardes and Bruns 1993) and ITS4 (White et al. 1990), modified to include TAG sequences (Guerreiro et al. 2018), which allow for multiplexing of multiple samples. The second amplification step was performed with primers which included sequences of the sequencing primer, the indices and the P5 and P7 adapters for the Illumina sequencing (Guerreiro et al. 2018). PCR products were pooled equimolarily and the library purified by gel electrophoresis (BluePippin, Sage Science, Beverly, MA, USA) by selecting amplicon sizes >350 bp. The amplicon libraries were sequenced on an Illumina MiSeq® sequencer (Illumina, Inc., San Diego, CA, USA), with 2 × 250 bp paired end sequencing (MiSeq Reagent Kit v3 Chemistry).
Processing of metabarcoding data
The obtained raw Illumina metabarcoding reads were processed using the open-source software QIIME version 1.9.0 (Caporaso et al. 2010) as detailed by Röhl et al. (2017). Briefly, the TAG sequences were extracted and quality filters (one ambiguity allowed and reads were truncated after 19 consecutive low quality, i.e., Phred > 19, base calls) were applied. The quality-filtered reads were assigned to samples, based on their identifiers (i.e., indices and TAG sequences). The ITS1 barcode region was subsequently extracted and trimmed to exclude the majority of the SSU rRNA gene region. The pre-processed sequence reads were deposited in the European Nucleotide Archive database (http://www.ebi.ac.uk/ena, accession number PRJEB19870).
The reads were clustered into Operational Taxonomic Units (OTUs), using CD-HIT-OTU for Illumina reads version 0.0.1 (Fu et al. 2012) and a 97% similarity threshold, which were considered as proxies for species. The generated table listing the read count per sample and OTU was filtered and samples with less than 1000 reads were discarded (Table S5). OTUs were assigned to taxa (Table S6) based on matches to the representative sequence in the UNITE database version 7.0 (Kõljalg et al. 2013). Only fungal OTUs were selected for downstream analyses. Total and active fungal communities were distinguished according to the amplicons originating from DNA and RNA extracts, respectively (Anderson and Parkin 2007). The normalized amplicon read count indicated species abundance or activity for fungal OTUs in total and active communities, respectively.
Metabarcoding studies often deal with extensive amount of information and rely on read abundance for interpretation of the results. Comparing relative abundances of OTUs among similar samples processes appears reliable (Peršoh 2015; Sinclair et al. 2015; Röhl et al. 2017; Guerreiro et al. 2018), even though absolute counts of amplicon reads are prone to quantitative biases and do not necessarily reflect biological abundances (Peršoh 2013). Quantification by qPCR also recently confirmed results indicated by metabarcoding (Gossner et al. 2021). This indicates that metabarcoding is a suitable approach to address abundance or activity of individual fungal species among complex community data.
Metametabolome extraction
Leaf and 1-year-old litter samples were dried in a drying oven for about 8 weeks at 70 °C until constant weight. The dried samples were ground in a Mixer Mill MM 2000 (Retsch GmbH, Haan, Germany) for 2 min at 70% amplitude, with a titanium sphere, in titanium vessels. The pulverized material of leaves and litter, respectively, was pooled per plot. Two hundred fifty milligrams of pulverized material were mixed with glass beads (each 80 μl beads of Ø 0.1–0.25 mm, Ø 0.25–0.5 mm and Ø 1.25–1.55 mm) and 1000 μl of Chloroform (Roth, Karsruhe, Germany) or methanol (Roth) as solvent and incubated in a FastPrep®-24 Instrument (MP Biomedicals) for 80 s at 6.5 m s−1. The suspension was centrifuged (Eppendorf 5424, rotor FA-45-24-11; Eppendorf, Wesseling-Berzdorf, Germany) for 5 min at 15,000 rpm, the supernatant transferred into a spin-column (GeneON, Maxima Spin Column GF-F1 1.5 ml; GeneON, Ludwigshafen, Germany), centrifuged for 3 min at 6000 rpm, and the flow-through stored. Another 1000 μl of solvent was added, the extraction (incubation, centrifugation, and filtration) repeated and the two flow-throughs combined. The combined flow-through was dried in a Vacuum Concentrator Centrifuge UNIVAPO 100 ECH (UniEquip, Planegg, Germany) at 1250 rpm and 60°C until dried and resuspended in 800 μl of the corresponding but deuterated solvent (Sigma-Aldrich). The suspension was centrifuged for 5 min at 12,000 rpm and 700 μl of the supernatant was transferred into an NMR tube (5-mm diameter).
1H-NMR measurement and data processing
All solution spectra were recorded on Bruker DRX 400, DRX 600, and AVANCE IIIHD 700 spectrometers (Table S7), respectively. 1D 1H and 1D 1H NOESY spectra were acquired at 298 K and 303 K, applying Bruker standard pulse programs. Typically, 512 scans were applied for a spectral size of 64,000 data points with an acquisition time of 1.7039 s and a relaxation delay of 2 s.
The resulting spectra were processed in MestReNova software version 11.0.0–17609 (Mestrelab Research S.L., Santiago de Compostela, Spain). Spectra were manually corrected for the phase and baseline. Bins with a width of 0.005 ppm were generated, with the “peaks” method, without normalization (Table S8). The residual water influenced regions (from 1.28 to 1.29 ppm in chloroform-d), the solvent peak regions (7.285 to 7.29 ppm in chloroform-d and 3.32 to 3.375 ppm in methanol-d4), a highly intense peak (1.31 ppm) in methanol-d4, and peaks with an intensity < 1×10−9 were excluded from further analyses. The binned data were normalized by cumulative sum scaling (CSS) (Paulson et al. 2013). Normalization by total was applied to allow relative comparison of resonances among samples. The derived resonance data were categorized into functional chemical groups according to Pretsch et al. (2009). The chemical composition of the substrate was represented as the chemical shift (ppm) and height of 1H-NMR peaks in the soluble fraction of leaf and litter samples.
Spiking experiments
The identification of NMR peaks corresponding to glucose and galactose was achieved by performing spiking experiments. The presence of sugar compounds in two leaf and the corresponding litter extracts (Table S7) was determined by performing NMR measurements before and after the addition of 35 μl of d-(+)-glucose (50 mM), 35 μl of d-(+)-galactose (50 mM) (Sigma-Aldrich), and 3.5 μl of N-acetyl-d-galactosamine (50 mM) (Sigma-Aldrich). Due to the chemical properties of the sugars, the compounds for the spiking experiments were prepared in methanol-d4 and only the methanol-d4 fractions of the extracts were considered. Non-spiked and spike spectra were imported into MestReNova, where they were processed, aligned, and superimposed and the corresponding compound peaks identified.
Spiking experiments with d-(+)-glucose were repeated as detailed for the 14 leaf and litter methanol extracts. The line fitting method in MestReNova was used to calculate the intensity of the peaks. To determine the glucose content in each spectrum, a reference peak was selected (approximately at 5.147 ppm) and its intensity used as reference. The ratio of the glucose and the reference peak before spiking was divided by the ratio of the glucose increment and the reference peak after spiking.
Solid-state NMR
For two leaf samples and the respective litter samples (Table S7), 13C solid-state NMR spectra were acquired using magic angle spinning on a Bruker Avance III HD spectrometer with an external magnetic field of B0 = 9.4 T. All spectra are referenced with respect to TMS. The samples were packed in 4.0-mm ZrO2 rotors and spun 10 kHz. For the cross polarization experiments was a ramped cross-polarization (CP) experiment, where the nutation frequency νnut on the proton channel was varied linearly by 50%. For the excitation of the proton spin bath, we applied a νnut = 100 kHz. νnut on the 13C channel and the spin lock time was adjusted to 70 kHz and 3 ms, respectively. The single pulse (SP) experiments were carried out with a 90° pulse excitation where the pulse length was set to 3.5 μs. The recycle delays for the 13C CP and 13C SP MAS spectra were adjusted to 1 s and 120 s, respectively. During acquisition, proton broadband decoupling was applied using a spinal-64 sequence (Fung et al. 2000) with νnut = 70 kHz. Spectra were superimposed, manually corrected for the phase and baseline using MestReNova software. Bins were generated with a width of 0.005 ppm, using the “peaks” method and without normalization. The derived resonance data were categorized into functional chemical groups as described in Bonanomi et al. (2019).
Statistical analyses
Tables of OTU, contig, and NMR quantities were normalized by CSS (Paulson et al. 2013). Normalization by CSS outperforms other normalization methods by avoiding biases derived from uneven sequencing depth (Weiss et al. 2017). The CSS-standardized tables were subsequently imported into Primer 7 version 7.0.11 (Clarke and Gorley 2015) and further re-standardized per sample by total abundance or total intensity. For analyses considering the relative fungal abundance, relative CAZyme transcription, or chemical resonance intensity, Bray–Curtis similarity matrices were calculated (Bray and Curtis 1957). Analyses of similarity (ANOSIM) were calculated in Primer 7 based on Bray–Curtis similarity matrices and using 10,000 permutations. Distance to centroids among samples derived from plots of the same region was calculated using Primer 7 PERMDISP (distance-based test for homogeneity of multivariate dispersions) method, with 10,000 permutations.
Diversity indices for each CAZymes transcribed were calculated using Primer 7 DIVERSE methods, based on contig composition and relative abundance. Diversity data were imported into R version 3.4.3 (R Core Team 2017), where linear models were applied to calculate the grand mean effect of the forest management on each individual enzyme diversity. The effects of forest management on the diversity of each CAZyme family were calculated as the log response ratio of the enzyme diversity between natural and age-class plots (Supplementary script S1).
Ternary plots were calculated with the “ggtern” package (Hamilton 2018). Spearman coefficients and significance were obtained with the “psych” package (Revelle 2017) without adjustments. The resulting tables were filtered for significant correlations (P < 0.05) and imported into Cytoscape 3.8.2 (Shannon et al. 2003) for visualization.
Euler Diagrams with proportional areas were generated using the BioVenn web application (Hulsen et al. 2008). Kruskal–Wallis tests were performed using STATISTICA 64 version 13.2 (StatSoft, Inc., Tulsa, USA). All statistical tests were considered significant at P < 0.05.
Direct and indirect effects of leaf and litter chemical composition, total (i.e., DNA-derived) and active (i.e., RNA-derived) litter fungal communities, and respective enzyme transcription profiles were jointly modeled with a partial least square path model (PLS-PM) approach (Wold 1975). Briefly, the abundance data were Hellinger-transformed (Legendre and Gallagher 2001) and scaled (via subtraction of the mean and division by the standard deviation) in order to yield scaled and thus directly comparable coefficient estimates between the latent variables (Supplementary script S2). All analyses were conducted with R (R Core Team 2017), using the packages “plspm” (Sanchez et al. 2017) and “ggplot2” (Wickham 2009).
Data accessibility
The sequence data are available from the European Nucleotide Archive (http://www.ebi.ac.uk/ena/) under study PRJEB19870 and from the EBI Metagenomics database (https://www.ebi.ac.uk/metagenomics) under studies PRJEB7861 and PRJEB8419. Raw 1H-NMR spectra were archived in Open Science Framework (DOI 10.17605/OSF.IO/2TJC3; https://osf.io/2tjc3/?view_only=593854b6f52f45ebbe285374051a0d52). Corresponding metadata are provided in Tables S2, S3, and S7. All data generated or analyzed during this study are included in this published article and its supplementary information files.
Results
Regional influence on litter fungal community structure
ITS-metabarcoding yielded a total of 963,520 quality-filtered ITS1 reads from litter samples. They clustered into 492 fungal OTUs, 467 of which were found in the DNA and 442 in the RNA extracts. The majority of the OTUs (417 OTUs, 85%) were detected in both extracts (Fig. S1). Across all 14 plots, Ascomycota was the most frequent and abundant phylum in both DNA (271 OTUs, 65.7% of all reads) and RNA extracts (256 OTUs, 63.4% of all reads), followed by Basidiomycota (DNA: 137 OTUs, 28.9% reads; RNA: 127 OTUs, 30.0% reads), fungi of unknown phylum affiliation (DNA: 21 OTUs, 3.8% reads; RNA: 21 OTUs, 4.4% reads) and Mucoromycota (DNA: 5 OTUs, 1.9% reads; RNA: 5 OTUs, 2.3% reads). The DNA- and RNA-derived communities differed significantly in composition (R = 0.204, P = 0.0002), with 46 OTUs (4.9% reads) occurring exclusively in DNA communities and 21 OTUs (3.5% reads) exclusively in RNA communities. The DNA-derived fungal litter communities were regionally structured (R = 0.37, P = 0.0050), while the RNA-derived communities only differed by trend across the regions (R = 0.194, P = 0.0840) (Fig. 2A, Fig. S2). The latter were regionally indistinct for Basidiomycota (R = 0.05, P = 0.3030), while composition of the (sub)community of non-Basidiomycota OTUs differed significantly among regions also according to the RNA extracts (R = 0.26, P = 0.0400) (Fig. 2A). Forest management had no significant impact on any community structure (R < 0.094, P > 0.3140).
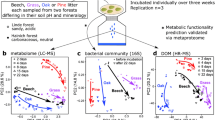
Regional preferences of fungal communities and CAZyme transcripts. Differences among regions according to total (DNA-based) and active (RNA-based) litter fungal community composition, transcription profiles of contigs and the inferred litter-degrading CAZyme families are represented (A). Magnitude (ANOSIM R-value) of the differences is depicted and significance (ANOSIM P) is indicated by asterisks (**P ≤ 0.01; *P < 0.05; ns: P ≥ 0.05). Differences according to whole (black bars) and non-Basidiomycetous fungal communities (gray bars) are shown separately. Distribution of the contigs encoding CAZymes (B) and corresponding CAZyme families (C). Symbol size indicates the average relative abundances. The CAZyme classes auxiliary activity (AA), glycoside hydrolases (GH), glycosyl transferases (GT), carbohydrate esterases (CE), and polysaccharide lyases (PL) and the carbohydrate-binding modules (CBM) are color-coded. Arrows indicate the orientation of the axes
Influence of geographical regions in the transcription of litter-degrading CAZymes
We obtained 89,068,690 mRNA reads, which were assembled into 1,287,115 contigs. Annotation of the contigs against the CAZyme database (dbCAN2 v9) revealed 216 CAZyme families (Table S9), 79 of which were reported to be involved in litter decomposition. The CAZyme families selected for further analyses were encoded by 4526 contigs and comprised 13 auxiliary activity (AA), 41 glycoside hydrolases (GH), 1 glycosyl transferase (GT), 9 carbohydrate esterases (CE), 3 polysaccharide lyases (PL) families, and 12 carbohydrate-binding modules (CBM) (Table S4).
The transcription profiles of the contigs encoding the selected litter-degrading CAZymes were regionally structured (R = 0.513, P = 0.0004) and highly variable (Fig. S3A). The similarity among the transcription profiles of the contigs decreased significantly (P < 0.0001) with increasing geographical distance (Fig. S2). However, summarizing the transcription rates of contigs coding for the same CAZyme families resulted in similar profiles within (Fig. S3A) and among regions (R = −0.046, P = 0.6170) (Fig. 2, Fig. S3B). Furthermore, the expression of the CAZyme families was independent of the geographical distance (Fig. S2). This is supported by the absence of many individual contigs from at least one region, while almost all CAZymes were transcribed in all regions (Fig. 2).
The predicted functional activity of the contigs differed significantly among regions for most functions involved in litter decomposition (cellulolytic, hemicellulolytic, ligninolytic, and pectinolytic activity), except for starch-degrading activity (Fig. S4). However, the geographical regions did not influence the predicted functional activity of the CAZyme families (Fig. S4).
Influence of management in transcription profiles and encoded CAZyme families
Almost all CAZyme families were transcribed in similar proportions in all forest management types (Fig. S5). However, diversity indicators of contigs encoding individual litter-degrading CAZyme families were sensitive to forestry measures (Fig. 3). While contigs of one CAZyme family were in general equally evenly distributed across management types, their diversity was higher in age-class (n = 4005) than in natural forests (n = 2723). Furthermore, individual families responded differently to forest management (Fig. S6); e.g., PL4 and GH30 were encoded by the most different contigs in age-class forests. Additionally, the highest richness for contigs encoding the CBM52 and GH63 was observed in natural forests.
Effects of management on the contig diversity within litter-decomposing CAZyme families. The grand mean of all CAZyme families is represented for each diversity index. Indices were calculated for each individual CAZyme family based on the relative abundance of the respective contigs. Only samples from age-class and natural forests in Hainich-Dün (2 age-class plots; 3 natural plots) and Schorfheide-Chorin (3 age-class plots; 2 natural plots) were considered. Points on the left and right side of the dotted line indicate lower and higher diversity, respectively, in natural as compared to age-class forests. Effects are deemed significant if their corresponding approximated 95% confidence interval (displayed as horizontal lines) does not cross the dotted line of zero effect
The transcription rate of some CAZyme families was also influenced by forest management. In natural forests, the expression profiles were on average dominated by GH43 (13%) and GH16 (12%), while in age-class forests GH16 (10%) and AA9 (9%) were the highest expressed CAZyme families (Fig. S7). Transcription of 35 and 27 CAZyme families was higher in age-class and natural forests, respectively. Age-class and natural forests differed most according to the expression levels of AA8 (32.9 times higher in age-class forests), GH26 (6.7 times higher in age-class forests), CBM63 (40.1 times higher in natural forests), CBM52 (6.4 times higher in natural forests), and GH44 (5.5 times higher in natural forests) (Fig. S8). The transcription of GH67, GH13, GH95, AA6, GH47, and GH131 differed less than 5% in both managements (Fig. S8). Despite the observed influences on the expression of individual CAZymes, the overall transcription profiles of the contigs (R = −0.068, P = 0.6510) and respective litter-degrading CAZyme families (R = 0.04, P = 0.2940) were similar for between age-class and natural forests.
Differences in NMR spectra according to region and decay state
Solid-state 13C-NMR spectra of leaves and litter were highly similar (Fig. 4A–B). Although some consistent patterns were visible in some of the spectral regions between leaves and litter (e.g., higher relative abundance of carboxyl C in litter), these were not consistent among the two locations (Fig. S9). Furthermore, quantitative solid-state 13C-NMR spectra of litter before and after methanol and chloroform extraction were also highly similar (Fig. 4C). In contrast, liquid NMR provided higher resolution and leaf samples were distinguishable from litter samples according to their 1H-NMR spectra, of both, methanol (i.e., polar fraction) (R = 0.99, P < 0.0001) and chloroform (i.e., apolar fraction) extracts (R = 1, P < 0.0001). Overall, 267 NMR peaks (132 peaks from chloroform and 135 peaks from methanol extracts) were responsible for 50% of the difference between leaves and litter (Table S10). In both fractions, we obtained 852 NMR peaks occurring in leaves and litter, while 592 peaks occurred only in leaves and 446 only in litter (Fig. S10). NMR peaks occurring exclusively in leaves, accounted for 25.5% of the total chemical composition of leaves, while peaks occurring exclusively in litter accounted for 16.1% of the litter chemical composition. Even though leaves and 1-year-old litter differed in the presence and height of NMR peaks (Fig. S10 and S11), the NMR peak heights summed for each of the major chemical classes were similar in leaves and litter (Fig. S12).
Solid-state NMR spectra of leaves and 1-year-old litter. 13C CP MAS NMR spectra (A, B) of leaves (solid line) and corresponding 1-year-old litter (dashed line) originating from 2 plots (HEW10 and AEW08). Quantitative SP MAS NMR measurements from samples of 1-year-old litter (C) before (dashed line) and after extraction of compounds soluble in methanol (gray line) or chloroform (black line)
1H-NMR spectra of leaves differed among the three regions for both extractives (R > 0.57, P < 0.0001) (Fig. 5A). For litter, the methanol extracts differed slightly among regions (R = 0.38, P = 0.007), while the chloroform extracts did not differ (R = 0.16, P = 0.118). The majority of NMR peaks was shared among regions and most of those occurred in at least two locations (Figs. 5B and S13). Despite the differences observed in the peak distribution among the spatial categories, no differences were found regarding chemical functional groups in leaf or litter composition (Figs. 5C and S14). Forest management showed no influence in the chemical composition of leaves and litter (R < −0.012, P > 0.503), where most NMR peaks were shared among practices and among and within regions (Figs. S15 and S16).
Regional distribution of chemical compounds. Differences among regions according to chemical composition of leaves and litter (A). Magnitude (ANOSIM R-value) of the differences is depicted and significance (ANOSIM P) is indicated by asterisks (***P ≤ 0.0001; **P ≤ 0.01; ns: P ≥ 0.05). Distribution of 1H-NMR peaks in the polar fraction of litter among the regions (B). Relative abundances of the peaks are indicated by symbol size; chemical shifts (from −1 to 12 ppm) are color-coded. Distribution of main chemical functional groups in the polar fraction of litter among the regions (C). Relative abundances of the groups are indicated by dot size; chemical functional groups are color-coded. Categorization of the compounds into functional chemical groups was performed according to Pretsch et al. (2009). Arrows indicate the orientation of the axes
Assignment and quantification of glucose and galactose
Spiking with d(+)-glucose resulted in an increase in heights of doublet peaks of the NMR-spectra at 4.48–4.51 ppm and 5.115–5.135 ppm (Fig. S17). Doublets at 5.155–5.169 ppm increased by the addition of d(+)-galactose. N-Acetyl-d-galactosamine addition entailed increases in peak heights at 2.015 ppm and 5.148 ppm. While these peaks were absent from the non-spiked spectra, those increased by glucose and galactose addition were also detected in the non-spiked spectra of the two analyzed leaf and litter extracts. In methanol extracts from litter collected across 14 plots, the glucose content (i.e., heights of corresponding peaks) was significantly lower for Hainich-Dün (P < 0.0079) than for Schorfheide-Chorin (Fig. S18). In methanol extracts from the corresponding autumn leaves, glucose was on average 2.7 times higher in age-class than in natural forests (P = 0.0317; Fig. S18).
Links between fungal activity, CAZyme transcription, and substrate chemistry
The orders Agaricales (Basidiomycota, Agaricomycetes), Helotiales (Ascomycota, Leotiomycetes), and Pleosporales (Ascomycota, Dothideomycetes) contributed the most to both DNA- and RNA-derived communities, accounting for 14%, 12–13%, and 8%, respectively, in both datasets (Table 1). These were also by far the most taxonomically diverse orders, contributing to the overall species richness with 51 and 49 (Agaricales), 48 and 44 (Helotiales), and 43 and 40 OTUs (Pleosporales), in the DNA- and RNA-derived datasets, respectively. For most OTUs within these orders, taxonomic assignment beyond order level was unreliable (Table S6). Abundances of individual RNA-derived OTUs were indirectly correlated to the heights of specific NMR peaks, via correlations with the transcription level of litter-degrading CAZyme families (Fig. 6). The abundances of OTU11 (Ciliciopodium sp., Hypocreales, Ascomycota), OTU23 (Chalara sp., Pezizomycotina, Ascomycota), and OTU47 (unidentified Helotiales, Ascomycota) (Table S6) in the RNA extracts were significantly (Spearman P < 0.05) and positively (0.61 ≤ R ≤ 0.73) correlated with the transcription level of the litter-degrading CAZyme family PL4 (rhamnogalacturonan endolyase, EC 4.2.2.23) (Fig. 6). The PL4 transcription level was again significantly (Spearman P < 0.05), positively and negatively correlated with the heights of two groups of NMR peaks, respectively, of the polar chemical compounds of falling autumn leaves. The NMR peak heights were negatively correlated between and positively within the groups.
Linkage between fungal activity and putative chemical compounds via correlation with enzyme transcription rates. Network of significant pairwise Spearman correlations (P < 0.05) between transcription of the litter-degrading CAZyme families (black) with RNA-derived fungal OTU abundances (brown) and leaf (A) and litter (B) 1H-NMR peak abundance (green) of the respective polar fraction. The polysaccharide lyase family 4 (PL4) and the glycoside hydrolase family 27 (GH27) were considered. Numbers indicate the chemical shifts (ppm) of compounds detected in the respective substrate. Glucose chemical shifts (yellow) were determined by spiking experiments. Positive significant correlations (R > 0, P < 0.05) are indicated in blue and negative significant correlations (R < 0, P < 0.05) in red
The abundances of OTU7 (Mycosphaerella sp., Mycosphaerellales, Ascomycota), OTU31 (unidentified Ascomycota), and OTU35 (Mycena sp., Agaricales, Basidiomycota) in the RNA extracts were significantly (Spearman P < 0.05) and positively (0.53 ≤ R ≤ 0.59) correlated with the transcription level of the litter-degrading GH9 CAZyme family, a multifunctional enzyme which includes endoglucanase (EC 3.2.1.4), endo-β-1,3(4)-glucanase/lichenase-laminarinase (EC 3.2.1.6), β-glucosidase (EC 3.2.1.21), lichenase/endo-β-1,3-1,4-glucanase (EC 3.2.1.73), exo-β-1,4-glucanase/cellodextrinase (EC 3.2.1.74), cellobiohydrolase (EC 3.2.1.91), xyloglucan-specific endo-β-1,4-glucanase/endo-xyloglucanase (EC 3.2.1.151), exo-β-glucosaminidase (EC 3.2.1.165), and endo-β-1,4-glucanase (xanthanase) (EC 3.2.1.-) activity (Fig. 6). The GH9 transcription level was significantly (Spearman P < 0.05) and negatively correlated with one group of NMR peaks including peaks expected to be caused by glucose molecules (Fig. 6) and positively correlated with a group of NMR peaks not related to glucose. The two groups of NMR peaks were detected in the polar fraction of 1-year-old litter and were correlated negatively between and positively within them (Fig. 6).
A partial least squares path model revealed that the NMR spectra of apolar chemical (i.e., chloroform extracts) extracts from litter are best explained by the corresponding NMR spectra of leaf chemistry (Fig. 7). This chemical fraction is only weakly linked to the other variables. The spectra of polar compounds (i.e., methanol extracts) of leaf and litter were not tightly linked, but explained together the composition of the DNA-derived fungal community well. Together, the total fungal community and the litter polar chemistry explained the composition of the RNA-derived community. Expression of litter-degrading CAZymes was moderately explained by NMR spectra of litter and composition of the RNA-derived fungal community.
Interconnection among the analyzed parameters. Directed graph of the partial least squares path model (PLS-PM). Direct and indirect effects of leaf and litter polar and apolar chemical composition, total (i.e., DNA-derived) and active (i.e., RNA-derived) litter fungal communities and respective transcription profiles of litter-degrading CAZyme families were jointly modeled. Positive (blue) and negative (red) effects among the considered latent variables are plotted
Discussion
The chemical compositions of leaves and litter are major drivers of fungal community assembly (Bray et al. 2012), while fungal exoenzymes change chemical composition of the litter (Liers et al. 2011). Expression and catalytic efficiency of litter-decomposing enzymes depend on the fungi active at a certain time (Li et al. 1999; Benoit et al. 2012). These represent a fraction of the whole fungal community detected in the litter (Baldrian et al. 2012; Peršoh 2015). In this study, we revealed a close interconnection between leaf and litter chemistry, fungal community composition, and their transcriptional activity (Fig. 1).
Total and active fungal communities are differently structured
Total and active communities often differ in composition (e.g., Fig. 2A), because not all detected fungi are metabolically active at a certain point in time (Baldrian et al. 2012; Peršoh 2015). Beyond that, we found that composition and CAZyme expression of the active communities differed at different spatial scales. The pool of expressed CAZyme-coding genes differed clearly among regions, while composition of the active fungal communities was not regionally structured (Fig. 2 and Fig. S3). This may be ascribed to different degrees of adaptation reflected by the CAZyme content and diversity (Rytioja et al. 2014; Sista Kameshwar and Qin 2017). While main litter decomposers degrade plant material by expressing and releasing CAZymes (Barbi et al. 2020), part of the metabolically active fungal community may not produce CAZymes (Fig. 2A). This may apply for less specialized fungi, such as secondary decomposers and “cheaters,” which also inhabit the litter but are not involved in primary degradation (Frankland 1998; Dighton 2007; van der Wal et al. 2013). They take advantage of products from the initial degradation or of other community-generated resources, respectively (see Velicer 2003). They are metabolically highly active while consuming readily available and easily accessible compounds, without producing notable amounts of CAZymes on their own. Our findings would be compatible with the hypothesis that main decomposers are structured by regional factors, and other litter-inhabiting fungi by local factors. This is further supported by the fact that the active community of non-Basidiomycota fungi differs significantly among regions. Early stages of decomposition are usually dominated by Ascomycetes, while Basidiomycetes dominate late stages (Schneider et al. 2012; Kuramae et al. 2013; Voříšková and Baldrian 2013; Bonanomi et al. 2019). In our study, the Agaricales (Basidiomycota) were the most active fungal order in decomposing leaf litter, but all taxa of Ascomycota together were roughly twice as abundant as Basidiomycota (Table 1). The different structures of active communities, total communities, and transcripts (Table 1, Figs. 2, S2, and S3) might therefore indicate the initial stage of a community shift from Ascomycota towards Basidiomycota. However, studies including chronologically higher resolved measurements will be required to reveal details of this potential transition phase.
Decomposer communities are functionally redundant
Metatranscriptome sequencing of litter communities provides insights into the activity of the entire community, but it has also the potential to place individual members of the community in context (Kuske et al. 2015). However, low availability of fungal genome data is a limitation in assigning eukaryotic transcripts to fungal taxa (Schneider et al. 2021). By applying a protocol to enrich the poly(A) in mRNA, we reduced possible contamination by non-eukaryotic transcripts in our data.
Extracellular enzymes produced by fungi are the major agents of litter decomposition in forest ecosystems (Baldrian et al. 2012; de Boer et al. 2005; Stursová et al. 2012; Žifčáková et al. 2016). Even though secreted by differently composed fungal communities, the enzymes are functionally redundant with regard to the degradation of the major organic compounds (Talbot et al. 2014). Samples from the three analyzed geographical regions were indeed not separable according to the CAZyme transcription profiles (Fig. 2), while composition of the diverse transcripts encoding for each CAZyme family clearly differed among regions. Interspecific sequence variation in the transcripts encoding the same enzymes (Table S4) (Aspeborg et al. 2012; Zhao et al. 2013) is likely to cause this, as taxonomic composition of the fungal communities was also regionally distinct (Fig. 2). Functional redundancy of decomposer communities is accordingly visible on the transcriptional level.
Producing the same CAZymes but with different substrate specificities and efficiencies (Collins et al. 2005; Benoit et al. 2012; Lombard et al. 2014; Wang et al. 2014), fungi with regional preferences might cause the apparent discrepancy between functional redundancy (Fig. 2) and convergence of substrate chemistry (Fig. 5); i.e., initially distinct leaf chemistry converges with time as discussed below (see the “Leaf chemical compositions reflect host origin and converge during decomposition” section), despite transcription of the same CAZymes. Although litter-decomposing fungal communities are functionally redundant (Fig. 2), these results suggest that litter communities are better adapted to their native litter (i.e., “Home-field advantage”) (Gholz et al. 2000; Prescott and Grayston 2013), while being additionally adapted to local factors (Talbot et al. 2014; Guerreiro et al. 2018). In addition, regionally different soil properties might affect persistence and activity of secreted CAZymes in the extracellular environment (Allison and Vitousek 2005; DeForest 2009). Even though these interactions are not known in detail, it is reasonable to assume that transcriptional diversity within functional fungal guilds is important for maintaining ecosystem services under varying conditions (Hooper et al. 2005; Folke 2006; Allison and Martiny 2008). By performing differently under different environmental conditions, different species of the same functional guild assure sustained overall ecosystem functioning within a certain range of conditions (Hooper et al. 2005). This resilience appears to be extended by forest management, as indicated by the higher diversity of transcripts per CAZyme in managed compared to natural forests (Fig. 3).
Leaf chemical compositions reflect host origin and converge during decomposition
Easily accessible soluble compounds, such as sugars, are quickly consumed by the decomposer organisms, followed by degradation of more complex largely insoluble compounds, such as (hemi)cellulose and lignin (Berg and McClaugherty 2014). These dynamic changes have been observed by the decrease of relative labile C compounds and respective relative increase of lignin throughout decomposition time (Purahong et al. 2014a; Bonanomi et al. 2019). Chemical composition of leaf litter thus changes with time. In our study, such changes were unclear (Fig. 4) or inconsistent (Fig. S9) across locations in the complete unfractionated material by solid-state NMR spectroscopy. According to the higher-resolved liquid NMR spectra, the decomposition process entailed a shift from leaves towards chemically distinct composed litter (Figs. S10 and S11). The richness of regionally specific compounds decreased with time, suggesting that leaves are chemically more complex than litter (Fig. S10). However, many compounds were shared between freshly fallen and 1-year-old decaying leaves. This overlap may be due to undegraded or only partly degraded soluble compounds, suggesting that decomposition of the soluble compounds was not completed within 1 year and still ongoing (Fig. S11).
Environmental conditions or abiotic stress influence the metabolome of the host and accordingly the quality of its litter (Bharti et al. 2011). This is supported by our findings, indicating that the overall chemical composition of leaves differed between geographic regions (Figs. 5 and S13), while the content of glucose, representing an omnipresent chemical compound, was regionally more distinct in litter than in leaves (Fig. S18). Plants change their phytochemistry and physiology to adapt to the surrounding conditions, which will be reflected by the plant metabolome (Shirouzu et al. 2009; Bharti et al. 2011). However, the mechanisms behind this adaptation process remain unclear. After senescence, chemical compounds derived from environmental adaptations are preferably degraded in earlier stages (Preston et al. 2009; Wallenstein et al. 2013). This is supported by our findings, where a large part of regionally discriminative chemical compounds in leaves were degraded during early decomposition (Fig. S13). Consequently, we detected a convergence of the chemical composition of leaf litter during decomposition across the different regions (Fig. 5).
Complex chemical compounds may become fragmented into smaller molecules during early decomposition (Šnajdr et al. 2011). The functional groups detected by 1H-NMR in these fragments may largely correspond to those in the original substrate (Figs. S12 and S14). For example, degradation of polysaccharides, such as cellulose, results in monosaccharide units (de Souza, 2013), in which most of the hydrogen bounds detected by 1H-NMR remain unmodified. On the other hand, enzymatic digestion may also generate products with a polarity differing from that of the original substrate. Accordingly, reactant and product would be detectable in different solvents. Such a process is known, for example, from the lipolysis pathway, in which fatty acids (apolar compounds) are converted into citric acid (polar compounds) (Nelson and Cox 2013; Haynes 2014). Such processes would result in a decrease in concentration in the apolar (chloroform soluble) fraction and an increase in signals in the polar (methanol soluble) fraction, or vice versa. It is reasonable to assume that, in agreement with our results (e.g., Fig. 5A), primarily apolar compounds are degraded into simpler water-soluble polar compounds during early stages of decomposition and become more easily absorbable by fungi (Berg and McClaugherty 2014). The temporal shifts in chemical composition of leaf litter observed in our pioneer study are therefore largely in agreement with the current general concept of the decomposition process.
Linking fungal activity, enzyme transcription, and chemical shifts
The substrate chemistry regulates the transcription of extracellular enzymes (Allison et al. 2011). However, the secreted enzymes will change the chemistry of the litter (Šnajdr et al. 2011), generating a complex regulatory feedback loop, as indicated by the direct linkage between litter chemistry and CAZyme expression levels (Fig. 7). Furthermore, composition of the active part of the fungal litter community is directly linked to transcription profiles of CAZyme families (Fig. 7). These direct linkages represent immediate effects, which are complemented by indirect, mostly long-term effects: Leaf chemistry affects composition of the total fungal community and thereby indirectly affecting the active fungal community. The indirect linkage appears reasonable, because the original, regionally discriminative leaf compounds have already been degraded at the time of litter sampling (Fig. 5). The leaf chemistry and CAZyme family transcription profiles are therefore supposed to be directly linked only in the very early stages of decomposition, i.e., before regionally discriminative compounds are degraded. In that early stage, leaf chemistry probably had a strong impact on assembly of the initial community (Šnajdr et al. 2011) and thereby influenced succession towards current litter community composition via priority effects (Cline and Zak 2015; Hiscox et al. 2015). The current composition and transcription profiles of the fungal communities are probably as much influenced by chemical changes and species successions in the past, as by the currently prevailing conditions.
Combining metametabolomics, metabarcoding, and metatranscriptomics approaches revealed to be a promising holistic approach to reveal interconnections between substrate chemistry, fungal community composition, and their transcriptional activity (Fig. 1). Correlative analyses may eventually link species to enzyme transcription and specific chemical shifts in litter (Fig. 6). In our study, Helotiales, Agaricales, and Pleosporales were the most active and diverse fungal groups in 1-year-old litter (Table 1). Furthermore, OTUs assigned to the genera Mycena and Chalara correlated directly with the activity of litter-degrading CAZymes (Fig. 6). Chalara species live mostly as litter saprotrophs (Koukol 2011), while several Mycena species are reported to switch between saprotrophic and biotrophic lifestyles (Thoen et al. 2020). These results further highlight the robustness of our approach to link fungal identity with ecological processes. Nevertheless, some of the OTUs correlating with litter-degrading CAZymes were assigned to ecologically ambiguous taxa, such as Mycosphaerella, a group of fungi known to include saprobes but also plant pathogens (Crous et al. 2001). This is due to the known limitations in taxonomic assignment of environmental OTUs in current metabarcoding approaches (Iquebal et al. 2021). The application of long-read sequencing approaches might improve the accuracy of taxonomic assignment in the medium term (Heeger et al. 2018; Jamy et al. 2020; Tedersoo et al. 2020). In the long term, increasing availability of genome data might allow to directly link CAZymes to fungal species. This will provide new insights into fungal ecology based on the role of CAZyme functioning in environmental processes.
Suitable spatial and temporal scales are a prerequisite for a conclusive picture, because the processes involved in decomposition, such as community assembly, fungal responses, enzyme activity, and chemical conversion, are closely interconnected (Fig. 1B) but occur at different time scales. While assessment of most major components (i.e., metagenome, proteome, and transcriptome) involved in decomposition is now well established microbial ecology (Franzosa et al. 2015; Peršoh 2015; Rastogi and Sani 2011), holistic assessment of the chemical properties and dynamics in the substrate is usually missing. Intensive monitoring is therefore required to causally link community succession, respective activity and changes in chemical composition. By enclosing fresh leaf litter in mesh bags and incubating them in the forest ground, litter bags represent a method close to natural conditions, while allowing accurate dating, high replication, and subsampling over time. However, litter bags are still artificial; they represent disturbances in the microclimate, might exclude macroinvertebrates from the process, and do not allow to determine losses by leaching (Karberg et al. 2008). Hypotheses arising from such experimental approaches should therefore be verified by analyzing natural litter. We were able to show that 1H-NMR spectroscopy provides meaningful fingerprints analyzable in context with other meta’omics data. Assigning additional NMR peaks to chemical compounds by spiking experiments (Fig. S17) in future studies will provide valuable reference data which will massively enhance the value of environmental 1H-NMR spectra.
Conclusions
In this study, we applied for the first time environmental NMR fingerprinting to infer ecological contexts from chemical substrate composition analyzed concertedly with metatranscriptomic and community metabarcoding data. The NMR spectra successfully distinguished the chemical composition of freshly fallen leaves from that of 1-year-old litter. We determined that chemical composition of leaves converges with progressing decomposition, while litter decomposing fungal communities were functionally redundant according to the expression of genes encoding litter-degrading enzymes (i.e., carbohydrate-active enzymes (CAZymes)). However, forestry measures influenced the diversity of transcribed litter-decomposing CAZymes. Since functional resilience of decomposer communities likely depends on transcript diversity, the findings are of major relevance for environmental ecology and applied forestry. Environmental NMR fingerprinting proved suitable for inferring ecological contexts, because the CAZyme expression profiles in litter could be shown to be influenced directly by litter and indirectly by leaf chemistry. The comprehensive and quantitative information provided NMR spectra was even accessible by spiking chemical compounds of interest. Altogether, we provide a conceptual framework that combines metametabolomics, metabarcoding, and metatranscriptomics approaches with the potential to unveil mechanistic interactions between substrate chemistry, fungal community composition, and their transcriptional activity. Determining how generalizable these findings are is crucial for future ecological studies in decomposition. Experimental incubations with mock communities, considering single or multiple species, will help determine the role of individuals within the communities and their influence on the decomposition process.
References
Albright MBN, Thompson J, Kroeger ME, Johansen R, Ulrich DEM, La Gallegos-Graves V, Munsky B, Dunbar J (2020) Differences in substrate use linked to divergent carbon flow during litter decomposition. FEMS Microbiol Ecol 96. https://doi.org/10.1093/femsec/fiaa135
Allison SD, Martiny JBH (2008) Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci USA 105(Suppl 1):11512–11519. https://doi.org/10.1073/pnas.0801925105
Allison SD, Vitousek PM (2005) Responses of extracellular enzymes to simple and complex nutrient inputs. Soil Biol Biochem 37:937–944. https://doi.org/10.1016/j.soilbio.2004.09.014
Allison SD, Weintraub MN, Gartner TB, Waldrop MP (2011) Evolutionary-economic principles as regulators of soil enzyme production and ecosystem function. In: Shukla G, Varma A (eds) Soil Enzymology, vol 22. Springer Berlin Heidelberg, Berlin, Heidelberg, pp 229–243. https://doi.org/10.1007/978-3-642-14225-3_12
Anderson IC, Parkin PI (2007) Detection of active soil fungi by RT-PCR amplification of precursor rRNA molecules. J Microbiol Methods 68:248–253. https://doi.org/10.1016/j.mimet.2006.08.005
Aspeborg H, Coutinho PM, Wang Y, Brumer H, Henrissat B (2012) Evolution, substrate specificity and subfamily classification of glycoside hydrolase family 5 (GH5). BMC Evol Biol 12:186. https://doi.org/10.1186/1471-2148-12-186
Baldrian P, Kolařík M, Stursová M, Kopecký J, Valášková V, Větrovský T, Zifčáková L, Snajdr J, Rídl J, Vlček C, Voříšková J (2012) Active and total microbial communities in forest soil are largely different and highly stratified during decomposition. ISME J 6:248–258. https://doi.org/10.1038/ismej.2011.95
Barbi F, Kohler A, Barry K, Baskaran P, Daum C, Fauchery L, Ihrmark K, Kuo A, LaButti K, Lipzen A, Morin E, Grigoriev IV, Henrissat B, Lindahl BD, Martin F (2020) Fungal ecological strategies reflected in gene transcription - a case study of two litter decomposers. Environ Microbiol 22:1089–1103. https://doi.org/10.1111/1462-2920.14873
Benoit I, Coutinho PM, Schols HA, Gerlach JP, Henrissat B, de Vries RP (2012) Degradation of different pectins by fungi. Correlations and contrasts between the pectinolytic enzyme sets identified in genomes and the growth on pectins of different origin. BMC Genomics 13:321. https://doi.org/10.1186/1471-2164-13-321
Berg B, McClaugherty C (2014) Plant litter. Decomposition, humus formation, carbon sequestration, Third edition. Springer, Heidelberg. https://doi.org/10.1007/978-3-642-38821-7
Bharti SK, Bhatia A, Tewari SK, Sidhu OP, Roy R (2011) Application of HR-MAS NMR spectroscopy for studying chemotype variations of Withania somnifera (L.) Dunal. Magn Reson Chem 49:659–667. https://doi.org/10.1002/mrc.2817
Bonanomi G, de Filippis F, Cesarano G, La Storia A, Zotti M, Mazzoleni S, Incerti G (2019) Linking bacterial and eukaryotic microbiota to litter chemistry. Combining next generation sequencing with 13C CPMAS NMR spectroscopy. Soil Biol Biochem 129:110–121. https://doi.org/10.1016/j.soilbio.2018.11.013
Bradford MA, Berg B, Maynard DS, Wieder WR, Wood SA, Cornwell W (2016) Understanding the dominant controls on litter decomposition. J Ecol 104:229–238. https://doi.org/10.1111/1365-2745.12507
Bray JR, Curtis JT (1957) An ordination of the upland forest communities of Southern Wisconsin. Ecol Monogr 27:325–349. https://doi.org/10.2307/1942268
Bray SR, Kitajima K, Mack MC (2012) Temporal dynamics of microbial communities on decomposing leaf litter of 10 plant species in relation to decomposition rate. Soil Biol Biochem 49:30–37. https://doi.org/10.1016/j.soilbio.2012.02.009
Burns RG, DeForest JL, Marxsen J, Sinsabaugh RL, Stromberger ME, Wallenstein MD, Weintraub MN, Zoppini A (2013) Soil enzymes in a changing environment. Current knowledge and future directions. Soil Biol Biochem 58:216–234. https://doi.org/10.1016/j.soilbio.2012.11.009
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Clarke KR, Gorley RN (2015) PRIMER v7: user manual/tutorial. PRIMER-E, Plymouth, 296pp
Cline LC, Zak DR (2015) Initial colonization, community assembly and ecosystem function. Fungal colonist traits and litter biochemistry mediate decay rate. Mol Ecol 24:5045–5058. https://doi.org/10.1111/mec.13361
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23. https://doi.org/10.1016/j.femsre.2004.06.005
Crous PW, Kang J-C, Braun U (2001) A phylogenetic redefinition of anamorph genera in Mycosphaerella based on ITS rDNA sequence and morphology. Mycologia 93:1081–1101. https://doi.org/10.1080/00275514.2001.12063243
Davidson EA, Janssens IA (2006) Temperature sensitivity of soil carbon decomposition and feedbacks to climate change. Nature 440:165–173. https://doi.org/10.1038/nature04514
de Boer W, Folman LB, Summerbell RC, Boddy L (2005) Living in a fungal world. Impact of fungi on soil bacterial niche development. FEMS Microbiol Rev 29:795–811. https://doi.org/10.1016/j.femsre.2004.11.005
DeForest JL (2009) The influence of time, storage temperature, and substrate age on potential soil enzyme activity in acidic forest soils using MUB-linked substrates and l-DOPA. Soil Biol Biochem 41:1180–1186. https://doi.org/10.1016/j.soilbio.2009.02.029
Dence CW (1992) The Determination of Lignin. In: Timell TE, Lin SY, Dence CW (eds) Methods in lignin chemistry. Springer Berlin Heidelberg, Berlin, Heidelberg, pp 33–61. https://doi.org/10.1007/978-3-642-74065-7_3
Dighton J (2007) Nutrient cycling by saprotrophic fungi in terrestrial habitats. In: Kubicek CP, Druzhinina IS (eds) The Mycota. A comprehensive treatise on fungi as experimental systems for basic and applied research, 2nd ed., vol 4. Springer Berlin, Berlin, pp 287–300. https://doi.org/10.1007/978-3-540-71840-6_16
Dunn WB, Bailey NJC, Johnson HE (2005) Measuring the metabolome. Current analytical technologies. Analyst 130:606–625. https://doi.org/10.1039/b418288j
Dunn WB, Ellis DI (2005) Metabolomics. Current analytical platforms and methodologies. Trends Analyt Chem 24:285–294. https://doi.org/10.1016/j.trac.2004.11.021
Eichlerová I, Homolka L, Žifčáková L, Lisá L, Dobiášová P, Baldrian P (2015) Enzymatic systems involved in decomposition reflects the ecology and taxonomy of saprotrophic fungi. Fungal Ecol 13:10–22. https://doi.org/10.1016/j.funeco.2014.08.002
Fiehn O (2002) Metabolomics – the link between genotypes and phenotypes. Plant Mol Biol 48:155–171. https://doi.org/10.1023/A:1013713905833
Fischer M, Bossdorf O, Gockel S, Hänsel F, Hemp A, Hessenmöller D, Korte G, Nieschulze J, Pfeiffer S, Prati D, Renner S, Schöning I, Schumacher U, Wells K, Buscot F, Kalko EKV, Linsenmair KE, Schulze E-D, Weisser WW (2010) Implementing large-scale and long-term functional biodiversity research. The Biodiversity Exploratories. Basic Appl Ecol 11:473–485. https://doi.org/10.1016/j.baae.2010.07.009
Folke C (2006) Resilience. The emergence of a perspective for social–ecological systems analyses. Glob Environ Chang 16:253–267. https://doi.org/10.1016/j.gloenvcha.2006.04.002
Frankland JC (1998) Fungal succession — unravelling the unpredictable. Mycol Res 102:1–15. https://doi.org/10.1017/S0953756297005364
Franzosa EA, Hsu T, Sirota-Madi A, Shafquat A, Abu-Ali G, Morgan XC, Huttenhower C (2015) Sequencing and beyond. Integrating molecular 'omics' for microbial community profiling. Nat Rev Microbiol 13:360–372. https://doi.org/10.1038/nrmicro3451
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT. Accelerated for clustering the next-generation sequencing data. Bioinform 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Fung BM, Khitrin AK, Ermolaev K (2000) An improved broadband decoupling sequence for liquid crystals and solids. J Magn Reson 142:97–101. https://doi.org/10.1006/jmre.1999.1896
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes--application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
Gholz HL, Wedin DA, Smitherman SM, Harmon ME, Parton WJ (2000) Long-term dynamics of pine and hardwood litter in contrasting environments. Toward a global model of decomposition. Glob Chang Biol 6:751–765. https://doi.org/10.1046/j.1365-2486.2000.00349.x
Gossner MM, Beenken L, Arend K, Begerow D, Peršoh D (2021) Insect herbivory facilitates the establishment of an invasive plant pathogen. ISME Commun 1:492. https://doi.org/10.1038/s43705-021-00004-4
Guerreiro MA, Brachmann A, Begerow D, Peršoh D (2018) Transient leaf endophytes are the most active fungi in 1-year-old beech leaf litter. Fungal Divers 89:237–251. https://doi.org/10.1007/s13225-017-0390-4
Hamilton N (2018) ggtern. : an extension to 'ggplot2', for the Creation of Ternary Diagrams. R package version 2.2.2
Haynes WM (2014) CRC handbook of chemistry and physics. A ready-reference book of chemical and physical data, 95th edition / edited by William M. Haynes. 100 Key Points. CRC Press, Boca Raton
Heeger F, Bourne EC, Baschien C, Yurkov A, Bunk B, Spröer C, Overmann J, Mazzoni CJ, Monaghan MT (2018) Long-read DNA metabarcoding of ribosomal RNA in the analysis of fungi from aquatic environments. Mol Ecol Resour 18:1500–1514. https://doi.org/10.1111/1755-0998.12937
Hiscox J, Savoury M, Müller CT, Lindahl BD, Rogers HJ, Boddy L (2015) Priority effects during fungal community establishment in beech wood. ISME J 9:2246–2260. https://doi.org/10.1038/ismej.2015.38
Holmes E, Tang H, Wang Y, Seger C (2006) The assessment of plant metabolite profiles by NMR-based methodologies. Planta Med 72:771–785. https://doi.org/10.1055/s-2006-946682
Hooper DU, Chapin ES III, Ewel JJ, Hector A, Inchausti P, Lavorel S, Lawton JH, Lodge DM, Loreau M, Naeem S, Schmid B, Setala H, Symstad AJ, Vandermeer J, Wardle DA (2005) Effects of biodiversity on ecosystem functioning a consensus of current knowledge. Ecol Monogr 75:3–35
Hulsen T, de Vlieg J, Alkema W (2008) BioVenn - a web application for the comparison and visualization of biological lists using area-proportional Venn diagrams. BMC Genomics 9:488. https://doi.org/10.1186/1471-2164-9-488
Iquebal MA, Jaiswal S, Mishra VK, Jasrotia RS, Angadi UB, Singh BP, Passari AK, Deka P, Prabha R, Singh DP, Gupta VK, Tomar RS, Oberoi HS, Rai A, Kumar D (2021) Fungal Genomic resources for strain identification and diversity analysis of 1900 fungal species. J Fungi 7. https://doi.org/10.3390/jof7040288
Jamy M, Foster R, Barbera P, Czech L, Kozlov A, Stamatakis A, Bending G, Hilton S, Bass D, Burki F (2020) Long-read metabarcoding of the eukaryotic rDNA operon to phylogenetically and taxonomically resolve environmental diversity. Mol Ecol Resour 20:429–443. https://doi.org/10.1111/1755-0998.13117
Kameshwar AKS, Ramos LP, Qin W (2019) CAZymes-based ranking of fungi (CBRF). An interactive web database for identifying fungi with extrinsic plant biomass degrading abilities. Bioresour Bioprocess 6:1602. https://doi.org/10.1186/s40643-019-0286-0
Karberg NJ, Scott NA, Giardina CP (2008) Methods for estimating litter decomposition. In: Hoover CM (ed) Field measurements for forest carbon monitoring. A landscape-scale approach. Springer, Netherlands, pp 103–111
Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M, Bates ST, Bruns TD, Bengtsson-Palme J, Callaghan TM, Douglas B, Drenkhan T, Eberhardt U, Dueñas M, Grebenc T, Griffith GW, Hartmann M, Kirk PM, Kohout P et al (2013) Towards a unified paradigm for sequence-based identification of fungi. Mol Ecol 22:5271–5277. https://doi.org/10.1111/mec.12481
Koukol O (2011) New species of Chalara occupying coniferous needles. Fungal Divers 49:75–91. https://doi.org/10.1007/s13225-011-0092-2
Krishnan P, Kruger NJ, Ratcliffe RG (2005) Metabolite fingerprinting and profiling in plants using NMR. J Exp Bot 56:255–265. https://doi.org/10.1093/jxb/eri010
Kubicek CP, Messner R, Gruber F, Mach RL, Kubicek-Pranz EM (1993) The Trichoderma cellulase regulatory puzzle. From the interior life of a secretory fungus. Enzym Microb Technol 15:90–99. https://doi.org/10.1016/0141-0229(93)90030-6
Kuramae EE, Hillekens RHE, de Hollander M, van der Heijden MGA, van den Berg M, van Straalen NM, Kowalchuk GA (2013) Structural and functional variation in soil fungal communities associated with litter bags containing maize leaf. FEMS Microbiol Ecol 84:519–531. https://doi.org/10.1111/1574-6941.12080
Kuske CR, Hesse CN, Challacombe JF, Cullen D, Herr JR, Mueller RC, Tsang A, Vilgalys R (2015) Prospects and challenges for fungal metatranscriptomics of complex communities. Fungal Ecol 14:133–137. https://doi.org/10.1016/j.funeco.2014.12.005
Legendre P, Gallagher ED (2001) Ecologically meaningful transformations for ordination of species data. Oecologia 129:271–280. https://doi.org/10.1007/s004420100716
Li K, Xu F, Eriksson KE (1999) Comparison of fungal laccases and redox mediators in oxidation of a nonphenolic lignin model compound. Appl Environ Microbiol 65:2654–2660
Liers C, Arnstadt T, Ullrich R, Hofrichter M (2011) Patterns of lignin degradation and oxidative enzyme secretion by different wood- and litter-colonizing basidiomycetes and ascomycetes grown on beech-wood. FEMS Microbiol Ecol 78:91–102. https://doi.org/10.1111/j.1574-6941.2011.01144.x
Lladó S, López-Mondéjar R, Baldrian P (2017) Forest soil bacteria. Diversity, Involvement in Ecosystem Processes, and Response to Global Change. Microbiol Mol Biol Rev 81. https://doi.org/10.1128/MMBR.00063-16
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:D490–D495. https://doi.org/10.1093/nar/gkt1178
Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M (2007) KAAS. An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35:W182–W185. https://doi.org/10.1093/nar/gkm321
Nelson DL, Cox MM (2013) Lehninger principles of biochemistry, Sixth edition, international edition. W.H. Freeman and Company, New York
Osono T (2020) Functional diversity of ligninolytic fungi associated with leaf litter decomposition. Ecol Res 35:30–43. https://doi.org/10.1111/1440-1703.12063
Parton W, Silver WL, Burke IC, Grassens L, Harmon ME, Currie WS, King JY, Adair EC, Brandt LA, Hart SC, Fasth B (2007) Global-scale similarities in nitrogen release patterns during long-term decomposition. Science 315:361–364. https://doi.org/10.1126/science.1134853
Pauli GF, Jaki BU, Lankin DC (2005) Quantitative 1H NMR. Development and potential of a method for natural products analysis. J Nat Prod 68:133–149. https://doi.org/10.1021/np0497301
Paulson JN, Stine OC, Bravo HC, Pop M (2013) Differential abundance analysis for microbial marker-gene surveys. Nat Methods 10:1200–1202. https://doi.org/10.1038/nmeth.2658
Peršoh D (2013) Factors shaping community structure of endophytic fungi–evidence from the Pinus-Viscum-system. Fungal Divers 60:55–69. https://doi.org/10.1007/s13225-013-0225-x
Peršoh D (2015) Plant-associated fungal communities in the light of meta’omics. Fungal Divers 75:1–25. https://doi.org/10.1007/s13225-015-0334-9
Peršoh D, Theuerl S, Buscot F, Rambold G (2008) Towards a universally adaptable method for quantitative extraction of high-purity nucleic acids from soil. J Microbiol Methods 75:19–24. https://doi.org/10.1016/j.mimet.2008.04.009
Persson T, Bååth E, Clarholm M, Lundkvist H, Söderström BE, Sohlenius B (1980) Trophic structure, biomass dynamics and carbon metabolism of soil organisms in a scots pine forest. Ecol Bull:419–459
Prescott CE, Grayston SJ (2013) Tree species influence on microbial communities in litter and soil. Current knowledge and research needs. For Ecol Manage 309:19–27. https://doi.org/10.1016/j.foreco.2013.02.034
Preston CM, Nault JR, Trofymow JA (2009) Chemical changes during 6 years of decomposition of 11 litters in some Canadian forest sites. Part 2. 13C Abundance, Solid-State 13C NMR Spectroscopy and the Meaning of “Lignin”. Ecosystems 12:1078–1102. https://doi.org/10.1007/s10021-009-9267-z
Pretsch E, Bühlmann P, Badertscher M (2009) Structure determination of organic compounds. Tables of spectral data, 4th, rev. and enl. ed. Springer, Berlin. https://doi.org/10.1007/978-3-540-93810-1
Purahong W, Kapturska D, Pecyna MJ, Jariyavidyanont K, Kaunzner J, Juncheed K, Uengwetwanit T, Rudloff R, Schulz E, Hofrichter M, Schloter M, Krüger D, Buscot F (2015) Effects of forest management practices in temperate beech forests on bacterial and fungal communities involved in leaf litter degradation. Microb Ecol 69:905–913. https://doi.org/10.1007/s00248-015-0585-8
Purahong W, Kapturska D, Pecyna MJ, Schulz E, Schloter M, Buscot F, Hofrichter M, Krüger D (2014a) Influence of different forest system management practices on leaf litter decomposition rates, nutrient dynamics and the activity of ligninolytic enzymes. A case study from central European forests. PLoS ONE 9:e93700. https://doi.org/10.1371/JOURNAL.PONE.0093700
Purahong W, Schloter M, Pecyna MJ, Kapturska D, Däumlich V, Mital S, Buscot F, Hofrichter M, Gutknecht JLM, Krüger D (2014b) Uncoupling of microbial community structure and function in decomposing litter across beech forest ecosystems in Central Europe. Sci Rep 4:7014. https://doi.org/10.1038/srep07014
Purahong W, Wubet T, Lentendu G, Schloter M, Pecyna MJ, Kapturska D, Hofrichter M, Krüger D, Buscot F (2016) Life in leaf litter. Novel insights into community dynamics of bacteria and fungi during litter decomposition. Mol Ecol 25:4059–4074. https://doi.org/10.1111/mec.13739
R Core Team (2017) R. A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rastogi G, Sani RK (2011) Molecular techniques to assess microbial community structure, function, and dynamics in the environment. In: Ahmad I, Ahmad F, Pichtel J (eds) Microbes and Microbial Technology. Springer New York, New York, NY, pp 29–57. https://doi.org/10.1007/978-1-4419-7931-5_2
Revelle W (2017) psych: procedures for psychological, psychometric, and personality research. R package version 1.8.4
Röhl O, Peršoh D, Mittelbach M, Elbrecht V, Brachmann A, Nuy J, Boenigk J, Leese F, Begerow D (2017) Distinct sensitivity of fungal freshwater guilds to water quality. Mycol Prog 16:155–169. https://doi.org/10.1007/s11557-016-1261-1
Rytioja J, Hildén K, Yuzon J, Hatakka A, de Vries RP, Mäkelä MR (2014) Plant-polysaccharide-degrading enzymes from Basidiomycetes. Microbiol Mol Biol Rev 78:614–649. https://doi.org/10.1128/MMBR.00035-14
Sanchez G, Trinchera L, Russolillo G (2017) plspm. Tools for partial least squares path modeling (PLS-PM). R package version 0.4.9
Schneider AN, Sundh J, Sundström G, Richau K, Delhomme N, Grabherr M, Hurry V, Street NR (2021) Comparative fungal community analyses using metatranscriptomics and internal transcribed spacer amplicon sequencing from Norway spruce. mSystems 6. https://doi.org/10.1128/mSystems.00884-20
Schneider T, Gerrits B, Gassmann R, Schmid E, Gessner MO, Richter A, Battin T, Eberl L, Riedel K (2010) Proteome analysis of fungal and bacterial involvement in leaf litter decomposition. Proteomics 10:1819–1830. https://doi.org/10.1002/pmic.200900691
Schneider T, Keiblinger KM, Schmid E, Sterflinger-Gleixner K, Ellersdorfer G, Roschitzki B, Richter A, Eberl L, Zechmeister-Boltenstern S, Riedel K (2012) Who is who in litter decomposition? Metaproteomics reveals major microbial players and their biogeochemical functions. ISME J 6:1749–1762. https://doi.org/10.1038/ismej.2012.11
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA 109:6241–6246. https://doi.org/10.1073/pnas.1117018109
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape. A software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Shirouzu T, Hirose D, Fukasawa Y, Tokumasu S (2009) Fungal succession associated with the decay of leaves of an evergreen oak, Quercus myrsinaefolia. Fungal Divers 34:87–109
Sinclair L, Osman OA, Bertilsson S, Eiler A (2015) Microbial community composition and diversity via 16S rRNA gene amplicons. Evaluating the illumina platform. PLoS ONE 10:e0116955. https://doi.org/10.1371/journal.pone.0116955
Sinsabaugh RL, Antibus RK, Linkins AE (1991) An enzymic approach to the analysis of microbial activity during plant litter decomposition. Agric Ecosyst Environ 34:43–54. https://doi.org/10.1016/0167-8809(91)90092-C
Sista Kameshwar AK, Qin W (2017) Comparative study of genome-wide plant biomass-degrading CAZymes in white rot, brown rot and soft rot fungi. Mycology 60:1–13. https://doi.org/10.1080/21501203.2017.1419296
Šnajdr J, Cajthaml T, Valášková V, Merhautová V, Petránková M, Spetz P, Leppänen K, Baldrian P (2011) Transformation of Quercus petraea litter. Successive changes in litter chemistry are reflected in differential enzyme activity and changes in the microbial community composition. FEMS Microbiol Ecol 75:291–303. https://doi.org/10.1111/j.1574-6941.2010.00999.x
Souza WR de (2013) Microbial degradation of lignocellulosic biomass. In: Chandel A (ed) Sustainable Degradation of Lignocellulosic Biomass - Techniques, Applications and Commercialization. InTech. https://doi.org/10.5772/54325
Stielow JB, Lévesque CA, Seifert KA, Meyer W, Iriny L, Smits D, Renfurm R, Verkley GJM, Groenewald M, Chaduli D, Lomascolo A, Welti S, Lesage-Meessen L, Favel A, Al-Hatmi AMS, Damm U, Yilmaz N, Houbraken J, Lombard L et al (2015) One fungus, which genes? Development and assessment of universal primers for potential secondary fungal DNA barcodes. Persoonia 35:242–263. https://doi.org/10.3767/003158515X689135
Stursová M, Zifčáková L, Leigh MB, Burgess R, Baldrian P (2012) Cellulose utilization in forest litter and soil. Identification of bacterial and fungal decomposers. FEMS Microbiol Ecol 80:735–746. https://doi.org/10.1111/j.1574-6941.2012.01343.x
Talbot JM, Bruns TD, Taylor JW, Smith DP, Branco S, Glassman SI, Erlandson S, Vilgalys R, Liao H-L, Smith ME, Peay KG (2014) Endemism and functional convergence across the North American soil mycobiome. Proc Natl Acad Sci USA 111:6341–6346. https://doi.org/10.1073/pnas.1402584111
Talbot JM, Martin F, Kohler A, Henrissat B, Peay KG (2015) Functional guild classification predicts the enzymatic role of fungi in litter and soil biogeochemistry. Soil Biol Biochem 88:441–456. https://doi.org/10.1016/j.soilbio.2015.05.006
Tedersoo L, Anslan S, Bahram M, Kõljalg U, Abarenkov K (2020) Identifying the ‘unidentified’ fungi. A global-scale long-read third-generation sequencing approach. Fungal Divers 103:273–293. https://doi.org/10.1007/s13225-020-00456-4
Thoen E, Harder CB, Kauserud H, Botnen SS, Vik U, Taylor AFS, Menkis A, Skrede I (2020) In vitro evidence of root colonization suggests ecological versatility in the genus Mycena. New Phytol 227:601–612. https://doi.org/10.1111/nph.16545
van der Wal A, Geydan TD, Kuyper TW, Wd B (2013) A thready affair. Linking fungal diversity and community dynamics to terrestrial decomposition processes. FEMS Microbiol Rev 37:477–494. https://doi.org/10.1111/1574-6976.12001
Velicer GJ (2003) Social strife in the microbial world. Trends Microbiol 11:330–337. https://doi.org/10.1016/S0966-842X(03)00152-5
Voříšková J, Baldrian P (2013) Fungal community on decomposing leaf litter undergoes rapid successional changes. ISME J 7:477–486. https://doi.org/10.1038/ismej.2012.116
Wallenstein MD, Haddix ML, Ayres E, Steltzer H, Magrini-Bair KA, Paul EA (2013) Litter chemistry changes more rapidly when decomposed at home but converges during decomposition–transformation. Soil Biol Biochem 57:311–319. https://doi.org/10.1016/j.soilbio.2012.09.027
Wang W, Li Y, Wang H, Zu Y (2014) Differences in the activities of eight enzymes from ten soil fungi and their possible influences on the surface structure, functional groups, and element composition of soil colloids. PLoS ONE 9:e111740. https://doi.org/10.1371/journal.pone.0111740
Weiss S, Xu ZZ, Peddada S, Amir A, Bittinger K, Gonzalez A, Lozupone C, Zaneveld JR, Vázquez-Baeza Y, Birmingham A, Hyde ER, Knight R (2017) Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5:27. https://doi.org/10.1186/s40168-017-0237-y
White T, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Gelfand D, Shinsky J, White T (eds) Innis M. A Guide to Methods and Applications. Academic Press, PCR Protocols, pp 315–322
Wickham H (2009) ggplot2. Elegant graphics for data analysis. R package version 2.2.1. Springer-Verlag New York
Wishart DS (2008) Quantitative metabolomics using NMR. Trends Analyt Chem 27:228–237. https://doi.org/10.1016/j.trac.2007.12.001
Wold H (1975) Path Models with Latent Variables. The NIPALS Approach. In: Blalock HM (ed) Quantitative sociology. International perspectives on mathematical and statistical modeling. Academic Press, New York, London, pp 307–357. https://doi.org/10.1016/B978-0-12-103950-9.50017-4
Wolfe BE, Tulloss RE, Pringle A (2012) The irreversible loss of a decomposition pathway marks the single origin of an ectomycorrhizal symbiosis. PLoS ONE 7:e39597. https://doi.org/10.1371/journal.pone.0039597
Xiao W, Chen C, Chen X, Huang Z, Chen HYH, Soininen J (2020) Functional and phylogenetic diversity promote litter decomposition across terrestrial ecosystems. Glob Ecol Biogeogr 29:2261–2272. https://doi.org/10.1111/geb.13181
Zhang H, Yohe T, Huang L, Entwistle S, Wu P, Yang Z, Busk PK, Xu Y, Yin Y (2018) dbCAN2. A meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 46:W95–W101. https://doi.org/10.1093/nar/gky418
Zhao Z, Liu H, Wang C, Xu J-R (2013) Comparative analysis of fungal genomes reveals different plant cell wall degrading capacity in fungi. BMC Genomics 14:274. https://doi.org/10.1186/1471-2164-14-274
Žifčáková L, Větrovský T, Howe A, Baldrian P (2016) Microbial activity in forest soil reflects the changes in ecosystem properties between summer and winter. Environ Microbiol 18:288–301. https://doi.org/10.1111/1462-2920.13026
Žifčáková L, Větrovský T, Lombard V, Henrissat B, Howe A, Baldrian P (2017) Feed in summer, rest in winter. Microbial carbon utilization in forest topsoil. Microbiome 5:122. https://doi.org/10.1186/s40168-017-0340-0
Acknowledgements
We thank all the members of the Geobotany working group (Ruhr-Universität Bochum) for their assistance during sample collection, nucleic acid extractions, and library preparation. We are grateful to Martin Gartmann for his NMR expertise and technical help. We also thank the managers of the three Exploratories, and all former managers for their work in maintaining the plot and project infrastructure, for giving support through the central office, for managing the central data base, and Markus Fischer, Eduard Linsenmair, Dominik Hessenmöller, Daniel Prati, Ingo Schöning, François Buscot, Ernst-Detlef Schulze, Wolfgang W. Weisser, and the late Elisabeth Kalko for their role in setting up the Biodiversity Exploratories project. We thank the administration of the Hainich national park, the UNESCO Biosphere Reserve Swabian Alb, and the UNESCO Biosphere Reserve Schorfheide-Chorin as well as all land owners for the excellent collaboration. The work has been (partly) funded by the DFG Priority Program 1374 “Biodiversity-Exploratories” (BE 2201/16-1 and PE 1673/5-1). RS gratefully acknowledges support from the DFG (grant nos. INST 213/757-1 FUGG and INST 213/843-1 FUGG). Field work permits were issued by the responsible state environmental offices of Baden-Württemberg, Thüringen, and Brandenburg (according to § 72 BbgNatSchG).
Availability of data and material
The sequence data are available from the European Nucleotide Archive (http://www.ebi.ac.uk/ena/) under study PRJEB19870 and from the EBI Metagenomics database (https://www.ebi.ac.uk/metagenomics) under studies PRJEB7861 and PRJEB8419. Raw 1H-NMR spectra were archived in Open Science Framework (DOI 10.17605/OSF.IO/2TJC3; https://osf.io/2tjc3/?view_only=593854b6f52f45ebbe285374051a0d52). Corresponding metadata are provided in Tables S2, S3, and S7. All data generated or analyzed during this study are included in this published article and its supplementary information files.
Funding
Open Access funding enabled and organized by Projekt DEAL. The work has been (partly) funded by the DFG Priority Program 1374 “Biodiversity-Exploratories” (BE 2201/16-1 and PE 1673/5-1). RS gratefully acknowledges support from the DFG (grant nos. INST 213/757-1 FUGG and INST 213/843-1 FUGG).
Author information
Authors and Affiliations
Contributions
The study was designed by DB and DP; DP, MG, and student helpers performed the sampling; MG conducted the laboratory work; AB supported preparation of the amplicon and mRNA libraries and performed the sequencing; RS and JS supported and performed the solvent and solid-state NMR measurements, respectively; SK developed the R scripts; DP and MG analyzed the data and wrote the manuscript. All authors discussed and contributed to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Section Editor: Andrey M Yurkov
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guerreiro, M.A., Kambach, S., Stoll, R. et al. Linking processes to community functions—insights into litter decomposition combining fungal metatranscriptomics and environmental NMR profiling. Mycol Progress 22, 10 (2023). https://doi.org/10.1007/s11557-022-01859-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-022-01859-0