Abstract
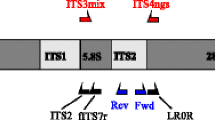
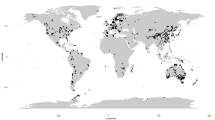
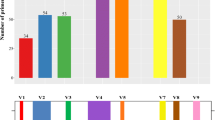
Molecular identification methods, in particular high-throughput sequencing tools, have greatly improved our knowledge about fungal diversity and biogeography, but many of the recovered taxa from natural environments cannot be identified to species or even higher taxonomic levels. This study addresses the phylogenetic placement of previously unrecognized fungal groups by using two complementary approaches: (i) third-generation amplicon sequencing analysis of DNA from global soil samples, screening out ITS reads of < 90% similarity to other available Sanger sequences, and (ii) analysis of common fungal taxa that were previously indicated to be enigmatic in terms of taxonomic placement based on the ITS sequences alone (so-called top50 sequences). For the global soil samples, we chose to amplify the full rRNA gene operon using four partly overlapping amplicons and multiple newly developed primers or primer combinations that cover nearly all fungi and a vast majority of non-fungal eukaryotes. We extracted the rRNA 18S (SSU) and 28S (LSU) genes and performed phylogenetic analyses against carefully selected reference material. Both SSU and LSU analyses placed most soil sequences and top50 sequences to known orders and classes, but tens of monophyletic groups and single sequences remained outside described taxa. Furthermore, the LSU analyses recovered a few small groups of sequences that may potentially represent novel phyla. We conclude that rRNA genes-based phylogenetic analyses are efficient tools for determining phylogenetic relationships of fungal taxa that cannot be placed to any order or class using ITS sequences alone. However, in many instances, longer rRNA gene sequences and availability of both SSU and LSU reads are needed to improve taxonomic resolution. By leveraging third-generation sequencing from global soil samples, we successfully provided phylogenetic placement for many previously unidentified sequences and broadened our view on the fungal tree of life, with 10–20% new order-level taxa. In addition, the PacBio sequence data greatly extends fungal class-level information in reference databases.









Similar content being viewed by others
References
Ahrendt SR, Quandt CA, Ciobanu D, Clum A, Salamov A, Andreopoulos B, Cheng JF, Woyke T, Pelin A, Henrissat B, Reynolds NK (2018) Leveraging single-cell genomics to expand the fungal tree of life. Nat Microbiol 3:1417
Anslan S, Bahram M, Hiiesalu I, Tedersoo L (2017) PipeCraft: flexible open-source toolkit for bioinformatics analysis of custom high-throughput amplicon sequencing data. Mol Ecol Res 17:e234–e240
Ardila-Garcia AM, Raghuram N, Sihota P, Fast NM (2013) Microsporidian diversity in soil, sand, and compost of the Pacific Northwest. J Euk Microbiol 60:601–608
Arnold AE (2007) Understanding the diversity of foliar endophytic fungi: progress, challenges, and frontiers. Fung Biol Rev 21:51–66
Bass D, Czech L, Williams BA, Berney C, Dunthorn M, Mahé F, Torruella G, Stentiford GD, Williams TA (2018) Clarifying the relationships between Microsporidia and Cryptomycota. J Euk Microbiol 65:773–782
Bauer R, Garnica S, Oberwinkler F, Riess K, Weiß M, Begerow D (2015) Entorrhizomycota: a new fungal phylum reveals new perspectives on the evolution of Fungi. PLoS ONE 10:e0128183
Benny GL, Humber AR, Voigt K (2014) Zygomycetous Fungi: Phylum Entomophthoromycota and Subphyla Kickxellomycotina, Mortierellomycotina, Mucoromycotina, and Zoopagomycotina. The Mycota 14:209–250
Blackwell M (2011) The Fungi: 1, 2, 3 … 5.1 million species? Am J Bot 98:426–438
Bruns TD, Corradi N, Redecker D, Taylor JW, Öpik M (2018) Glomeromycotina: what is a species and why should we care? New Phytol 220:963–967
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST + : architecture and applications. BMC Bioinform 10:421
Chang Y, Desirò A, Na H, Sandor L, Lipzen A, Clum A, Barry K, Grigoriev IV, Martin FM, Stajich JE, Smith ME, Spatafora J (2019) Phylogenomics of Endogonaceae and evolution of mycorrhizas within Mucoromycota. New Phytol 222:511–525
Colin S, Coelho LP, Sunagawa S, Bowler C, Karsenti E, Bork P, Pepperkok R, DeVargas C (2017) Quantitative 3D-imaging for cell biology and ecology of environmental microbial eukaryotes. eLife 6:e26066
Corsaro D, Wylezich C, Venditti D, Michel R, Walochnik J, Wegensteiner R (2019) Filling gaps in the microsporidian tree: rDNA phylogeny of Chytridiopsis typographi (Microsporidia: Chytridiopsida). Parasitol Res 23:169–180
Davison J, Moora M, Öpik M, Ainsaar L, Ducousso M, Hiiesalu I, Jairus T, Johnson N, Jourand P, Kalamees R, Koorem K, Zobel M (2018) Microbial island biogeography: isolation shapes the life history characteristics but not diversity of root-symbiotic fungal communities. ISME J 12:2211–2224
Desiro A, Rimington WR, Jacob A, Pol NV, Smith ME, Trappe JM, Bidartondo MI, Bonito G (2017) Multigene phylogeny of Endogonales, an early diverging lineage of fungi associated with plants. IMA Fung 8:245–264
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Egidi E, Delgado-Baquerizo M, Plett JM, Wang J, Eldridge DJ, Bardgett RD, Maestre FT, Singh BK (2019) A few Ascomycota taxa dominate soil fungal communities worldwide. Nat Commun 10:2369
Grossart HP, Van den Wyngaert S, Kagami M, Wurzbacher C, Cunliffe M, Rojas-Jimenez K (2019) Fungi in aquatic ecosystems. Nat Rev Microbiol 17:339–354
Hawksworth DL, Lücking R (2017) Fungal diversity revisited: 2.2 to 3.8 million species. Microbiol Spectr 1:79–95
Heeger F, Bourne EC, Baschien C, Yurkov A, Bunk B, Spröer C, Overmann J, Mazzoni CJ, Monaghan MT (2018) Long-read DNA metabarcoding of ribosomal rRNA in the analysis of fungi from aquatic environments. Mol Ecol Res 18:1500–1514
Hibbett DS, Binder M, Bischoff J (2007) A higher-level phylogenetic classification of the Fungi. Mycol Res 111:509–547
James TY, Kauff F, Schoch CL (2006) Reconstructing the early evolution of fungi using a six-gene phylogeny. Nature 443:818–822
Jones MDM, Forn I, Gadelha C, Egan MJ, Bass D, Massana R, Richards TA (2011) Discovery of novel intermediate forms redefines the fungal tree of life. Nature 474:200–203
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Ko TWK, Stephenson SL, Bahkali AH, Hyde KD (2011) From morphology to molecular biology: can we use sequence data to identify fungal endophytes? Fung Divers 50:113
Kõljalg U, Larsson K-H, Abarenkov K, Nilsson RH, Alexander IJ, Eberhardt U, Erland S, Høiland K, Kjøller R, Larsson E, Pennanen T, Sen R, Taylor AFS, Tedersoo L, Vrålstad T, Ursing BM (2005) UNITE: a database providing web-based methods for the molecular identification of ectomycorrhizal fungi. New Phytol 166:1063–1068
Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M et al (2013) Towards a unified paradigm for sequence-based identification of Fungi. Mol Ecol 22:5271–5277
Kõljalg U, Tedersoo L, Nilsson RH, Abarenkov K (2016) Digital identifiers for fungal species. Science 352:1182–1183
Larsson K-H (2007) Re-thinking the classification of corticioid fungi. Mycol Res 111:1040–1063
Liu XZ, Wang QM, Göker M, Groenewald M, Kachalkin AV, Lumbsch HT, Millanes AM, Wedin M, Yurkov AM, Boekhout T, Bai FY (2015) Towards an integrated phylogenetic classification of the Tremellomycetes. Stud Mycol 81:85–147
Loit K, Adamson K, Bahram M, Puusepp R, Anslan S, Kiiker R, Drenkhan R, Tedersoo L (2019) Relative performance of MinION (Oxford Nanopore Technologies) versus Sequel (Pacific Biosciences) third-generation sequencing instruments in identification of agricultural and forest fungal pathogens. Appl Environ Microbiol 85:e01368-19
Menkis A, Urbina H, James TY, Rosling A (2014) Archaeorhizomyces borealis sp. nov. and a sequence-based classification of related soil fungal species. Fung Ecol 118:943–945
Monchy S, Sanciu G, Jobard M, Rasconi S, Gerphagnon M, Chabe M, Cian A, Meloni D, Niquil N, Christaki U, Viscogliosi E, Sime-Ngando T (2011) Exploring and quantifying fungal diversity in freshwater lake ecosystems using rDNA cloning/sequencing and SSU tag pyrosequencing. Environ Microbiol 13:1433–1453
Nagy LG, Petkovits T, Kovacs G, Voigt K, Vagvölgyi C, Papp T (2011) Where is the unseen fungal diversity hidden? A study of Mortierella reveals a large contribution of reference collections to the identification of fungal environmental sequences. New Phytol 191:789–794
Nilsson RH, Wurzbacher C, Bahram M, Coimbra VRM, Larsson E, Tedersoo L, Eriksson J, Duarte Ritter C, Svantesson S, Sánchez-García M, Ryberg M, Kristiansson E, Abarenkov K (2017) Top 50 most wanted fungi. MycoKeys 12:29–40
Nilsson RH, Larsson KH, Taylor AF, Bengtsson-Palme J, Jeppesen TS, Schigel D, Kennedy P, Picard K, Glöckner FO, Tedersoo L, Saar I, Kõljalg U, Abarenkov K (2018) The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucl Acids Res 47:D259–D264
Nilsson RH, Anslan S, Bahram M, Wurzbacher C, Baldrian P, Tedersoo L (2019) Mycobiome diversity: high-throughput sequencing and identification of fungi. Nat Rev Microbiol 17:95–109
Öpik M, Vanatoa A, Vanatoa E, Moora M, Davison J, Kalwij JM, Reier Ü, Zobel M (2010) The online database MaarjAM reveals global and ecosystemic distribution patterns in arbuscular mycorrhizal fungi (Glomeromycota). New Phytol 188:223–241
Powell MJ, Letcher PM (2014) Chytridiomycota, Monoblepharidomycota and Neocallimastigomycota. Mycota 9:141–176
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl Acids Res 41:D590–D596
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JS, Strez B, Thallinger GG, van Horn DJ, Weber CF (2009) Introducing Mothur: open-source, platform-independent. Community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Bolchacova E, Voigt K (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA 109:6241–6246
Spatafora JW, Chang Y, Benny GL, Lazarus K, Smith ME, Berbee ML, Bonito G, Corradi N, Grigoriev I, Gryganskyi A, James TY (2016) A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia 108:1028–1046
Spatafora JW, Aime MC, Grigoriev IV, Martin F, Stajich JE, Blackwell M (2017) The fungal tree of life: from molecular systematics to genome-scale phylogenies. Microbiol Spectr 5:FUNK-0053-2016
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 10:1093
Taylor DL, Hollingsworth TN, McFarland JW, Lennon NJ, Nussbaum C, Ruess RW (2014) A first comprehensive census of fungi in soil reveals both hyperdiversity and fine-scale niche partitioning. Ecol Monogr 84:3–20
Tedersoo L, Anslan S (2019) Towards PacBio-based pan-eukaryote metabarcoding using full-length ITS sequences. Environ Microbiol Rep 11:659–668
Tedersoo L, Lindahl B (2016) Fungal identification biases in microbiome projects. Environ Microbiol Rep 8:774–779
Tedersoo L, Nilsson RH (2016) Molecular identification of fungi. In: Martin F (ed) Molecular Mycorrhizal Symbiosis. Wiley, Hoboken, pp 301–322
Tedersoo L, Bahram M, Põlme S, Kõljalg U, Yorou NS, Wijesundera R, Villarreal-Ruiz L, Vasco-Palacios A, Quang Thu P, Suija A, Smith ME, Abarenkov K (2014) Global diversity and geography of soil fungi. Science 346:1078
Tedersoo L, Bahram M, Põlme S, Anslan S, Riit T, Kõljalg U, Nilsson RH, Hildebrand F, Abarenkov K (2015) Response to comment on “Global diversity and geography of soil fungi”: analytical biases in microbial diversity studies. Science 359:936
Tedersoo L, Bahram M, Puusepp R, Nilsson RH, James TY (2017) Novel soil-inhabiting clades fill gaps in the fungal tree of life. Microbiome 5:42
Tedersoo L, Kõljalg U, Bahram M, Döring M, Schigel D, May T, Sanchez-Ramirez S, Ryberg M, Abarenkov K (2018a) High-level classification of the Fungi and a tool for evolutionary ecological analyses. Fung Divers 90:135–159
Tedersoo L, Tooming-Klunderud A, Anslan S (2018b) PacBio metabarcoding of fungi and other eukaryotes: biases and perspectives. New Phytol 217:1370–1385
Tedersoo L, Anslan S, Bahram M, Drenkhan R, Pritsch K, Buegger F, Padari A, Hagh-Doust N, Mikryukov V, Kõljalg U, Abarenkov K (2020) Regional-scale in-depth analysis of soil fungal diversity reveals strong pH and plant species effects in Northern Europe. Front Microbiol. https://doi.org/10.3389/fmicb.2020.01953
U’Ren JM, Lutzoni F, Miadlikowska J, Zimmerman NB, Carbone I, May G, Arnold AE (2019) Host availability drives distributions of fungal endophytes in the imperilled boreal realm. Nat Ecol Evol 3:1430–1437
Varga T, Krizsán K, Földi C, Dima B, Sánchez-García M, Sánchez-Ramírez S, Szöllősi GJ, Szarkándi JG, Papp V, Albert L, Andreopoulos W, Nagy L (2019) Megaphylogeny resolves global patterns of mushroom evolution. Nat Ecol Evol 3:668–678
Walder F, Schlaeppi K, Wittwer R, Held AY, Vogelgsang S, van der Heijden MG (2017) Community profiling of Fusarium in combination with other plant-associated fungi in different crop species using SMRT sequencing. Front Plant Sci 2017:8
Wijayawardene NN, Hyde KD, Al-Ani LK, Tedersoo L, Haelewaters D, Rajeshkumar KC, Zhao RL, Aptroot A, Leontyev D, Saxena RK, Tokarev YS (2020) Outline of Fungi and fungus-like taxa. Mycosphere 11:1060–1456
Wurzbacher C, Larsson E, Bengtsson-Palme J, Van den Wyngaert S, Svantesson S, Kristiansson E, Kagami M, Nilsson RH (2019) Introducing ribosomal tandem repeat barcoding for fungi. Mol Ecol Res 19:118–127
Acknowledgements
We thank multiple colleagues for providing samples and Rasmus Puusepp for molecular analyses. This study was funded by the Estonian Science Foundation (Grants PUT1399, PRG632, MOBERC21).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tedersoo, L., Anslan, S., Bahram, M. et al. Identifying the ‘unidentified’ fungi: a global-scale long-read third-generation sequencing approach. Fungal Diversity 103, 273–293 (2020). https://doi.org/10.1007/s13225-020-00456-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-020-00456-4