Abstract
Neurodegenerative diseases with tau pathology, or tauopathies, include Alzheimer’s disease and related dementia disorders. Previous work has shown that loss of the poly(A) RNA-binding protein gene sut-2/MSUT2 strongly suppressed tauopathy in Caenorhabditis elegans, human cell culture, and mouse models of tauopathy. However, the mechanism of suppression is still unclear. Recent work has shown that MSUT2 protein interacts with the THO complex and ALYREF, which are components of the mRNA nuclear export complex. Additionally, previous work showed ALYREF homolog Ref1 modulates TDP-43 and G4C2 toxicity in Drosophila melanogaster models. We used transgenic C. elegans models of tau or TDP-43 toxicity to investigate the effects of loss of ALYREF function on tau and TDP-43 toxicity. In C. elegans, three genes are homologous to human ALYREF: aly-1, aly-2, and aly-3. We found that loss of C. elegans aly gene function, especially loss of both aly-2 and aly-3, suppressed tau-induced toxic phenotypes. Loss of aly-2 and aly-3 was also able to suppress TDP-43-induced locomotor behavior deficits. However, loss of aly-2 and aly-3 had divergent effects on mRNA and protein levels as total tau protein levels were reduced while mRNA levels were increased, but no significant effects were seen on total TDP-43 protein or mRNA levels. Our results suggest that although aly genes modulate both tau and TDP-43-induced toxicity phenotypes, the molecular mechanisms of suppression are different and separated from impacts on mRNA and protein levels. Altogether, this study highlights the importance of elucidating RNA-related mechanisms in both tau and TDP-43-induced toxicity.








Similar content being viewed by others
Data availability
The data generated during this study is included in the published article and linked supplementary files.
Abbreviations
- ALS:
-
amyotrophic lateral sclerosis
- gRNAs:
-
guide RNAs
- NGM:
-
nematode growth media
- non Tg:
-
non-transgenic
- TDP-43:
-
TAR DNA-binding protein 43
- Tg:
-
transgenic
References
Götz J, Halliday G, Nisbet RM. Molecular Pathogenesis of the Tauopathies. Annu Rev Pathol Mech Dis. 2019;14:239–61.
Bierer LM, Hof PR, Purohit DP, Carlin L, Schmeidler J, Davis KL, et al. Neocortical neurofibrillary tangles correlate with dementia severity in Alzheimer’s disease. Arch Neurol. 1995;52:81–8.
Riley KP, Snowdon DA, Markesbery WR. Alzheimer’s neurofibrillary pathology and the spectrum of cognitive function: findings from the Nun Study. Ann Neurol. 2002;51:567–77.
Tiraboschi P, Hansen LA, Thal LJ, Corey-Bloom J. The importance of neuritic plaques and tangles to the development and evolution of AD. Neurology. 2004;62:1984–9.
MAPT ( tau ) Protein C-terminus pathogenic benign protective unclear. :441. Available from: https://www.alzforum.org/mutations/mapt.
Chang CW, Shao E, Mucke L. Tau: Enabler of diverse brain disorders and target of rapidly evolving therapeutic strategies. Science (80- ). 2021;371.
Soeda Y, Takashima A. New insights into drug discovery targeting tau protein. Front Mol Neurosci. 2020;13:1–24.
Kraemer BC, Zhang B, Leverenz JB, Thomas JH, Trojanowski JQ, Schellenberg GD. Neurodegeneration and defective neurotransmission in a Caenorhabditis elegans model of tauopathy. Proc Natl Acad Sci U S A [Internet]. 2003;100:9980–5. Available from: http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=187908&tool=pmcentrez&rendertype=abstract
Guthrie CR, Schellenberg GD, Kraemer BC. SUT-2 potentiates tau-induced neurotoxicity in Caenorhabditis elegans. Hum Mol Genet. 2009;18:1825–38.
Wheeler JM, McMillan P, Strovas TJ, Liachko NF, Amlie-Wolf A, Kow RL, et al. Activity of the poly(A) binding protein MSUT2 determines susceptibility to pathological tau in the mammalian brain. Sci Transl Med. 2019;11:eaao6545.
Kelly SM, Leung SW, Pak C, Banerjee A, Moberg KH, Corbett AH. A conserved role for the zinc finger polyadenosine RNA binding protein, ZC3H14, in control of poly(A) tail length. Rna. 2014;20:681–8.
Kow RL, Strovas TJ, McMillan PJ, Jacobi AM, Behlke MA, Saxton AD, et al. Distinct Poly(A) nucleases have differential impact on sut-2 dependent tauopathy phenotypes. Neurobiol Dis. 2021;147:105148. https://doi.org/10.1016/j.nbd.2020.105148.
Morris KJ, Corbett AH. The polyadenosine RNA-binding protein ZC3H14 interacts with the THO complex and coordinately regulates the processing of neuronal transcripts. Nucleic Acids Res. 2018;46:6561–75.
Zhou Z, Luo MJ, Straesser K, Katahira J, Hurt E, Reed R. The protein Aly links pre-messenger-RNA splicing to nuclear export in metazoans. Nature. 2000;407:401–5.
Stubbs SH, Conrad NK. Depletion of REF/Aly alters gene expression and reduces RNA polymerase II occupancy. Nucleic Acids Res. 2015;43:504–19.
Shi M, Zhang H, Wu X, He Z, Wang L, Yin S, et al. ALYREF mainly binds to the 5′ and the 3′ regions of the mRNA in vivo. Nucleic Acids Res. 2017;45:9640–53.
Yang X, Yang Y, Sun BF, Chen YS, Xu JW, Lai WY, et al. 5-methylcytosine promotes mRNA export-NSUN2 as the methyltransferase and ALYREF as an m 5 C reader. Cell Res. 2017;27:606–25. https://doi.org/10.1038/cr.2017.55.
Freibaum BD, Lu Y, Lopez-Gonzalez R, Kim NC, Almeida S, Lee KH, et al. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature. 2015;525:129–33.
Berson A, Goodman LD, Sartoris AN, Otte CG, Aykit JA, Lee VMY, et al. Drosophila Ref1/ALYREF regulates transcription and toxicity associated with ALS/FTD disease etiologies. Acta Neuropathol Commun. 2019;7:65.
Prpar Mihevc S, Baralle M, Buratti E, Rogelj B. TDP-43 aggregation mirrors TDP-43 knockdown, affecting the expression levels of a common set of proteins. Sci Rep. 2016;6:1–9.
Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974;77:71–94.
Liachko NF, Guthrie CR, Kraemer BC. Phosphorylation promotes neurotoxicity in a Caenorhabditis elegans model of TDP-43 proteinopathy. J Neurosci. 2010;30:16208–19.
Paix A, Folkmann A, Rasoloson D, Seydoux G. High efficiency, homology-directed genome editing in Caenorhabditis elegans using CRISPR-Cas9ribonucleoprotein complexes. Genetics. 2015;201:47–54.
Arribere JA, Bell RT, Fu BXH, Artiles KL, Hartman PS, Fire AZ. Efficient marker-free recovery of custom genetic modifications with CRISPR/Cas9 in caenorhabditis elegans. Genetics. 2014;198:837–46.
Kraemer BC, Schellenberg GD. SUT-1 enables tau-induced neurotoxicity in C. elegans. Hum Mol Genet. 2007;16:1959–71.
Kow RL, Sikkema C, Wheeler JM, Wilkinson CW, Kraemer BC. DOPA decarboxylase modulates tau toxicity. Biol Psychiatry. 2018;83:438–46. https://doi.org/10.1016/j.biopsych.2017.06.007.
Brignull HR, Moore FE, Tang SJ, Morimoto RI. Polyglutamine proteins at the pathogenic threshold display neuron-specific aggregation in a pan-neuronal Caenorhabditis elegans model. J Neurosci. 2006;26:7597–606.
Longman D, Johnstone IL, Cáceres JF. The REF/Aly proteins are dispensable for mRNA export and development in Caenorhabditis elegans. RNA. 2003;9:881–91.
Macmorris M, Brocker C, Blumenthal T. UAP56 levels affect viability and mRNA export in Caenorhabditis elegans. RNA. 2003;9:847–57.
Castellano-Pozo M, García-Muse T, Aguilera A. The Caenorhabditis elegans THO complex is required for the mitotic cell cycle and development. PLoS One. 2012;7.
Gatfield D, Izaurralde E. REF1/Aly and the additional exon junction complex proteins are dispensable for nuclear mRNA export. J Cell Biol. 2002;159:579–88.
Straesser K, Hurt E. Yra1p, a conserved nuclear RNA-binding protein, interacts directly with Mex67p and is required for mRNA export. EMBO J. 2000;19:410–20.
Okada M, Jang SW, Ye K. Akt phosphorylation and nuclear phosphoinositide association mediate mRNA export and cell proliferation activities by ALY. Proc Natl Acad Sci U S A. 2008;105:8649–54.
Dias AP, Dufu K, Lei H, Reed R. A role for TREX components in the release of spliced mRNA from nuclear speckle domains. Nat Commun. 2010;1.
Maeder CI, Kim JI, Liang X, Kaganovsky K, Shen A, Li Q, et al. The THO complex coordinates transcripts for synapse development and dopamine neuron survival. Cell. 2018;174:1436–1449.e20. https://doi.org/10.1016/j.cell.2018.07.046.
Kuersten S, Segal SP, Verheyden J, LaMartina SM, Goodwin EB. NXF-2, REF-1, and REF-2 affect the choice of nuclear export pathway for tra-2 mRNA in C. elegans. Mol Cell. 2004;14:599–610.
Tollervey JR, Curk T, Rogelj B, Briese M, Cereda M, Kayikci M, et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat Neurosci. 2011;14:452–8.
Polymenidou M, Lagier-Tourenne C, Hutt KR, Huelga SC, Moran J, Liang TY, et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat Neurosci. 2011;14:459–68.
Arnold ES, Ling SC, Huelga SC, Lagier-Tourenne C, Polymenidou M, Ditsworth D, et al. ALS-linked TDP-43 mutations produce aberrant RNA splicing and adult-onset motor neuron disease without aggregation or loss of nuclear TDP-43. Proc Natl Acad Sci U S A. 2013;110.
Apicco DJ, Ash PEA, Maziuk B, Leblang C, Medalla M, Al Abdullatif A, et al. Reducing the RNA binding protein TIA1 protects against tau-mediated neurodegeneration in vivo. Nat Neurosci. 2018;21:72–82.
McMillan PJ, Strovas TJ, Baum M, Mitchell BK, Eck RJ, Hendricks N, et al. Pathological tau drives ectopic nuclear speckle scaffold protein SRRM2 accumulation in neuron cytoplasm in Alzheimer’s disease. Acta Neuropathol Commun. 2021;9:117. https://doi.org/10.1186/s40478-021-01219-1.
Montalbano M, McAllen S, Puangmalai N, Sengupta U, Bhatt N, Johnson OD, et al. RNA-binding proteins Musashi and tau soluble aggregates initiate nuclear dysfunction. Nat Commun. 2020;11:1–16. https://doi.org/10.1038/s41467-020-18022-6.
Lester E, Ooi FK, Bakkar N, Ayers J, Woerman AL, Wheeler J, et al. Tau aggregates are RNA-protein assemblies that mislocalize multiple nuclear speckle components. Neuron. 2021;109:1675–1691.e9. https://doi.org/10.1016/j.neuron.2021.03.026.
Hsieh YC, Guo C, Yalamanchili HK, Abreha M, Al-Ouran R, Li Y, et al. Tau-mediated disruption of the spliceosome triggers cryptic RNA splicing and neurodegeneration in Alzheimer’s disease. Cell Rep. 2019;29:301–316.e10. https://doi.org/10.1016/j.celrep.2019.08.104.
Guo Q, Dammer EB, Zhou M, Kundinger SR, Gearing M, Lah JJ, et al. Targeted quantification of detergent-insoluble RNA-binding proteins in human brain reveals stage and disease specific Co-aggregation in Alzheimer’s disease. Front Mol Neurosci. 2021;14:1–17.
Bryan J, Nagle BW, Doenges KH. Inhibition of tubulin assembly by RNA and other polyanions: Evidence for a required protein. Proc Natl Acad Sci U S A. 1975;72:3570–4.
Chakraborty P, Rivière G, Liu S, de Opakua AI, Dervişoğlu R, Hebestreit A, et al. Co-factor-free aggregation of tau into seeding-competent RNA-sequestering amyloid fibrils. Nat Commun. 2021;12:1–12.
Ginsberg SD, Crino PB, Lee VMY, Eberwine JH, Trojanowski JQ. Sequestration of RNA in Alzheimer’s disease neurofibrillary tangles and senile plaques. Ann Neurol. 1997;41:200–9.
Ranganathan R, Haque S, Coley K, Shepheard S, Cooper-Knock J, Kirby J. Multifaceted genes in amyotrophic lateral sclerosis-frontotemporal dementia. Front Neurosci. 2020;14:1–21.
Lee J, Kim M, Itoh TQ, Lim C. Ataxin-2: a versatile posttranscriptional regulator and its implication in neural function. Wiley Interdiscip Rev RNA. 2018;9:1–13.
Purice MD, Taylor JP. Linking hnRNP function to ALS and FTD pathology. Front Neurosci. 2018;12:1–12.
Tomé SO, Gomes LA, Li X, Vandenberghe R, Tousseyn T, Thal DR. TDP-43 interacts with pathological τ protein in Alzheimer’s disease. Acta Neuropathol. 2021;141:795–9. https://doi.org/10.1007/s00401-021-02295-2.
Latimer CS, Burke BT, Liachko NF, Currey HN, Kilgore MD, Gibbons LE, et al. Resistance and resilience to Alzheimer’s disease pathology are associated with reduced cortical pTau and absence of limbic-predominant age-related TDP-43 encephalopathy in a community-based cohort. Acta Neuropathol Commun. 2019;7:91.
Acknowledgements
We thank the reviewers for their useful suggestions and commentary. We thank Heather Currey, Brandon Henderson, and Jade Stair for exceptional technical assistance. We thank Dr. Yishi Jin for the CZ1200 C. elegans strain. We thank the Developmental Studies Hybridoma Bank (NICHD) for the β-Tubulin antibody E7. We thank WormBase (WS251) for C. elegans genetic information. We thank the CGC (funded by the NIH Office of Research Infrastructure Programs (P40 OD010440)) and the National Bioresource Project (Japan) for the provision of C. elegans strains used for initial studies.
Funding
This work was supported by grants from the Department of Veterans Affairs [Merit Review Grants #I01BX002619 and #I01BX004044, and Career Development Award Level-2 #BX004341-01A1] and National Institutes of Health [RF1AG055474 and R01AG066729].
Author information
Authors and Affiliations
Contributions
Conceptualization: BK, RK. Methodology: AB, AS, BK, NL, RK. Resources: AB, AS, BK, NL, RK. Performing experiments: AB, RK. Data analysis: AB, RK. Visualization: AB, RK. Supervision: RK, BK; Funding acquisition: RK, BK; Project administration: RK, BK. Writing—original draft: BK, RK. Writing—revision and editing: All.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information

Fig. S1
Loss of aly genes on swimming behavior of tau-transgenic C. elegans. (PNG 207 kb)

Fig. S2
Loss of aly genes on swimming behavior of non-transgenic C. elegans. (PNG 232 kb)

Fig. S3
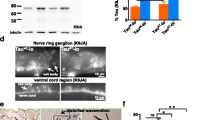
Total tau, RAB-soluble tau, RIPA-soluble tau, insoluble (formic acid) tau, and tubulin (loading control) immunoblots. (PNG 496 kb)

Fig. S4
TDP-43 immunoblots: phosphorylated (pS409/S410) TDP-43, total TDP-43, and tubulin (loading control). (PNG 260 kb)

Fig. S5
SUT-2, tau or TDP-43, and tubulin (loading control) immunoblots. (PNG 506 kb)
About this article
Cite this article
Kow, R.L., Black, A.H., Saxton, A.D. et al. Loss of aly/ALYREF suppresses toxicity in both tau and TDP-43 models of neurodegeneration. GeroScience 44, 747–761 (2022). https://doi.org/10.1007/s11357-022-00526-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11357-022-00526-2