Abstract
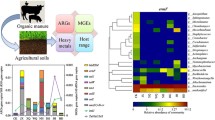
Massive antibiotic resistance genes (ARG) were detected in the soil modified by manure, which may affect human life safety through the food chain. However, the transmission of ARGs through the soil–plant-animal food chain is still unclear. Therefore, this study used high-throughput quantitative PCR technology to explore the effects of pig manure application on ARGs and bacterial communities in soil, lettuce phyllosphere, and snail excrement. The results showed that a total of 384 ARGs and 48 MEGs were detected in all samples after 75 days of incubation. The diversity of ARGs and MGEs in soil components increased significantly by 87.04% and 40% with the addition of pig manure. The absolute abundance of ARGs in the phyllosphere of lettuce was significantly higher than that of the control group, with a growth rate of 212.5%. Six common ARGs were detected between the three components of the fertilization group, indicating that there was internal transmission of fecal ARGs between the trophic levels of the food chain. Firmicutes and Proteobacteria were identified as the dominant host bacteria in the food chain system, which were more likely to be used as carriers of ARGs to promote the spread of resistance in the food chain. The results were used to assess the potential ecological risks of livestock and poultry manure. It provides theoretical basis and scientific support for the formulation of ARG prevention and control policies.













Similar content being viewed by others

Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments [J]. Nat Rev Microbiol 8(4):251–259. https://doi.org/10.1038/nrmicro2312
An X-L, Su J-Q (2022) Soil antibiotic resistance group : source, diffusion and driving factors [J ]. Sci Technol Bull 40(03):64–74
Angela HAMVH, Christiaan V, Wendy MVO, Gretta L, Ana MDRH, Hetty B (2015) Prevalence and characterization of ESBL- and AmpC-producing Enterobacteriaceae on retail vegetables [J]. Int J Food Microbiol 204:1–8. https://doi.org/10.3390/ANTIBIOTICS10111333
Bao C-X, Zhang Y, Chen M-L et al (2022) Study on the reduction of antibiotics and antibiotic resistance genes during pig manure composting by different biochars [J ]. Environ Pollut Prev 44(02):156–232
Bombatwala S, Mandpe A, Paliya S et al (2021) Antibiotic resistance in the environment: a critical insight on its occurrence, fate, and eco-toxicity [J]. Environ Sci Pollut Res 28(20):1–28. https://doi.org/10.1007/S11356-021-13143-X
Cerqueira F, Christou A, Fatta-kassinos D et al (2020) Effects of prescription antibiotics on soil- and root-associated microbiomes and resistomes in an agricultural context [J]. J Hazard Mater 400(prepublish):123208. https://doi.org/10.1016/j.jhazmat.2020.123208
Charlop PZ, Owen JG, Reddy BVB, Ternei MA et al (2015) Global biogeographic sampling of bacterial secondary metabolism [J]. eLife 4:1–10. https://doi.org/10.7554/eLife.05048
Chen B-W, Liang X-P, Nie X-P et al (2015) The role of class I integrons in the dissemination of sulfonamide resistance genes in the Pearl River and Pearl River Estuary, South China [J]. J Hazard Mater 282:61–67. https://doi.org/10.1016/j.jhazmat.2014.06.010
Chen Q-L, An X-L, Zhu Y-G, Su J-Q, Gillings-Michael R, Ye Z-L, Cui L (2017) Application of struvite alters the antibiotic resistome in soil, rhizosphere, and phyllosphere.[J]. Environ Sci Technol 51(14):8149–8157. https://doi.org/10.1021/acs.est.7b01420
Chen Q-L, An X-L, Zheng B-X et al (2018) Long-term organic fertilization increased antibiotic resistome in phyllosphere of maize [J]. Sci Total Environ 645:1230–1237. https://doi.org/10.1016/j.scitotenv.2018.07.260
Chen Q-L, AN X-L, LI H et al (2017) Do manure-borne or indigenous soil microorganisms influence the spread of antibiotic resistance genes in manured soil? [J]. Soil Biol Biochem 114:229–237. https://doi.org/10.1016/j.soilbio.2017.07.022
Cheng J-H, Tang X-Y, Liu C (2021) Bacterial communities regulate temporal variations of the antibiotic resistome in soil following manure amendment [J]. Environ Sci Pollut Res 28(23):1–12. https://doi.org/10.1007/S11356-021-12746-8
Ding J, Zhu D, Hong B et al (2019) Long-term application of organic fertilization causes the accumulation of antibiotic resistome in earthworm gut microbiota [J]. Environ Int 124:145–152. https://doi.org/10.1016/j.envint.2019.01.017
Duan Y-J, Chen Z-Y, Tan L, Wang X-L, Xue Y-G, Wang S-P, Wang Q, Das RJ, Lin H-A, Hou J, Li L-Y, Mao D-Q, Luo Y (2020) Gut resistomes, microbiota and antibiotic residues in Chinese patients undergoing antibiotic administration and healthy individuals [J]. Sci Total Environ 705(C):135674. https://doi.org/10.1016/j.scitotenv.2019.135674
Durán P, Thiergart T, Garrido-oter R et al (2018) Microbial interkingdom interactions in roots promote arabidopsis survival [J]. Cell 175(4):973–983. https://doi.org/10.1016/j.cell.2018.10.020
Fan X-T, Su J-Q, Zhou S-Y-D, An X-L, Li H (2023) Plant cultivar determined bacterial community and potential risk of antibiotic resistance gene spread in the phyllosphere[J]. J Environ Sci 127:508–518. https://doi.org/10.1016/J.JES.2022.06.006
Feng T-S, Yu Q-L, Zhou R et al (2020) Comparison of gut microbiota and common antibiotic resistance genes between plateau pika and plateau zokor [J ]. J Wildl 41(04):897–911
Florent M, Antoine G, Wegener PL (2023) Transmission mode and dispersal traits correlate with host specificity in mammalian gut microbes [J]. Mol Ecol. https://doi.org/10.1111/MEC.16862
Gao M, Jia R-Z, Qiu T-L et al (2017) Size-related bacterial diversity and tetracycline resistance gene abundance in the air of concentrated poultry feeding operations [J]. Environ Pollut 220(Pt B):1342–1348. https://doi.org/10.1016/j.envpol.2016.10.101
Guo Y-J, Qiu T-L, Gao M, Ru S-H, Gao H-Z, Wang X-M (2023) Does increasing the organic fertilizer application rate always boost the antibiotic resistance level in agricultural soils?[J]. Environ Pollut (Barking, Essex : 1987) 322:121251–121251. https://doi.org/10.1016/J.ENVPOL.2023.121251
He Y, Zhao X, Zhu S-M, Yuan L, Li X-Y, Feng Z-H, Yang X, Luo L, Xiao Y-L, Liu Y, Wang L-L, Deng O-P (2023) Conversion of swine manure into biochar for soil amendment: efficacy and underlying mechanism of dissipating antibiotic resistance genes[J]. Sci Total Environ 871:162046–162046. https://doi.org/10.1016/J.SCITOTENV.2023.162046
Heuer H, Schmitt H, Smakka K (2011) Antibiotic resistance gene spread due to manure application on agricultural fields [J]. Curr Opin Microbiol 14(3):246–243. https://doi.org/10.1016/j.mib.2011.04.009
Hu H-W, Wang J-T, Singh BK, Liu Y-R, Chen Y-L, Zhang Y-J, He J-Z (2018) Diversity of herbaceous plants and bacterial communities regulates soil resistome across forest biomes.[J]. Environ Microbiol 20(9):3186–3200. https://doi.org/10.1111/1462-2920.14248
Ishi K, Lauren W, Partha R et al (2022) Metagenomic tracking of antibiotic resistance genes through a pre-harvest vegetable production system: an integrated lab-, microcosm-, and greenhouse-scale study [J]. Environ Microbiol 24(8):3705–3721. https://doi.org/10.1111/1462-2920.16022
Jechalke S, Heuer H, Siemens J et al (2014) Fate and effects of veterinary antibiotics in soil [J]. Trends Microbiol 22(9):536–545. https://doi.org/10.1016/j.tim.2014.05.005
Kevin JF, Alejandro R, Bin W, Elizabeth MS, Morten OAS, Gautam D (2012) The shared antibiotic resistome of soil bacteria and human pathogens [J]. Science 337(6098):1107–1111. https://doi.org/10.1126/science.1220761
Li C, Chen H-Q, Gao P, Huang X-H, Zhu Y-X, Xu M, Yuan Q, Gao Y, Shen X-X (2023) Distribution and drivers of antibiotic resistance genes in brackish water aquaculture sediment [J]. Sci Total Environ 860:160475–160475. https://doi.org/10.1016/J.SCITOTENV.2022.160475
Liu S-H, Han Z-M, Zhu D, Luan X, Deng L-J, Dong L-P, Yang M, Zhang Y (2024) Field-based evidence for the enrichment of intrinsic antibiotic resistome stimulated by plant-derived fertilizer in agricultural soil [J]. J Environ Sci 135:728–740. https://doi.org/10.1016/J.JES.2022.08.009
Man Y, Li W-X, Wang J-X, Tam NF, Tai Y-P, Tao R, Yang Y (2022) Plants inhibit the relative abundance of sulfonamide resistance genes and class 1 integron by influencing bacterial community in rhizosphere of constructed wetlands [J]. Sci Total Environ 824:153977–153977. https://doi.org/10.1016/J.SCITOTENV.2022.153977
Mannanov RN, Sattarova RK (2001) Antibiotics produced by bacillus bacteria [J]. Chem Nat Compd 37(2):117–123. https://doi.org/10.1023/A:1012314516354
Meng F, Sun S-J, Geng J-L, Ma L-X, Jiang J-P, Li B, Dauda YS, Lu L, Fu D-L, Shen J-M, Qi H (2023) The source distribution and seasonal discrepancies of antibiotic resistance genes among multiple environment media in a cold megacity [J]. J Clean Prod 384:135164. https://doi.org/10.1016/J.JCLEPRO.2022.135164
Peng S, Wang Y-M, Zhou B-B et al (2015) Long-term application of fresh and composted manure increase tetracycline resistance in the arable soil of eastern China [J]. Sci Total Environ 279–286. https://doi.org/10.1016/j.scitotenv.2014.11.010
Penha FRAC, Ferreira JC, Galetti R, Kanashiro AMI, Berchieri A, da Costa Darini AL (2023) The rise of multidrug resistant Salmonella isolates in healthy chickens in Brazil by successful establishment of plasmid IncHI2A carrying several antibiotic resistance genes.[J]. Braz J Microbiol : [publication of the Brazilian Society for Microbiology] 54(1):469–474. https://doi.org/10.1007/S42770-022-00893-0
Pu C-J, Yu Y, Diao J-X et al (2019) Exploring the persistence and spreading of antibiotic resistance from manure to biocompost, soils and vegetables [J]. Sci Total Environ 688:262–269. https://doi.org/10.1016/j.scitotenv.2019.06.081
Pu Q, Zhao L-X, Li Y-T et al (2020) Manure fertilization increase antibiotic resistance in soils from typical greenhouse vegetable production bases, China [J]. J Hazard Mater 391(C):122267. https://doi.org/10.1016/j.jhazmat.2020.122267
Qiao M, Ying G-G, Singer AC et al (2018) Review of antibiotic resistance in China and its environment [J]. Environ Int 110:160–172. https://doi.org/10.1016/j.envint.2017.10.016
Rupert B, Annika N, Johann B et al (2022) Spread of antimicrobial resistance genes via pig manure from organic and conventional farms in the presence or absence of antibiotic use [J]. J Appl Microbiol 133(4):2457–2465. https://doi.org/10.1111/JAM.15717
Sarah H, Gangan W, Coelho KJ et al (2023) The fate of sulfonamide resistance genes and anthropogenic pollution marker intI1 after discharge of wastewater into a pristine river stream [J]. Front Microbiol 14:1058350–1058350. https://doi.org/10.3389/FMICB.2023.1058350
Stefanie M, Ye X, Nodwell JR (2014) The expression of antibiotic resistance genes in antibiotic-producing bacteria [J]. Mol Microbiol 93(3):391–402. https://doi.org/10.1111/mmi.12689
Su J-Q, Wei B, Ou Yang WY, Huang F-Y, Zhao Y, Xu H-J, Zhu Y-G (2015) Antibiotic resistome and its association with bacterial communities during sewage sludge composting.[J]. Environ Sci Technol 49(12):7356–7363. https://doi.org/10.1021/acs.est.5b01012
Sui Q-W, Meng X-S, Wang R et al (2018) Effects of endogenous inhibitors on the evolution of antibiotic resistance genes during high solid anaerobic digestion of swine manure [J]. Biores Technol 270:328–336. https://doi.org/10.1016/j.biortech.2018.09.043
Sun D-L, Jiang X, Wu Q-L-L, Zhou N-Y (2013) Intragenomic heterogeneity of 16S rRNA genes causes overestimation of prokaryotic diversity.[J]. Appl Environ Microbiol 79(19):5962–5969. https://doi.org/10.1128/AEM.01282-13
Tang M-J, Wu Z-Y, Li W-Z, Shoaib M, Aqib AI, Shang R-F, Yang Z, Pu W-X (2022) Effects of different composting methods on antibiotic-resistant bacteria, antibiotic resistance genes, and microbial diversity in dairy cattle manures.[J]. J Dairy Sci 106(1):257–273. https://doi.org/10.3168/JDS.2022-22193
Telma F, Ivone V-M, Manaia CM (2019) Neighbor urban wastewater treatment plants display distinct profiles of bacterial community and antibiotic resistance genes [J]. Environ Sci Pollut Res Int 26(11):11269–11278. https://doi.org/10.1007/s11356-019-04546-y
Wang F-H, Qiao M, Chen Z et al (2015) Antibiotic resistance genes in manure-amended soil and vegetables at harvest [J]. J Hazard Mater 299:215–221. https://doi.org/10.1016/j.jhazmat.2015.05.028
Wang B (2021) The distribution characteristics of tetracycline resistance genes in soil-vegetable [ D ]. Shenyang University. https://doi.org/10.27692/d.cnki.gsydx.2021.000399.
Whitehead TR, Cotta MA (2013) Stored swine manure and swine faeces as reservoirs of antibiotic resistance genes [J]. Lett Appl Microbiol 56(4):264–267. https://doi.org/10.1111/1am.12043
Wu Z-W, Zhang L, Lin H, Zhou S-G (2024) Enhanced removal of antibiotic resistance genes during chicken manure composting after combined inoculation of Bacillus subtilis with biochar [J]. J Environ Sci 135:274–284. https://doi.org/10.1016/J.JES.2022.12.002
Xiang Q, Chen Q-L, Yang X-R, Li G, Zhu D (2022) Microbial multitrophic communities drive the variation of antibiotic resistome in the gut of soil woodlice (Crustacea: Isopoda).[J]. Environ Sci Technol. https://doi.org/10.1021/ACS.EST.2C02471
Xie W-Y, Su J-Q, Zhu Y-G (2015) Phyllosphere bacterial community of floating macrophytes in paddy soil environments as revealed by illumina high-throughput sequencing.[J]. Appl Environ Microbiol 81(2):522–532. https://doi.org/10.1128/AEM.03191-14
Yan Q-F, Li X-Y, Ma B-H, Zou Y-D, Wang Y, Liao X-D, Liang J, Mi J-D, Wu Y-B (2018) Different concentrations of doxycycline in swine manure affect the microbiome and degradation of doxycycline residue in soil [J]. Front Microbiol 9:3129. https://doi.org/10.3389/fmicb.2018.03129
Yang Q-X, Wang R-F, Ren S-W, Szoboszlay M, Moe LA (2016) Practical survey on antibiotic-resistant bacterial communities in livestock manure and manure-amended soil [J]. J Environ Sci Health 51:14–23. https://doi.org/10.1080/03601234.2015.1080481
Yang Q-X, Ren S-W, Niu T-Q, Guo Y-H, Qi S-Y, Han X-K, Liu D, Pan F (2014) Distribution of antibiotic-resistant bacteria in chicken manure and manure-fertilized vegetables [J]. Environ Sci Pollut Res Int 21(2):1231–1241. https://doi.org/10.1007/s11356-013-1994-1
Ye Z-X, Tang Y-J, He L-Q et al (2022) Research progress on tetracycline antibiotic resistance: plasmid-mediated tigecycline resistance mechanism [J ]. ACTA Ecotoxicol 17(04):122–40
Yin Y, Zhu D, Yang G, Su J-Q, Duan G-L (2022) Diverse antibiotic resistance genes and potential pathogens inhabit in the phyllosphere of fresh vegetables [J]. Sci Total Environ 815:152851–152851. https://doi.org/10.1016/J.SCITOTENV.2021.152851
Yuan X-X, Zhang Y, Fan L-X, Wang W-B, Wu Y-J (2022) Temporal dynamics of antibiotic resistance genes in vegetable greenhouse soils following different manure applications [J]. J Soil Sci Plant Nutr 22(4):5144–5158. https://doi.org/10.1007/S42729-022-00990-X
Yuan X-X, Zhang Y, Sun C-X, Wang W-B, Wu Y-J, Fan L-X, Liu B (2022) Profile of bacterial community and antibiotic resistance genes in typical vegetable greenhouse soil [J]. Int J Environ Res Public Health 19(13):7742–7742. https://doi.org/10.3390/IJERPH19137742
Zhang H (2018) Transmission and transfer of antibiotics and their resistance in livestock manure-soil-vegetables [ D ]. Henan Normal University, pp 1–144
Zhang Y-J, Hu H-W, Chen Q-L et al (2019) Transfer of antibiotic resistance from manure-amended soils to vegetable microbiomes [J]. Environ Int 130:104912. https://doi.org/10.1016/j.envint.2019.104912
Zhang H, Dong M, Zhou Y et al (2021) Animal manure fertilization measures promote the spread of drug-resistant genes between manure-soil-vegetables [J ]. Environ Sci 42(04):2080–2088
Zhang Y, Zhou J, Wu J, Hua Q-W, Bao C-X (2022) Distribution and transfer of antibiotic resistance genes in different soil-plant systems[J]. Environ Sci Pollut Res Int 29(39):59159–59172. https://doi.org/10.1007/S11356-021-17465-8
Zhao C-X, Su X-X, Xu M-R, An X-L, Su J-Q (2022) Uncovering the diversity and contents of gene cassettes in class 1 integrons from the endophytes of raw vegetables.[J]. Ecotoxicol environ Saf 247:114282–114282. https://doi.org/10.1016/J.ECOENV.2022.114282
Zheng F, Zhu D, Giles M et al (2019) Mineral and organic fertilization alters the microbiome of a soil nematode Dorylaimus stagnalis and its resistome [J]. Sci Total Environ 680:70–78. https://doi.org/10.1016/j.scitotenv.2019.04.384
Zhu B, Chen Q, Chen S et al (2017) Does organically produced lettuce harbor higher abundance of antibiotic resistance genes than conventionally produced? [J]. Environ Int 98:152–159. https://doi.org/10.1016/j.envint.2016.11.001
Zhu D, Chen Q, Ding J et al (2019) Antibiotic resistance genes in soil ecosystems and planetary health: progress and prospects [J ]. Chin Sci : Life Sci 49(12):1652–1663
Zhu D, Xiang Q, Yang X-R, Ke X, Onnor P, Zhu Y-G (2019) Trophic transfer of antibiotic resistance genes in a soil detritus food chain.[J]. Environ Sci Technol 53(13):7770–7781. https://doi.org/10.1021/acs.est.9b00214
Zhu Y-G, Zhao Y, Zhu D et al (2019) Soil biota, antimicrobial resistance and planetary health [J]. Environ Int 131(12):105059. https://doi.org/10.1016/j.envint.2019.105059
Zou Y, Zhang Y, Zhou J, Bao C-X, Chen M-L, He W-C, Shi X-C (2022) Effects of composting pig manure at different mature stages on ARGs in different types of soil-vegetable systems.[J]. J Environ Manag 321:116042–116042. https://doi.org/10.1016/J.JENVMAN.2022.116042
Acknowledgements
We are grateful to the Youth Program of the National Natural Science Foundation of China (41701564), Suzhou Science and Technology Plan Program (SNG201613), and the Natural Science Foundation of the Higher Education institutions of Jiangsu Province, China (17KJB610010) for providing financial support for this study. We would like to thank Editage (www.editage.cn) for English language editing.
Funding
This research was funded by the Youth Program of the National Natural Science Foundation of China (41701564), Suzhou Science and Technology Plan Program (SNG201613), and Natural Science Foundation of the Higher Education Institutions of Jiangsu Province, China (17KJB610010).
Author information
Authors and Affiliations
Contributions
Conceptualization, Yuan Zhang; methodology, Yuan Zhang; software, Jiayi Zhao; validation, Minglong Chen, Xinyue Tang, Yijia Wang, and Yun Zou; formal analysis, Jiayi Zhao; investigation, Jiayi Zhao and Minglong Chen; resources, Minglong Chen; data curation, Jiayi Zhao; writing—original draft preparation, Jiayi Zhao; writing—review and editing, Yuan Zhang; visualization, Jiayi Zhao; supervision, Yuan Zhang; project administration, Yuan Zhang; funding acquisition, Yuan Zhang. All the authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Y., Zhao, J., Chen, M. et al. Fecal antibiotic resistance genes were transferred through the distribution of soil-lettuce-snail food chain. Environ Sci Pollut Res 30, 87793–87809 (2023). https://doi.org/10.1007/s11356-023-28606-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-28606-6