Abstract
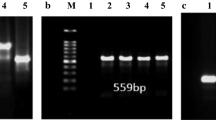
Campylobacters, especially C. jejuni and C. coli, have become one of the leading causes of acute gastroenteritis in humans worldwide in recent years. We aimed to investigate the presence, antimicrobial resistance, putative virulence genes, and molecular characterization of C. jejuni and C. coli recovered from human acute gastroenteritis cases in the study. In the study, suspected Campylobacter spp. isolates were obtained in 43 (5%) feces samples collected from a total of 850 patients who applied to the Erciyes University Medical Faculty acute clinic between March 2019 and February 2020. As a result of the phenotypic tests, these isolates were determined to be Campylobacter spp. According to the multiplex PCR, 33 of 43 Campylobacter spp. isolates were identified as C. jejuni (76%) and ten isolates were as C. coli (24%). In the disc diffusion test, the highest resistance rate was found in the trimethoprim/sulfamethoxazole (90.1%) and ciprofloxacin (90.1%), and the lowest rate was found in the amoxicillin-clavulanic acid (9.3%). In Campylobacter spp. isolates, the virulence genes cdtA, virB11, cdtB, cadF, iam, ceu, and flaA were found to be positive at rates of 26 (60%), 28 (65.6%), 13 (30%), 4 (9%), 27 (62%), 17 (39%), and 7 (16%), respectively. However, the cdtC gene was not detected in any of the isolates. The study searched tetO gene to examine the genetic aspect of tetracycline resistance, which was found in all Campylobacter spp. isolates. In the PCR reactions to investigate A2074C and A2075G mutations of macrolide resistance, it was determined as 7 (16%) and 21 (48%) of the isolates. To detect quinolone resistance, the rates of quinolone resistance-determining regions (QRDR) were 20 (45.4%) and the gyrA gene mutations in the mismatch amplification mutation assay PCR (MAMA-PCR), were 19 (43.1%) of isolates. In addition, the quinolone resistance gene (qnr) carried by plasmid in Campylobacter isolates was not found in the study. BlaOXA-61 and CmeB (multi-drug efflux pump) genes were detected as 28 (63.6%) and 30 (68.1), respectively. The Enterobacterial Repetitive Intergenic Consensus PCR (ERIC-PCR) used for typing the isolates revealed that the band profiles obtained from the isolates were different. In conclusion, this was a very comprehensive study revealing the presence of antibiotic-resistant C. jejuni and C. coli with various virulence genes in patients admitted to a university hospital with acute gastroenteritis. This is of utmost significance for public health. Since campylobacteria are foodborne, zoonotic pathogens and transmission occurs mostly through food. People should have detailed information about the transmission routes of campylobacteria and risky foods. In addition, staff, food processors and caterers, should be trained in hygiene.

Similar content being viewed by others
Data availability
The available (OM883911, OM883915, OM883913, OM883914, OM883912, OM906857, OM906854, OM906855, and OM906856) accession numbers at the link; https://blast.ncbi.nlm.nih.gov/Blast.cgi
References
Alfredson DA, Korolik V (2007) Antibiotic resistance and resistance mechanisms in Campylobacter jejuni and Campylobacter coli. Fems Microbiol Lett 277:123–132. https://doi.org/10.1111/j.1574-6968.2007.00935.x
Alonso R, Mateo E, Churruca E, Martinez I, Girbau C, Fernandez-Astorga A (2005) MAMA-PCR assay for the detection of point mutations associated with high-level erythromycin resistance in Campylobacter jejuni and Campylobacter coli strains. J Micro Methods 63:99–103. https://doi.org/10.1016/j.mimet.2005.03.013
Aslantaş Ö (2019) Isolation and molecular characterization of thermophilic Campylobacter spp. in dogs and cats. Kafkas Univ Vet Fak Derg 25(3):341–348. https://doi.org/10.9775/kvfd.2018.20952
Chansiripornchai N, Sasipreeyajan J (2009) PCR detection of four virulence-associated genes of Campylobacter jejuni isolates from Thai broilers and their abilities of adhesion to and invasion of INT-407 cells. J Vet Med Sci 71(6):839–844. https://doi.org/10.1292/jvms.71.839
CLSI. M100Ed31 (2021) Performance standards for antimicrobial susceptibility testing: 31st informational supplement. Clinical and Laboratory Standards Institute, Wayne
Cokal Y, Caner V, Sen A, Cetin C, Karagenc N (2009) Campylobacter spp. and their antimicrobial resistance patterns in poultry: an epidemiological survey study in Turkey. Zoonoses Public Health 56(3):105–110. https://doi.org/10.1111/j.1863-2378.2008.01155.x. (Epub 2008 Sep 22. PMID: 18811676)
Dai L, Sahin O, Grover M, Zhang Q (2020) New and alternative strategies for the prevention, control, and treatment of antibiotic-resistant Campylobacter. Transl Res 223:76–88. https://doi.org/10.1016/j.trsl.2020.04.009. (Epub 2020 May 11. PMID: 32438073; PMCID: PMC7423705)
Datta S, Niwa H, Itoh K (2003) Prevalence of 11 pathogenic genes of Campylobacter jejuni by PCR in strains isolated from humans, poultry meat and broiler and bovine faeces. J Med Microbiol 52(Pt 4):345–348. https://doi.org/10.1099/jmm.0.05056-0
Deogratias AP, Mushi MF, Paterno L, Tappe D, Seni J, Kabymera R, Mshana SE (2014) Prevalence and determinants of Campylobacter infection among under five children with acute watery diarrhea in Mwanza, North Tanzania. Arch Public Health 72:1–6
Elhadidy M, Ali MM, El-Shibiny A, Miller WG, Elkhatib WF, Botteldoorn N, Dierick K (2020) Antimicrobial resistance patterns and molecular resistance markers of Campylobacter jejuni isolates from human diarrheal cases. PLoS ONE 15(1):e0227833. https://doi.org/10.1371/journal.pone.0227833
Elmalı M, Can HY (2019) Antimicrobial susceptibility and virulence-associated genes in Campylobacter isolates from milk and wastewater in Hatay, Turkey. Ciência Rural 49(5):e20180227. https://doi.org/10.1590/0103-8478cr20180227
Garcia-Vallvé S, Pere P (2009) DendroUPGMA: a dendrogram construction utility. Universitat Rovira i Virgili, pp 1–14
Gibreel A, Tracz DM, Nonaka L, Ngo TM, Connell SR et al (2004) Incidence of antibiotic resistance in Campylobacter jejuni isolated in Alberta, Canada, from 1999 to 2002, with special reference to tet (O)-mediated tetracycline resistance. Antimicrob Agents Chemother 48:3442–3450
Guney M, Basustaoglu AC (2010) Gülhane Askeri Tıp Akademisi Egitim Hastanesi’nde akut bakteriyel gastroenterit etkenleri arasinda Campylobacter jejuni ve Campylobacter coli’nin yeri ve bunların antimikrobiklere duyarlılıklarının aras¸tırılması. Türk Mikrobiyol Cem Derg 40:183–192
Houf K, De Zutter L, Van Hoof J, Vandamme P (2002) Assessment of the genetic diversity among arcobacters isolated from poultry products by using two PCR-based typing methods. Appl Environ Microbiol 68(5):2172–2178. https://doi.org/10.1128/AEM.68.5.2172-2178.2002
Iovine NM (2013) Resistance mechanisms in Campylobacter jejuni. Virulence 4(3):230–240. https://doi.org/10.4161/viru.23753
Jacoby GA, Chow N, Waites KB (2003) Prevalence of plasmid-mediated quinolone. Antimicrob Agents Chemother 47(2):559–562. https://doi.org/10.1128/aac.47.2.559-562.2003
Kayman T, Abay S, Hizlisoy H (2013) Campylobacter türlerinin fenotipik yöntemler ve multipleks polimeraz zincir reaksiyonu ile tanımlanması ve antibiyotik duyarlılıkları. Mikrobiyol Bul 47(2):230–239
Kayman T, Abay S, Aydin F, Sahin O (2019) Antibiotic resistance of Campylobacter jejuni isolates recovered from humans with diarrhoea in Turkey. J Med Microbiol 68(2):136–142. https://doi.org/10.1099/jmm.0.000890
Komba EV, Mdegela RH, Msoffe PLM, Nielsen LN, Ingmer H (2015) Prevalence, antimicrobial resistance and risk factors for thermophilic Campylobacter infections in symptomatic and asymptomatic humans in Tanzania. Zoo Public Health 62(7):557–568. https://doi.org/10.1111/zph.12185
Krutkiewicz A, Klimuszko D (2010) Genotyping and PCR detection of potential virulence genes in Campylobacter jejuni and Campylobacter coli isolates from different sources in Poland. Folia Microbiol 55(2):167–175
Lehtopolku M, Hakanen AJ, Siitonen A, Huovinen P, Kotilainen P (2005) In vitro activities of 11 fluoroquinolones against 226 Campylobacter jejuni strains isolated from Finnish patients, with special reference to ciprofloxacin resistance. J Antimicrob Chemother 56:1134–1138. https://doi.org/10.1093/jac/dki380
Lengerh A, Moges F, Unakal C, Anagaw B (2013) Prevalence, associated risk factors and antimicrobial susceptibility pattern of Campylobacter species among under five diarrheic children at Gondar University Hospital Northwest Ethiopia. BMC Pediatr 13:9
Letunic I, Bork P (2019) Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47(W1):W256–W259. https://doi.org/10.1093/nar/gkz239
Lin J, Akiba M, Sahin O, Zhang Q (2005) CmeR functions as a transcriptional repressor for the multidrug efflux pump CmeABC in Campylobacter jejuni. Antimicrob Agents Chemother 49(3):1067–1075. https://doi.org/10.1128/aac.49.3.1067-1075.2005
Marotta F, Garofolo G, Di Marcantonio L, Di Serafino G, Neri D, Romantini R, Di Giannatale E (2019) Antimicrobial resistance genotypes and phenotypes of Campylobacter jejuni isolated in Italy from humans, birds from wild and urban habitats, and poultry. PLoS ONE 14(10):e0223804. https://doi.org/10.1371/journal.pone.0223804
Obeng AS, Rickard H, Sexton M, Pang Y, Peng H, Barton M (2012) Antimicrobial susceptibilities and resistance genes in Campylobacter strains isolated from poultry and pigs in Australia. J Appl Microbiol 113(2):294–307. https://doi.org/10.1111/j.1365-2672.2012.05354.x
Olah PA, Doetkott C, Fakhr MK, Logue CM (2006) Prevalence of the Campylobacter multi-drug efflux pump (CmeABC) in Campylobacter spp. isolated from freshly processed turkeys. Food Microbiol 23(5):453–460. https://doi.org/10.1016/j.fm.2005.06.004
Ongen B, Nazik H (2008) Investigation of plasmid mediated quinolone resistance among Campylobacter strains. Farmaci e Terapia 25:37–39
Osbjer K, Tano EVA, Chhayheng L, Mac-Kwashie AO, Fernström LL, Ellström P, Seng S, Choup S, Veng M, Kannanrath C, Sorn S, Holl D, Sofia B, Hilpi R, Ulf M, Magnusson U (2016) Detection of Campylobacter in human and animal field samples in Cambodia. APMIS 124(6):508–515. https://doi.org/10.1111/apm.12531
Taylor DE, Tracz DM (2005) Mechanisms of antimicrobial resistance in Campylobacter. In: Ketley JM, Konkel ME (eds) Campylobacter molecular and cellular biology. Horizon Bioscience, Norfolk, pp 193–204
Thames HT, Sukumaran AT (2020) A review of salmonella and campylobacter in broiler meat: emerging challenges and food safety measures. Foods 9(6):776. https://doi.org/10.3390/foods9060776
Tryjanowski P, Nowakowski JJ, Indykiewicz P, Andrzejewska M, Śpica D, Sandecki R, Cezary M, Artur G, Beata D, Joanna D, Tomasz J, Piotr M, Stanislaw S, Marcin T, Radoslaw W, Bernadeta S, Klawe JJ (2020) Campylobacter in wintering great tits Parus major in Poland. Environ Sci Pollut Res 27:7570–7577
Wang G, Clifford GC, Tracy MT et al (2022) Colony multiplex PCR assay for identification and differentiation of Campylobacter jejuni, C. coli, C. lari, C. upsaliensis, and C. fetus subsp. J Clinical Microbiol 40:4744–4747. https://doi.org/10.1128/jcm.40.12.4744-4747.2002
Wieczorek K, Osek J (2013) Antimicrobial resistance mechanisms among Campylobacter. Biomed Res Int. https://doi.org/10.1155/2013/340605
Wysok B, Wojtacka J, Hänninen ML, Kivistö R (2020) Antimicrobial resistance and virulence-associated markers in Campylobacter strains from diarrheic and non-diarrheic humans in Poland. Front Microbiol 11:1799. https://doi.org/10.3389/fmicb.2020.01799
Yılmaz AT, Tugrul HM (2005) Edirne’de ishal etkenleri arasında Campylobacter türlerinin yerinin ve antimikrobiklere duyarlılıklarının araştırılması. Infeksiyon Derg 19:53–59
Zhang X, Zhou Q, Tang M, Pu J, Zhang J, Lu J, Zhang Y, Gao Y (2021) Aminoglycoside resistance and possible mechanisms in Campylobacter Spp. isolated from chicken and swine in Jiangsu. China. Front Microbiol. 12:716185. https://doi.org/10.3389/fmicb.2021.716185
Zirnstein G, Li Y, Swaminathan B, Angulo F (1999) Ciprofloxacin resistance in Campylobacter jejuni isolates: detection of gyrA resistance mutations by mismatch amplification mutation assay PCR and DNA sequence analysis. J Clin Microbiol 37(10):3276–3280. https://doi.org/10.1128/jcm.37.10.3276-3280.1999
Funding
No funding was used in the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
HH, MB, AD MAA, PS; Methodology, Writing- Original draft preparation, KK, SH, PS, MAA, CG; Analyzes, Methodology, Writing- Original draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hizlisoy, H., Sagiroglu, P., Barel, M. et al. Campylobacter jejuni and Campylobacter coli in human stool samples: antibiotic resistance profiles, putative virulence determinants and molecular characterization of the isolates. World J Microbiol Biotechnol 39, 353 (2023). https://doi.org/10.1007/s11274-023-03786-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03786-y