Abstract
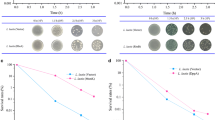
Molecular chaperone CbpA from extreme acidophile Acidithiobacillus caldus was applied to improve acid tolerance of Escherichia coli via CRISPR/Cas9. Cell growth and viability of plasmid complementary strain indicated the importance of cbpAAc for bacteria acid tolerance. With in situ gene replacement by CRISPR/Cas9 system, colony formation unit (CFU) of genome recombinant strain BL21-ΔcbpA/AccbpA showed 7.7 times higher cell viability than deficient strain BL21-ΔcbpA and 2.3 times higher than wild type. Cell morphology observation using Field Emission Scanning Electron Microscopy (FESEM) revealed cell breakage of BL21-ΔcbpA and significant recovery of BL21-ΔcbpA/AccbpA. The intracellular ATP level of all strains gradually decreased along with the increased stress time. Particularly, the value of recombinant strain was 56.0% lower than that of deficient strain after 5 h, indicating that the recombinant strain consumed a lot of energy to resist acid stress. The arginine concentration in BL21-ΔcbpA/AccbpA was double that of BL21-ΔcbpA, while the aspartate and glutamate contents were 14.8% and 6.2% higher, respectively, compared to that of wild type. Moreover, RNA-Seq analysis examined 93 genes down-regulated in BL21-ΔcbpA compared to wild type strain, while 123 genes were up-regulated in BL21-ΔcbpA/AccbpA compared to BL21-ΔcbpA, with an emphasis on energy metabolism, transport, and cell components. Finally, the working model in response to acid stress of cbpA from A. caldus was developed. This study constructed a recombinant strain resistant to acid stress and also provided a reference for enhancing microorganisms’ robustness to various conditions.







Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within this article.
References
Akram M (2014) Citric acid cycle and role of its Intermediates in Metabolism. Cell Biochem Biophys 68(3):475–478. https://doi.org/10.1007/s12013-013-9750-1
Arnold RJ, Reilly JP (1999) Observation of Escherichia coli ribosomal proteins and their posttranslational modifications by mass spectrometry. Anal Biochem 269(1):105–112. https://doi.org/10.1006/abio.1998.3077
Baker P, Hillis C, Carere J, Seah SYK (2012) Protein–protein interactions and substrate channeling in Orthologous and chimeric aldolase–dehydrogenase complexes. Biochemistry 51(9):1942–1952. https://doi.org/10.1021/bi201832a
Bausch C, Peekhaus N, Utz C, Blais T, Murray E, Lowary T, Conway T (1998) Sequence analysis of the GntII (subsidiary) system for gluconate metabolism reveals a novel pathway for L-idonic acid catabolism in Escherichia coli. J Bacteriol 180(14):3704–3710. https://doi.org/10.1128/jb.180.14.3704-3710.1998
Bird JG, Sharma S, Roshwalb SC, Hoskins JR, Wickner S (2006) Functional analysis of CbpA, a DnaJ homolog and nucleoid-associated DNA-binding protein. J Biol Chem 281(45):34349–34356
Bishop RE, Leskiw BK, Hodges RS, Kay CM, Weiner JH (1998) The entericidin locus of Escherichia coli and its implications for programmed bacterial cell death. J Mol Biol 280(4):583–596. https://doi.org/10.1006/jmbi.1998.1894
Brameyer S, Schumacher K, Kuppermann S, Jung K (2022) Division of labor and collective functionality in Escherichia coli under acid stress. Commun Biology 5(1):327. https://doi.org/10.1038/s42003-022-03281-4
Brock M, Maerker C, Schütz A, Völker U, Buckel W (2002) Oxidation of propionate to pyruvate in Escherichia coli. Involvement of methylcitrate dehydratase and aconitase. Eur J Biochem 269(24):6184–6194. https://doi.org/10.1046/j.1432-1033.2002.03336.x
Bubunenko M, Baker T, Court DL (2007) Essentiality of ribosomal and transcription antitermination proteins analyzed by systematic gene replacement in Escherichia coli. J Bacteriol 189(7):2844–2853. https://doi.org/10.1128/jb.01713-06
Chae C, Sharma S, Hoskins JR, Wickner S (2004) CbpA, a DnaJ homolog, is a DnaK co-chaperone, and its activity is modulated by CbpM. J Biol Chem 279(32):33147–33153. https://doi.org/10.1074/jbc.M404862200
Chen MJ, Tang HY, Chiang ML (2017) Effects of heat, cold, acid and bile salt adaptations on the stress tolerance and protein expression of kefir-isolated probiotic Lactobacillus kefiranofaciens M1. Food Microbiol 66:20–27. https://doi.org/10.1016/j.fm.2017.03.020
Chen XK, Li XY, Ha YF, Lin JQ, Liu XM, Pang X, Lin JQ, Chen LX (2020) Ferric Uptake Regulator provides a new strategy for Acidophile Adaptation to Acidic Ecosystems. Appl Environ Microbiol 86(11). https://doi.org/10.1128/aem.00268-20
Chen J, Liu Y, Diep P, Mahadevan R (2022) Genetic engineering of extremely acidophilic Acidithiobacillus species for biomining: progress and perspectives. J Hazard Mater 438:129456. https://doi.org/10.1016/j.jhazmat.2022.129456
Chittori S, Savithri HS, Murthy MR (2011) Preliminary X-ray crystallographic studies on acetate kinase (AckA) from Salmonella typhimurium in two crystal forms. Acta Crystallogr Sect F Struct Biol Cryst Commun 67(Pt 12):1658–1661. https://doi.org/10.1107/s1744309111043740
Cosgriff S, Chintakayala K, Chim YT, Chen X, Allen S, Lovering AL, Grainger DC (2010) Dimerization and DNA-dependent aggregation of the Escherichia coli nucleoid protein and chaperone CbpA. Mol Microbiol 77(5):1289–1300. https://doi.org/10.1111/j.1365-2958.2010.07292.x
Diaz Ricci JC, Hernández ME (2000) Plasmid effects on Escherichia coli metabolism. Crit Rev Biotechnol 20(2):79–108. https://doi.org/10.1080/07388550008984167
Diez-Gonzalez F, Karaibrahimoglu Y (2004) Comparison of the glutamate-, arginine- and lysine-dependent acid resistance systems in Escherichia coli O157:H7. J Appl Microbiol 96(6):1237–1244. https://doi.org/10.1111/j.1365-2672.2004.02251.x
Dong T, Wang W, Xia M, Liang S, Hu G, Ye H, Cao Q, Dong Z, Zhang C, Feng D, Zuo J (2021) Involvement of the heat shock protein HtpG of Salmonella Typhimurium in infection and proliferation in hosts. Front Cell Infect Microbiol 11:758898. https://doi.org/10.3389/fcimb.2021.758898
Feng J, Li C, He H, Xu S, Wang X, Chen K (2022) Construction of cell factory through combinatorial metabolic engineering for efficient production of itaconic acid. Microb Cell Fact 21(1):275. https://doi.org/10.1186/s12934-022-02001-1
Fernández M, Zúñiga M (2006) Amino acid catabolic pathways of lactic acid bacteria. Crit Rev Microbiol 32(3):155–183. https://doi.org/10.1080/10408410600880643
Fountoulakis M, Lahm H-W (1998) Hydrolysis and amino acid composition analysis of proteins. J Chromatogr A 826(2):109–134. https://doi.org/10.1016/S0021-9673(98)00721-3
Fujino T, Kondo J, Ishikawa M, Morikawa K, Yamamoto TT (2001) Acetyl-CoA synthetase 2, a mitochondrial matrix enzyme involved in the oxidation of acetate. J Biol Chem 276(14):11420–11426. https://doi.org/10.1074/jbc.M008782200
Gao X, Yang X, Li J, Zhang Y, Chen P, Lin Z (2018) Engineered global regulator H-NS improves the acid tolerance of E. coli. Microb Cell Fact 17(1):118. https://doi.org/10.1186/s12934-018-0966-z
Genest O, Hoskins JR, Kravats AN, Doyle SM, Wickner S (2015) Hsp70 and Hsp90 of E. coli directly interact for collaboration in protein remodeling. J Mol Biol 427(24):3877–3889. https://doi.org/10.1016/j.jmb.2015.10.010
Grueninger D, Schulz GE (2006) Structure and reaction mechanism of l-Rhamnulose kinase from Escherichia coli. J Mol Biol 359(3):787–797. https://doi.org/10.1016/j.jmb.2006.04.013
Guan N, Liu L (2020) Microbial response to acid stress: mechanisms and applications. Appl Microbiol Biotechnol 104(1):51–65. https://doi.org/10.1007/s00253-019-10226-1
Guan N, Li J, Shin HD, Du G, Chen J, Liu L (2017) Microbial response to environmental stresses: from fundamental mechanisms to practical applications. Appl Microbiol Biotechnol 101(10):3991–4008. https://doi.org/10.1007/s00253-017-8264-y
Guo J, Ma Z, Gao J, Zhao J, Wei L, Liu J, Xu N (2019) Recent advances of pH homeostasis mechanisms in Corynebacterium glutamicum. World J Microbiol Biotechnol 35(12):192. https://doi.org/10.1007/s11274-019-2770-2
Harnischfeger J, Beutler M, Salzig D, Rahlfs S, Becker K, Grevelding CG, Czermak P (2021) Biochemical characterization of the recombinant schistosome tegumental protein SmALDH_312 produced in E. coli and baculovirus expression vector system. Electron J Biotechnol 54:26–36. https://doi.org/10.1016/j.ejbt.2021.08.002
He W, Li C, Lu CD (2011) Regulation and characterization of the dadRAX locus for D-amino acid catabolism in Pseudomonas aeruginosa PAO1. J Bacteriol 193(9):2107–2115. https://doi.org/10.1128/jb.00036-11
Isupov MN, Antson AA, Dodson EJ, Dodson GG, Dementieva IS, Zakomirdina LN, Wilson KS, Dauter Z, Lebedev AA, Harutyunyan EH (1998) Crystal structure of tryptophanase. J Mol Biol 276(3):603–623. https://doi.org/10.1006/jmbi.1997.1561
Itoh Y (1997) Cloning and characterization of the aru genes encoding enzymes of the catabolic arginine succinyltransferase pathway in Pseudomonas aeruginosa. J Bacteriol 179(23):7280–7290. https://doi.org/10.1128/jb.179.23.7280-7290.1997
Jiang Y, Chen B, Duan C, Sun B, Yang J, Yang S (2015) Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Appl Environ Microbiol 81(7):2506–2514. https://doi.org/10.1128/aem.04023-14
Jomaa A, Jain N, Davis JH, Williamson JR, Britton RA, Ortega J (2014) Functional domains of the 50S subunit mature late in the assembly process. Nucleic Acids Res 42(5):3419–3435. https://doi.org/10.1093/nar/gkt1295
Kang H-J, Heo D-H, Choi S-W, Kim K-N, Shim J, Kim C-W, Sung H-C, Yun C-W (2007) Functional characterization of Hsp33 protein from Bacillus psychrosaccharolyticus; additional function of HSP33 on resistance to solvent stress. Biochem Biophys Res Commun 358(3):743–750. https://doi.org/10.1016/j.bbrc.2007.04.184
Kapatral V, Bina X, Chakrabarty AM (2000) Succinyl coenzyme A synthetase of Pseudomonas aeruginosa with a broad specificity for nucleoside triphosphate (NTP) synthesis modulates specificity for NTP synthesis by the 12-kilodalton form of nucleoside diphosphate kinase. J Bacteriol 182(5):1333–1339. https://doi.org/10.1128/jb.182.5.1333-1339.2000
Kawazoe N, Kimata Y, Izawa S (2017) Acetic acid causes endoplasmic reticulum stress and induces the unfolded protein response in Saccharomyces cerevisiae. Front Microbiol 8:1192. https://doi.org/10.3389/fmicb.2017.01192
Krulwich TA, Sachs G, Padan E (2011) Molecular aspects of bacterial pH sensing and homeostasis. Nat Reviews: Microbiol 9(5):330–343. https://doi.org/10.1038/nrmicro2549
Kurihara S, Suzuki H, Oshida M, Benno Y (2011) A Novel Putrescine Importer required for type 1 Pili-driven Surface Motility Induced by Extracellular Putrescine in Escherichia coli K-12*. J Biol Chem 286(12):10185–10192. https://doi.org/10.1074/jbc.M110.176032
Lacroix JM, Lanfroy E, Cogez V, Lequette Y, Bohin A, Bohin JP (1999) The mdoC gene of Escherichia coli encodes a membrane protein that is required for succinylation of osmoregulated periplasmic glucans. J Bacteriol 181(12):3626–3631. https://doi.org/10.1128/jb.181.12.3626-3631.1999
Le Vo TD, Ko J-s, Park SJ, Lee SH, Hong SH (2013) Efficient gamma-aminobutyric acid bioconversion by employing synthetic complex between glutamate decarboxylase and glutamate/GABA antiporter in engineered Escherichia coli. J Ind Microbiol Biotechnol 40(8):927–933. https://doi.org/10.1007/s10295-013-1289-z
Li Z, Jiang B, Zhang X, Yang Y, Hardwidge PR, Ren W, Zhu G (2020) The role of bacterial cell envelope structures in acid stress resistance in E. coli. Appl Microbiol Biotechnol 104(7):2911–2921. https://doi.org/10.1007/s00253-020-10453-x
Lin Z, Li J, Yan X, Yang J, Li X, Chen P, Yang X (2021) Engineering of the small noncoding RNA (sRNA) DsrA together with the sRNA chaperone hfq enhances the Acid Tolerance of Escherichia coli. Appl Environ Microbiol 87(10). https://doi.org/10.1128/aem.02923-20
Liu Y, Tang H, Lin Z, Xu P (2015) Mechanisms of acid tolerance in bacteria and prospects in biotechnology and bioremediation. Biotechnol Adv 33(7):1484–1492. https://doi.org/10.1016/j.biotechadv.2015.06.001
Liu H, Qi Y, Zhou P, Ye C, Gao C, Chen X, Liu L (2021) Microbial physiological engineering increases the efficiency of microbial cell factories. Crit Rev Biotechnol 41(3):339–354. https://doi.org/10.1080/07388551.2020.1856770
Locher KP (2016) Mechanistic diversity in ATP-binding cassette (ABC) transporters. Nat Struct Mol Biol 23(6):487–493. https://doi.org/10.1038/nsmb.3216
Luna L, Bjørås M, Hoff E, Rognes T, Seeberg E (2000) Cell-cycle regulation, intracellular sorting and induced overexpression of the human NTH1 DNA glycosylase involved in removal of formamidopyrimidine residues from DNA. Mutat Res 460(2):95–104. https://doi.org/10.1016/s0921-8777(00)00015-x
Lund P, Tramonti A, De Biase D (2014) Coping with low pH: molecular strategies in neutralophilic bacteria. FEMS Microbiol Rev 38(6):1091–1125. https://doi.org/10.1111/1574-6976.12076
Marc J, Grousseau E, Lombard E, Sinskey AJ, Gorret N, Guillouet SE (2017) Over expression of GroESL in Cupriavidus necator for heterotrophic and autotrophic isopropanol production. Metab Eng 42:74–84. https://doi.org/10.1016/j.ymben.2017.05.007
Márquez J, de la Oliva AR, Matés JM, Segura JA, Alonso FJ (2006) Glutaminase: a multifaceted protein not only involved in generating glutamate. Neurochem Int 48(6–7):465–471. https://doi.org/10.1016/j.neuint.2005.10.015
Martínez-Reyes I, Chandel NS (2020) Mitochondrial TCA cycle metabolites control physiology and disease. Nat Commun 11(1):102. https://doi.org/10.1038/s41467-019-13668-3
Matuszewska M, Kuczyńska-Wiśnik D, Laskowska E, Liberek K (2005) The small heat shock protein IbpA of Escherichia coli cooperates with IbpB in stabilization of thermally aggregated proteins in a disaggregation competent state. J Biol Chem 280(13):12292–12298. https://doi.org/10.1074/jbc.M412706200
Mayer MP (2021) The Hsp70-Chaperone machines in Bacteria. Front Mol Biosci 8:694012. https://doi.org/10.3389/fmolb.2021.694012
Minárik P, Tomásková N, Kollárová M, Antalík M (2002) Malate dehydrogenases–structure and function. Gen Physiol Biophys 21(3):257–265
Molan K, Žgur Bertok D (2022) Small prokaryotic DNA-Binding proteins protect Genome Integrity throughout the life cycle. Int J Mol Sci 23(7). https://doi.org/10.3390/ijms23074008
Monteagudo-Cascales E, Martín-Mora D, Xu W, Sourjik V, Matilla MA, Ortega Á, Krell T (2022) The pH robustness of bacterial sensing. mBio 13(5):e0165022. https://doi.org/10.1128/mbio.01650-22
Moosavi B, Zhu XL, Yang WC, Yang GF (2020) Genetic, epigenetic and biochemical regulation of succinate dehydrogenase function. Biol Chem 401(3):319–330. https://doi.org/10.1515/hsz-2019-0264
Moxley MA, Tanner JJ, Becker DF (2011) Steady-state kinetic mechanism of the proline:ubiquinone oxidoreductase activity of proline utilization A (PutA) from Escherichia coli. Arch Biochem Biophys 516(2):113–120. https://doi.org/10.1016/j.abb.2011.10.011
Nogly P, Gushchin I, Remeeva A, Esteves AM, Borges N, Ma P, Ishchenko A, Grudinin S, Round E, Moraes I, Borshchevskiy V, Santos H, Gordeliy V, Archer M (2014) X-ray structure of a CDP-alcohol phosphatidyltransferase membrane enzyme and insights into its catalytic mechanism. Nat Commun 5:4169. https://doi.org/10.1038/ncomms5169
Ohide H, Miyoshi Y, Maruyama R, Hamase K, Konno R (2011) D-Amino acid metabolism in mammals: biosynthesis, degradation and analytical aspects of the metabolic study. J Chromatogr B Analyt Technol Biomed Life Sci 879(29):3162–3168. https://doi.org/10.1016/j.jchromb.2011.06.028
Okamura-Ikeda K, Ohmura Y, Fujiwara K, Motokawa Y (1993) Cloning and nucleotide sequence of the gcv operon encoding the Escherichia coli glycine-cleavage system. Eur J Biochem 216(2):539–548. https://doi.org/10.1111/j.1432-1033.1993.tb18172.x
Okochi M, Kanie K, Kurimoto M, Yohda M, Honda H (2008) Overexpression of prefoldin from the hyperthermophilic archaeum Pyrococcus horikoshii OT3 endowed Escherichia coli with organic solvent tolerance. Appl Microbiol Biotechnol 79(3):443–449. https://doi.org/10.1007/s00253-008-1450-1
Peetermans A, Foulquié-Moreno MR, Thevelein JM (2021) Mechanisms underlying lactic acid tolerance and its influence on lactic acid production in Saccharomyces cerevisiae. Microb Cell 8(6):111–130. https://doi.org/10.15698/mic2021.06.751
Pepe S, Scarlato V, Roncarati D (2020) The Helicobacter pylori HspR-Modulator CbpA is a multifunctional heat-shock protein. Microorganisms 8(2). https://doi.org/10.3390/microorganisms8020251
Piper P, Calderon CO, Hatzixanthis K, Mollapour M (2001) Weak acid adaptation: the stress response that confers yeasts with resistance to organic acid food preservatives. Microbiology 147(Pt 10):2635–2642. https://doi.org/10.1099/00221287-147-10-2635
Prud’homme-Généreux A, Beran RK, Iost I, Ramey CS, Mackie GA, Simons RW (2004) Physical and functional interactions among RNase E, polynucleotide phosphorylase and the cold-shock protein, CsdA: evidence for a ‘cold shock degradosome’. Mol Microbiol 54(5):1409–1421. https://doi.org/10.1111/j.1365-2958.2004.04360.x
Reeves PJ, Whitcombe D, Wharam S, Gibson M, Allison G, Bunce N, Barallon R, Douglas P, Mulholland V, Stevens S (1993) Molecular cloning and characterization of 13 out genes from Erwinia carotovora subspecies carotovora: genes encoding members of a general secretion pathway (GSP) widespread in gram-negative bacteria. Mol Microbiol 8(3):443–456. https://doi.org/10.1111/j.1365-2958.1993.tb01589.x
Roncarati D, Scarlato V (2017) Regulation of heat-shock genes in bacteria: from signal sensing to gene expression output. FEMS Microbiol Rev 41(4):549–574. https://doi.org/10.1093/femsre/fux015
Rouches MV, Xu Y, Cortes LBG, Lambert G (2022) A plasmid system with tunable copy number. Nat Commun 13(1):3908. https://doi.org/10.1038/s41467-022-31422-0
Saibil H (2013) Chaperone machines for protein folding, unfolding and disaggregation. Nat Reviews: Mol Cell Biology 14(10):630–642. https://doi.org/10.1038/nrm3658
Sarraf NS, Baardsnes J, Cheng J, O’Connor-McCourt M, Cygler M, Ekiel I (2010) Structural basis of the regulation of the CbpA co-chaperone by its specific modulator CbpM. J Mol Biol 398(1):111–121. https://doi.org/10.1016/j.jmb.2010.03.006
Sarraf NS, Shi R, McDonald L, Baardsnes J, Zhang L, Cygler M, Ekiel I (2014) Structure of CbpA J-domain bound to the regulatory protein Cbpm explains its specificity and suggests evolutionary link between Cbpm and transcriptional regulators. PLoS ONE 9(6):e100441. https://doi.org/10.1371/journal.pone.0100441
Shirai H, Mizuguchi K (2003) Prediction of the structure and function of AstA and AstB, the first two enzymes of the arginine succinyltransferase pathway of arginine catabolism. FEBS Lett 555(3):505–510. https://doi.org/10.1016/s0014-5793(03)01314-0
Silhavy TJ, Kahne D, Walker S (2010) The bacterial cell envelope. Cold Spring Harb Perspect Biol 2(5):a000414. https://doi.org/10.1101/cshperspect.a000414
Stewart JB, Hermodson MA (2003) Topology of RbsC, the membrane component of the Escherichia coli ribose transporter. J Bacteriol 185(17):5234–5239. https://doi.org/10.1128/jb.185.17.5234-5239.2003
Ueta M, Wada C, Bessho Y, Maeda M, Wada A (2017) Ribosomal protein L31 in Escherichia coli contributes to ribosome subunit association and translation, whereas short L31 cleaved by protease 7 reduces both activities. Genes Cells 22(5):452–471. https://doi.org/10.1111/gtc.12488
Ultee E, Ramijan K, Dame RT, Briegel A, Claessen D (2019) Stress-induced adaptive morphogenesis in bacteria. Adv Microb Physiol 74:97–141. https://doi.org/10.1016/bs.ampbs.2019.02.001
Valentin-Hansen P, Boëtius F, Hammer-Jespersen K, Svendsen I (1982) The primary structure of Escherichia coli K12 2-deoxyribose 5-phosphate aldolase. Nucleotide sequence of the deoC gene and the amino acid sequence of the enzyme. Eur J Biochem 125(3):561–566. https://doi.org/10.1111/j.1432-1033.1982.tb06719.x
VanOrsdel CE, Bhatt S, Allen RJ, Brenner EP, Hobson JJ, Jamil A, Haynes BM, Genson AM, Hemm MR (2013) The Escherichia coli CydX protein is a member of the CydAB cytochrome bd oxidase complex and is required for cytochrome bd oxidase activity. J Bacteriol 195(16):3640–3650. https://doi.org/10.1128/jb.00324-13
Vinnemeier J, Hagemann M (1999) Identification of salt-regulated genes in the genome of the cyanobacterium Synechocystis sp. strain PCC 6803 by subtractive RNA hybridization. Arch Microbiol 172(6):377–386. https://doi.org/10.1007/s002030050774
Wall D, Zylicz M, Georgopoulos C (1994) The NH2-terminal 108 amino acids of the Escherichia coli DnaJ protein stimulate the ATPase activity of DnaK and are sufficient for lambda replication. J Biol Chem 269(7):5446–5451. https://doi.org/10.1016/S0021-9258(17)37706-2
Wang J, Wang W, Wang H, Yuan F, Xu Z, Yang K, Li Z, Chen Y, Fan K (2019) Improvement of stress tolerance and riboflavin production of Bacillus subtilis by introduction of heat shock proteins from thermophilic bacillus strains. Appl Microbiol Biotechnol 103(11):4455–4465. https://doi.org/10.1007/s00253-019-09788-x
Wang C, Ren X, Yu C, Wang J, Wang L, Zhuge X, Liu X (2020) Physiological and transcriptional responses of Streptomyces albulus to acid stress in the biosynthesis of ε-Poly-L-lysine. Front Microbiol 11:1379. https://doi.org/10.3389/fmicb.2020.01379
Weidmann S, Maitre M, Laurent J, Coucheney F, Rieu A, Guzzo J (2017) Production of the small heat shock protein Lo18 from Oenococcus oeni in Lactococcus lactis improves its stress tolerance. Int J Food Microbiol 247:18–23. https://doi.org/10.1016/j.ijfoodmicro.2016.06.005
Yang M, Zhang X (2017) Construction of pyruvate producing strain with intact pyruvate dehydrogenase and genome-wide transcription analysis. World J Microbiol Biotechnol 33(3):59. https://doi.org/10.1007/s11274-016-2202-5
Yang DC, Blair KM, Salama NR (2016) Staying in shape: the impact of cell shape on bacterial survival in diverse environments. Microbiol Mol Biol Rev 80(1):187–203. https://doi.org/10.1128/mmbr.00031-15
Yin Y, Tong Y, Yang H, Feng S (2022) EpsR(ac) is a copper-sensing MarR family transcriptional repressor from Acidithiobacillus caldus. Appl Microbiol Biotechnol 106(9–10):3679–3689. https://doi.org/10.1007/s00253-022-11971-6
Zhu Z, Yang J, Yang P, Wu Z, Zhang J, Du G (2019) Enhanced acid-stress tolerance in Lactococcus lactis NZ9000 by overexpression of ABC transporters. Microb Cell Fact 18(1):136. https://doi.org/10.1186/s12934-019-1188-8
Acknowledgements
This study was supported by grants from the National Key Research and Development Program of China (2022YFC3401300), the National Natural Science Foundation of China (No. 21878128; 21776113; 21606110; 31701582), the funding of Key Laboratory of Industrial Biotechnology, Ministry of Education (KLIBKF202005), the Priority Academic Program Development of Jiangsu Higher Education Institutions, and Program of Introducing Talents of Discipline to Universities (No. 111-2-06).
Author information
Authors and Affiliations
Contributions
Zhenming Jiang and Shoushuai Feng designed the research and analyzed the data. Zhenming Jiang and Jie Lu conducted experiments. Zhenming Jiang wrote the first draft of the manuscript. Shoushuai Feng, Hailin Yang, and Yanjun Tong contributed to manuscript revision, read, and approved the submitted version. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiang, Z., Lu, J., Tong, Y. et al. Enhancement of acid tolerance of Escherichia coli by introduction of molecule chaperone CbpA from extremophile. World J Microbiol Biotechnol 39, 158 (2023). https://doi.org/10.1007/s11274-023-03613-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03613-4