Abstract
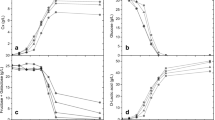
In lactobacilli, CcpA is known to modulate the expression of genes involved in sugar metabolism, stress response and aerobic adaptation. This study aimed to evaluate a ccpA mutant of Lacticaseibacillus casei BL23 to increase lactic acid production using cheese whey. The ccpA derivative (BL71) showed better growth than the L. casei wild-type in the whey medium. In a stirred tank reactor, at 48 h, lactate production by BL71 was eightfold higher than that by BL23. In batch fermentations, the final values reached were 44.23 g L−1 for BL71 and 27.58 g L−1 for BL23. Due to a decrease in the delay of lactate production in the mutant, lactate productivity increased from 0.17 g (L.h)−1 with BL23 to 0.80 g (L.h)−1 with BL71. We found that CcpA would play additional roles in nitrogen metabolism by the regulation of the proteolytic system. BL71 displayed higher activity of the PepX, PepQ and PrtP enzymes than BL23. Analysis of prtP expression confirmed this deregulation in BL71. Promoter analysis of the prtP gene revealed CcpA binding sites with high identity to the cre consensus sequence and the interaction of CcpA with this promoter was confirmed in vitro. We postulate that deregulation of the proteolytic system in BL71 allows a better exploitation of nitrogen resources in cheese whey, resulting in enhanced fermentation capacity. Therefore, the ccpA gene could be a good target for future technological developments aimed at effective and inexpensive lactate production from dairy industrial wastes.
Graphic abstract






Similar content being viewed by others
References
Abdel-Rahman MA, Tashiro Y, Sonomoto K (2013) Recent advances in lactic acid production by microbial fermentation processes. Biotechnol Adv 31:877–902. https://doi.org/10.1016/j.biotechadv.2013.04.002
Acedo-Félix E, Pérez-Martínez G (2003) Significant differences between Lactobacillus casei subsp. casei ATCC 393T and a commonly used plasmid-cured derivative revealed by a polyphasic study. Int J Syst Evol Microbiol 53:67–75. https://doi.org/10.1099/ijs.0.02325-0
Alcántara C, Bäuerl C, Revilla-Guarinos A, Pérez-Martínez G, Monedero V, Zúñiga M (2016) Peptide and amino acid metabolism is controlled by an OmpR-family response regulator in Lactobacillus casei. Mol Microbiol 100(1):25–41
Allievi MC, Palomino MM, Ruzal SM (2019) Production of lactate using Lactobacillus. In: Ruzal SM (ed) Lactobacillus genomics and metabolic engineering caister. Academic Press, UK, pp 61–80. https://doi.org/10.21775/9781910190890.04
Alvarez MM, Aguirre-Ezkauriatza EJ, Ramírez-Medrano A, Rodríguez-Sánchez Á (2010) Kinetic analysis and mathematical modeling of growth and lactic acid production of Lactobacillus casei var. rhamnosus in milk whey. J Dairy Sci 93:5552–5560
Bernardo MP, Coelho LF, Sass DC, Contiero J (2016) L-(+)-lactic acid production by Lactobacillus rhamnosus B103 from dairy industry waste. Braz J Microbiol 47(3):640–646. https://doi.org/10.1016/j.bjm.2015.12.001
Brown L, Villegas JM, Elean M, Fadda S, Mozzi F, Saavedra L, Hebert EM (2017) YebC, a putative transcriptional factor involved in the regulation of the proteolytic system of Lactobacillus. Sci Rep 7(1):8579. https://doi.org/10.1038/s41598-017-09124-1
Carvalho F, Prazeres AR, Rivas J (2013) Cheese whey wastewater: characterization and treatment. Sci Total Environ 445–446:385–396. https://doi.org/10.1016/j.scitotenv.2012.12.038
Coll-Marqués JM, Bäuerl C, Zúñiga M, Pérez-Martínez G (2020) Differences in the expression of cell envelope proteinases (CEP) in two Lactobacillus paracasei probiotic strains. FEMS Microbiol Lett 367(13):fnaa102. https://doi.org/10.1093/femsle/fnaa102
Dandoy D, Fremaux C, Henry de Frahan M, Horvath P, Boyaval P, Hols P et al (2011) The fast milk acidifying phenotype of Streptococcus thermophilus can be acquired by natural transformation of the genomic island encoding the cell-envelope proteinase PrtS. Microb Cell Fact 10:S21
Deutscher J (2008) The mechanisms of carbon catabolite repression in bacteria. Curr Opinion Microbiol 11:87–93
Francke C, Kerkhoven R, Wels M, Siezen RJ (2008) A generic approach to identify transcription factor-specific operator motifs; Inferences for LacI-family mediated regulation in Lactobacillus plantarum WCFS1. BMC Genomics 9:145
Fujita Y (2009) Carbon catabolite control of the metabolic network in Bacillus subtilis. Biosci Biotech Biochem 73:245–259
Ghosh P, Wasil LR, Hatfull GF (2006) Control of phage Bxb1 excision by a novel recombination directionality factor. PLoS Biol 4(6):e186
Guédon E, Martin C, Gobert FX, Ehrlich DS, Renault P, Delorme C (2001) Réseau de régulation de la transcription des genes du systeme protéolytique de Lactococcus lactis. Lait 81:65–74
Gutmann I, Wahlefeld AW (1974) Lactate measurements. In: Bergmeyer HU (ed) Methods of Enzymatic Analysis. Verlag Chemie, Weinhein, Academic Press, New York, pp 1464–1468
John RP, Nampoothiri KM, Pandey A (2007) Fermentative production of lactic acid from biomass: an overview on process developments and future perspectives. Appl Microbiol Biotechnol 74(3):524–534
Kleerebezem M, Boekhorst J, van Kranenburg R, Molenaar D, Kuipers OP et al (2003) Complete genome sequence of Lactobacillus plantarum WCFS1. Proc Natl Acad Sci USA 100(4):1990–1995
Küster E, Luesink EJ, de Vos WM, Hillen W (1996) Immunological crossreactivity to the catabolite control protein CcpA from Bacillus megaterium is found in many Gram-positive bacteria. FEMS Microbiol Lett 139(2–3):109–115
Larionov A, Krause A, Miller W (2005) A standard curve based method for relative real time PCR data processing. BMC Bioinformatics 6(1):62
Li C, Sun F, Cho H, Yelavarthi V, Sohn C et al (2010) CcpA mediates proline auxotrophy and is required for Staphylococcus aureus pathogenesis. J Bacteriol 192(15):3883–3892
Luesink EJ, Beumer CM, Kuipers OP, de Vos WM (1999) Molecular characterization of the Lactococcus lactis ptsHI operon and analysis of the regulatory role of HPr. J Bacteriol 181:764–771
Mahr K, Hillen W, Titgemeyer F (2000) Carbon catabolite repression in Lactobacillus pentosus: analysis of the ccpA region. Appl Environ Microbiol 66:277–283
Marciniak BC, Pabijaniak M, de Jong A, Dűhring R, Seidel G, Hillen W, Kuipers OP (2012) High- and low-affinity cre boxes for CcpA binding in Bacillus subtilis revealed by genome-wide analysis. BMC Genomics 13:401. https://doi.org/10.1186/1471-2164-13-401
Mazé A et al (2010) Complete genome sequence of the probiotic Lactobacillus casei strain BL23. J Bacteriol 192:2647–2648
Mazzeo MF, Cacace G, Peluso A, Zotta T, Muscariello L, Vastano V, Parente E, Siciliano RA (2012) Effect of inactivation of ccpA and aerobic growth in Lactobacillus plantarum: a proteomic perspective. J Proteomics 75:4050–4061. https://doi.org/10.1016/j.jprot.2012.05.019
Monedero V, Gosalbes MJ, Pérez-Martínez G (1997) Catabolite repression in Lactobacillus casei ATCC 393 is mediated by CcpA. J Bacteriol 179:6657–6664
Monedero V, Mazé A, Boël G, Zúñiga M, Beaufils S, Hartke A, Deutscher J (2007) The phosphotransferase system of Lactobacillus casei: regulation of carbon metabolism and connection to cold shock response. J Mol Microbiol Biotechnol 12(1–2):20–32
Morishita T, Fukada T, Shirota M, Yura T (1974) Genetic basis of nutritional requirements in Lactobacillus casei. J Bacteriol 120(3):1078–1084
Narayanan N, Roychoudhury PK, Srivastava A (2004) L (+) lactic acid fermentation and its product polymerization. Electron J Biotechnol 7(2):167–178
Palomino MM, Waehner PM, Fina Martin J, Ojeda P, Malone L et al (2016) Influence of osmotic stress on the profile and gene expression of surface layer proteins in Lactobacillus acidophilus ATCC 4356. Appl Microbiol Biotechnol 100(19):8475–8484
Panesar PS, Kennedy JF, Gandhi DN, Bunko K (2007) Bioutilisation of whey for lactic acid production. Food Chem 105:1–14. https://doi.org/10.1016/j.foodchem.2007.03.035
Petranovic D, Guédon E, Sperandio B, Delorme C, Ehrlich D, Renault P (2004) Intracellular effectors regulating the activity of the Lactococcus lactis CodY pleiotropic transcription regulator. Mol Microbiol 53:613–621
Piuri M, Sanchez-Rivas C, Ruzal SM (2003) Adaptation to high salt in Lactobacillus: role of peptides and proteolytic enzymes. J Appl Microbiol 95:372–379
Rabaioli Rama G, Kuhn D, Beux S, Jachetti Maciel M, Volken de Souza CF (2019) Potential applications of dairy whey for the production of lactic acid bacteria cultures. Int Dairy J 98:25–37. https://doi.org/10.1016/j.idairyj.2019.06.012
Rey DA, Pühler A, Kalinowski J (2003) The putative transcriptional repressor McbR, member of the TetR-family, is involved in the regulation of the metabolic network directing the synthesis of sulfur containing amino acids in Corynebacterium glutamicum. J Biotechnol 103:51–65
Ricciardi A, Zotta T, Ianniello RG, Boscaino F, Matera A, Parente E (2019) Effect of respiratory growth on the metabolite production and stress robustness of Lactobacillus casei N87 cultivated in cheese whey permeate medium. Front Microbiol 10:851. https://doi.org/10.3389/fmicb.2019.00851
Sahoo TK, Jayaraman G (2019) Co-culture of Lactobacillus delbrueckii and engineered Lactococcus lactis enhances stoichiometric yield of d-lactic acid from whey permeate. Appl Microbiol Biotechnol 103(14):5653–5662
Schick J, Weber B, Klein JR, Henrich B (1999) PepR1, a CcpA-like transcription regulator of Lactobacillus delbrueckii subsp. lactis. Microbiology 145(11):3147–3154
Siciliano RA, Pannella G, Lippolis R, Ricciardi A, Mazzeo F, Zotta T (2019) Impact of aerobic and respirative life-style on Lactobacillus casei N87 proteome. Int J Food Microbiol 298:51–62
Singh SK, Ahmed SU, Pandey A (2006) Metabolic engineering approaches for lactic acid production. Proc Biochem 41:991–1000
Sonenshein AL (2005) CodY, a global regulator of stationary phase and virulence in Gram-positive bacteria. Curr Opin Microbiol 8:203–207
Sonenshein AL (2007) Control of key metabolic intersections in Bacillus subtilis. Nat Rev Microbiol 5(12):917–927
Spatafora G, Rohrer K, Barnard D, Michalek S (1995) A Streptococcus mutans mutant that synthesizes elevated levels of intracellular polysaccharide is hypercariogenic in vivo. Infect Immun 63:2556–2563
Viana R, Pérez-Martínez G, Deutscher J, Monedero V (2005) The glycolytic genes pfk and pyk from Lactobacillus casei are induced by sugars transported by the phosphoenolpyruvate: sugar phosphotransferase system and repressed by CcpA. Arch Microbiol 183(6):385–393. https://doi.org/10.1007/s00203-005-0003-6
Wang J, Wang Q, Xu Z, Zhang W, Xiang J (2015) Effect of fermentation conditions on L-lactic acid production from soybean straw hydrolysate. J Microbiol Biotechnol 25(1):26–32
Wakai T, Yamamoto N (2013) A Novel branched chain amino acids responsive transcriptional regulator, BCARR, negatively acts on the proteolytic system in Lactobacillus helveticus. PLoS ONE 8(10):e75976. https://doi.org/10.1371/journal.pone.0075976
Wünsche A, Hammer E, Bartholomae M, Völker U, Burkovski A, Seidel G, Hillen W (2012) CcpA forms complexes with CodY and RpoA in Bacillus subtilis. FEBS J 279(12):2201–2214. https://doi.org/10.1111/j.1742-4658.2012.08604.x
Zhang G, Li LL, C, (2020) Effects of ccpA gene deficiency in Lactobacillus delbrueckii subsp. bulgaricus under aerobic conditions as assessed by proteomic analysis. Microb Cell Fact 19(1):1–12. https://doi.org/10.1186/s12934-020-1278-7
Zheng J, Wittouck S, Salvetti E, Franz C, Harris HMB, Mattarelli P et al (2020) A taxonomic note on the genus Lactobacillus: description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int J Syst Evol Microbiol 70:2782–2858. https://doi.org/10.1099/ijsem.0.004107
Zomer AL, Buist G, Larsen R, Kok J, Kuipers OP (2007) Time-resolved determination of the CcpA regulon of Lactococcus lactis subsp. cremoris MG1363. J Bacteriol 189:1366–1381
Zotta T, Ricciardi A, Guidone A, Sacco M, Muscariello L, Mazzeo MF (2012) Inactivation of ccpA and aeration affect growth, metabolite production and stress tolerance of Lactobacillus plantarum WCFS1. Int J Food Microbiol 155:51–59. https://doi.org/10.1016/j.ijfoodmicro.2012.01.017
Zotta T, Parente E, Ricciardi A (2017) Aerobic metabolism in the genus Lactobacillus: impact on stress response and potential applications in the food industry. J Appl Microbiol 122:857–869. https://doi.org/10.1111/jam.13399
Zotta T, Solieri L, Iacumin L, Picozzi C, Gullo M (2020) Valorization of cheese whey using microbial fermentations. Appl Microbiol Biotechnol 104:2749–2764. https://doi.org/10.1007/s00253-020-10408-2
Funding
The present report was supported by grants from the Universidad de Buenos Aires (UBA) (UBACyT 20020170200329BA and 20020170100019BA) and Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), Argentina. JFM is a graduate fellow of CONICET; VMG is a member of CSIC (Spain); MVC and DML are members of INTI (Argentina) and MMP, SMR and MCA are members of CONICET (Argentina).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Catone, M.V., Palomino, M.M., Legisa, D.M. et al. Lactic acid production using cheese whey based medium in a stirred tank reactor by a ccpA mutant of Lacticaseibacillus casei. World J Microbiol Biotechnol 37, 61 (2021). https://doi.org/10.1007/s11274-021-03028-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-021-03028-z