Abstract
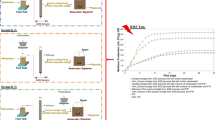
Biogas reactors are now a common part of wastewater treatment systems. The quality of produced biogas is the result of many factors, mainly the input substrate and microbial composition of the bioreactor. The aim of this research was to evaluate the microbial community of the Modřice biogas reactor together with the possible changes in biogas composition. The key microbial groups and their content in anaerobic digester were identified by sequencing techniques. The most dominant group were sulphate-reducing (45%), followed by methanogenic (19%), acetate (6%) and hydrogen-producing (11%) microorganisms. The remaining microorganisms were identified only to their order (19%). Phylogenetic trees were constructed to show evolutionary relationships of detected microorganisms. The volume of methane in biogas content was 60%, which corresponds with literature data regarding sewage digesters. None of the detected impurities have crossed the safe limits and their volume remained stable during the measurement period. Despite sulphate-reducing bacteria being the dominant group, their produced hydrogen sulphide (H2S) was detected only in a small quantity (2.43–7.46 ppm) and had no inhibitory effect on the methane production. The mechanism of inhibition by H2S and the perspective of its biological removal were discussed. Application of phototrophic sulphur bacteria, especially Chlorobiaceae and Chromatiaceae family, and the creation of new photobioreactor systems can be a promising pathway for hydrogen sulphide treatment in biogas plants.






Similar content being viewed by others
References
Altschul, S. F., Madden, T. L., Schäffer, A. A., Zhang, J., Zhang, Z., Miller, W., & Lipman, D. J. (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Research, 25(17), 3389–3402.
Barton, L. L., & Hamilton, W. A. (Eds.). (2007). Sulphate-reducing bacteria (1st ed.). Cambridge: Cambridge University Press.
Boone, D. R., Castenholz, R. W., & Garrity, G. M. (2012). Bergey’s manual of systematic bacteriology (2nd ed.). New York: Springer.
Bryant, M. P. (1979). Microbial methane production—theoretical aspects 2. Journal of Animal Science, 48(1), 193–201.
Buisman, C., Post, R., Ijspeert, P., Geraats, G., & Lettinga, G. (1989). Biotechnological process for sulphide removal with sulphur reclamation. Acta Biotechnologica, 9(3), 255–267.
Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nature Methods, 7(5), 335–336.
Černý, M., Vítězová, M., Vítěz, T., Bartoš, M., & Kushkevych, I. (2018). Variation in the distribution of hydrogen producers from the Clostridiales order in biogas reactors depending on different input substrates. Energies, 11(12).
Chen, Y., Cheng, J. J., & Creamer, K. S. (2008). Inhibition of anaerobic digestion process: a review. Bioresource Technology, 99(10), 4044–4064.
Colleran, E., Finnegan, S., & Lens, P. (1995). Anaerobic treatment of sulphate-containing waste streams. Antonie Van Leeuwenhoek, 67(1), 29–46.
Conrad, R. (1999). Contribution of hydrogen to methane production and control of hydrogen concentrations in methanogenic soils and sediments. FEMS Microbiology Ecology, 28(3), 193–202.
Copeland, A., Spring, S., Göker, M., Schneider, S., Lapidus, A., Del Rio, T. G., et al. (2009). Complete genome sequence of Desulfomicrobium baculatum type strain (XT). Standards in Genomic Sciences, 1(1), 29–37.
CSN EN 121761 Characterization of sludge—determination of pH-value. (1999). Prague: Czech Standards Institute.
CSN EN 14346 Characterization of waste—calculation of dry matter by determination of dry residue or water content. (2007). Prague: Czech Standards Institute.
CSN EN 15169 Characterization of waste—determination of loss on ignition in waste, sludge and sediments. (2007). Prague: Czech Standards Institute.
Dahl, C., Hell, R., Leustek, T., & Knaff, D. (2008). Introduction to sulfur metabolism in phototrophic organisms. In Sulfur metabolism in phototrophic organisms (pp. 1–14). Dordrecht: Springer.
Demirel, B., & Scherer, P. (2008). The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Reviews in Environmental Science and Bio/Technology, 7(2), 173–190.
Edgar, R. C. (2004). MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics, 5(1), 113–113.
Flörke, M., Kynast, E., Bärlund, I., Eisner, S., Wimmer, F., & Alcamo, J. (2013). Domestic and industrial water uses of the past 60 years as a mirror of socio-economic development: a global simulation study. Global Environmental Change, 23(1), 144–156.
Fotidis, I. A., Karakashev, D., & Angelidaki, I. (2013). Bioaugmentation with an acetate-oxidising consortium as a tool to tackle ammonia inhibition of anaerobic digestion. Bioresource Technology, 146, 57–62.
Gabriel, D., & Deshusses, M. A. (2003). Retrofitting existing chemical scrubbers to biotrickling filters for H2S emission control. Proceedings of the National Academy of Sciences, 100(11), 6308–6312.
Hansen, T. A. (1993). Carbon metabolism of sulfate-reducing bacteria. In The sulfate-reducing bacteria: contemporary perspectives (pp. 21–40). New York: Springer.
Henshaw, P. (2001). Biological conversion of hydrogen sulphide to elemental sulphur in a fixed-film continuous flow photo-reactor. Water Research, 35(15), 3605–3610.
Henshaw, P., Medlar, D., & McEwen, J. (1999). Selection of a support medium for a fixed-film green sulphur bacteria reactor. Water Research, 33(14), 3107–3110.
Hobson, P. N. (1982). Biogas production from agricultural wastes. Experientia, 38(2), 206–209.
Hurse, T. J., Kappler, U., & Keller, J. (2008). Using anoxygenic photosynthetic bacteria for the removal of sulfide from wastewater. In Sulfur metabolism in phototrophic organisms (pp. 437–460). Dordrecht: Springer.
Imachi, H., & Sakai, S. (2015). Methanolinea. In Bergey’s manual of systematics of archaea and bacteria (pp. 1–4). Chichester: Wiley.
Imhoff, J. F. (2008). Systematics of anoxygenic phototrophic bacteria. In Sulfur metabolism in phototrophic organisms (pp. 269–287). Dordrecht: Springer.
Imhoff, J. F. (2014a). The family Chlorobiaceae. In The prokaryotes (4 ed., pp. 501–514). Berlin: Springer.
Imhoff, J. F. (2014b). The family Chromatiaceae. In The prokaryotes (pp. 151–178). Berlin: Springer.
Imhoff, J. F. (2015). Chromatiaceae. In Bergey’s manual of systematics of archaea and bacteria (pp. 1–12). Chichester: Wiley.
Kim, B. W., & Chang, H. N. (1991). Removal of hydrogen sulfide by Chlorobium thiosulfatophilum in immobilized-cell and sulfur-settling free-cell recycle reactors. Biotechnology Progress, 7(6), 495–500.
Kimura, M. (1980). A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution, 16(2), 111–120.
Klok, J. B. M., de Graaff, M., van den Bosch, P. L. F., Boelee, N. C., Keesman, K. J., & Janssen, A. J. H. (2013). A physiologically based kinetic model for bacterial sulfide oxidation. Water Research, 47(2), 483–492.
Kobayashi, H. A., Stenstrom, M., & Mah, R. A. (1983). Use of photosynthetic bacteria for hydrogen sulfide removal from anaerobic waste treatment effluent. Water Research, 17(5), 579–587.
Koeck, D. E., Hahnke, S., & Zverlov, V. V. (2016). Herbinix luporum sp. nov., a thermophilic cellulose-degrading bacterium isolated from a thermophilic biogas reactor. International Journal of Systematic and Evolutionary Microbiology, 66(10), 4132–4137.
Koschorreck, M. (2008). Microbial sulphate reduction at a low pH. FEMS Microbiology Ecology, 64(3), 329–342.
Kumar, S., Stecher, G., Li, M., Knyaz, C., Tamura, K., & Battistuzzi, F. U. (2018). MEGA X: molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35(6), 1547–1549.
Kushkevych, I., Vítězová, M., Vítěz, T., & Bartoš, M. (2017). Production of biogas: relationship between methanogenic and sulfate-reducing microorganisms. Open Life Sciences, 12(1), 82–91.
Kushkevych, I., Vítězová, M., Vítěz, T., Kováč, J., Kaucká, P., Jesionek, W., et al. (2018a). A new combination of substrates: biogas production and diversity of the methanogenic microorganisms. Open Life Sciences, 13(1), 119–128.
Kushkevych, I., Kováč, J., Vítězová, M., Vítěz, T., & Bartoš, M. (2018b). The diversity of sulfate-reducing bacteria in the seven bioreactors. Archives of Microbiology, 200(6), 945–950.
Kushkevych, I., Dordević, D., & Vítězová, M. (2019a). Toxicity of hydrogen sulfide toward sulfate-reducing bacteria Desulfovibrio piger Vib-7. Archives of Microbiology, 201(3), 389–397.
Kushkevych, I., Kobzová, E., Vítězová, M., Vítěz, T., Dordević, D., & Bartoš, M. (2019b). Acetogenic microorganisms in operating biogas plants depending on substrate combinations. Biologia, 74, 1–8.
Laanbroek, H. J., Geerligs, H. J., Sijtsma, L., & Veldkamp, H. (1984). Competition for sulfate and ethanol among Desulfobacter, Desulfobulbus, and Desulfovibrio species isolated from intertidal sediments. Applied and Environmental Microbiology, 47(2), 329.
Laws, E. A. (2017). Aquatic pollution: an introductory text (4th ed.). Hoboken: Wiley.
Lin, S., Mackey, H. R., Hao, T., Guo, G., van Loosdrecht, M. C. M., & Chen, G. (2018). Biological sulfur oxidation in wastewater treatment: a review of emerging opportunities. Water Research, 143, 399–415.
Manzoor, S., Schnürer, A., Bongcam-Rudloff, E., & Müller, B. (2016). Complete genome sequence of Methanoculleus bourgensis strain MAB1, the syntrophic partner of mesophilic acetate-oxidising bacteria (SAOB). Standards in Genomic Sciences, 11(1).
McCartney, D. M., & Oleszkiewicz, J. A. (1993). Competition between methanogens and sulfate reducers: effect of COD. Water Environment Research, 65(5), 655–664.
Muyzer, G., & Stams, A. J. M. (2008). The ecology and biotechnology of sulphate-reducing bacteria. Nature Reviews Microbiology, 6(6), 441–454.
Naik, S. N., Goud, V. V., Rout, P. K., & Dalai, A. K. (2010). Production of first and second generation biofuels: a comprehensive review. Renewable and Sustainable Energy Reviews, 14(2), 578–597.
Nath, K., & Das, D. (2004). Improvement of fermentative hydrogen production: various approaches. Applied Microbiology and Biotechnology, 65(5).
Nossa, C. W. (2010). Design of 16S rRNA gene primers for 454 pyrosequencing of the human foregut microbiome. World Journal of Gastroenterology, 16(33).
Oren, A. (2014). The family Methanospirillaceae. In The prokaryotes (pp. 283–290). Berlin: Springer.
Oyarzún, P., Arancibia, F., Canales, C., & Aroca, G. E. (2003). Biofiltration of high concentration of hydrogen sulphide using Thiobacillus thioparus. Process Biochemistry, 39(2), 165–170.
Parawira, W., Read, J. S., Mattiasson, B., & Björnsson, L. (2008). Energy production from agricultural residues: high methane yields in pilot-scale two-stage anaerobic digestion. Biomass and Bioenergy, 32(1), 44–50.
Parkin, G. F., Lynch, N. A., Kuo, W. C., Van Keuren, E. L., & Bhattacharya, S. K. (1990). Interaction between sulfate reducers and methanogens fed acetate and propionate. Research Journal of the Water Pollution Control Federation, 62(6), 780–788.
Patel, G. B., Khan, A. W., Agnew, B. J., & Colvin, J. R. (1980). Isolation and characterization of an anaerobic, cellulolytic microorganism, Acetivibrio cellulolyticus gen. nov., sp. nov. International Journal of Systematic and Evolutionary Microbiology, 30(1), 179–185.
Podosokorskaya, O. A., Bonch-Osmolovskaya, E. A., Beskorovaynyy, A. V., Toshchakov, S. V., Kolganova, T. V., & Kublanov, I. V. (2014). Mobilitalea sibirica gen. nov., sp. nov., a halotolerant polysaccharide-degrading bacterium. International Journal of Systematic and Evolutionary Microbiology, 64(Pt 8), 2657–2661.
Pokorna, D., & Zabranska, J. (2015). Sulfur-oxidizing bacteria in environmental technology. Biotechnology Advances, 33(6), 1246–1259.
Schnurer, A., Schink, B., & Svensson, B. H. (1996). Clostridium ultunense sp. nov., a mesophilic bacterium oxidizing acetate in syntrophic association with a hydrogenotrophic methanogenic bacterium. International Journal of Systematic Bacteriology, 46(4), 1145–1152.
Sercu, B., Núñez, D., Van Langenhove, H., Aroca, G., & Verstraete, W. (2005). Operational and microbiological aspects of a bioaugmented two-stage biotrickling filter removing hydrogen sulfide and dimethyl sulfide. Biotechnology and Bioengineering, 90(2), 259–269.
Stams, A. J. M., & Plugge, C. M. (2009). Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nature Reviews Microbiology, 7(8), 568–577.
Stefanie, J. W. H. O. E., Visser, A., Pol, L. W. H., & Stams, A. J. M. (1994). Sulfate reduction in methanogenic bioreactors. FEMS Microbiology Reviews, 15(2–3), 119–136.
Struk, M., Kushkevych, I. (2018). Perspectives of application of phototrophic sulfur bacteria in hydrogen sulfide utilization. In MendelNet 2018: proceedings of 25th international PhD students conference. Brno, Czech Republic, 537–541.
Sublette, K. L., & Sylvester, N. D. (1987). Oxidation of hydrogen sulfide by mixed cultures of Thiobacillus denitrificans and heterotrophs. Biotechnology and Bioengineering, 29(6), 759–761.
Syed, M. A., & Henshaw, P. F. (2003). Effect of tube size on performance of a fixed-film tubular bioreactor for conversion of hydrogen sulfide to elemental sulfur. Water Research, 37(8), 1932–1938.
Syed, M., Soreanu, G., Falletta, P., & Béland, M. (2006). Removal of hydrogen sulfide from gas streams using biological processes—a review. Canadian Biosystems Engineering, 48, 2.
Tang, K., Baskaran, V., & Nemati, M. (2009). Bacteria of the sulphur cycle: an overview of microbiology, biokinetics and their role in petroleum and mining industries. Biochemical Engineering Journal, 44(1), 73–94.
Tursman, J. F., & Cork, D. J. (1989). Influence of sulfate and sulfate-reducing bacteria on anaerobic digestion technology. Advances in Biotechnological Processes, 12, 273–285.
Ullah Khan, I., Hafiz Dzarfan Othman, M., Hashim, H., Matsuura, T., Ismail, A. F., Rezaei-DashtArzhandi, M., & Wan Azelee, I. (2017). Biogas as a renewable energy fuel—a review of biogas upgrading, utilisation and storage. Energy Conversion and Management, 150, 277–294.
van den Brand, T. P. H., Roest, K., Brdjanovic, D., Chen, G. H., & van Loosdrecht, M. C. M. (2014). Influence of acetate and propionate on sulphate-reducing bacteria activity. Journal of Applied Microbiology, 117(6), 1839–1847.
Ziemiński, K., & Frąc, M. (2012). Methane fermentation process as anaerobic digestion of biomass: transformations, stages and microorganisms. African Journal of Biotechnology, 11(18), 4127–4139.
Funding
This study was supported by a grant agency of Masaryk University (MUNI/A/0902/2018).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Struk, M., Vítězová, M., Vítěz, T. et al. Modřice Plant Anaerobic Digester: Microbial Distribution and Biogas Production. Water Air Soil Pollut 230, 240 (2019). https://doi.org/10.1007/s11270-019-4289-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-019-4289-4