Abstract
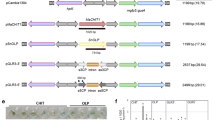
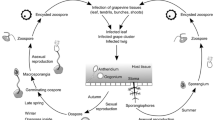
The fungi Botrytis cinerea and Erysiphe necator are responsible for gray mold and powdery mildew diseases, respectively, which are among the most devastating diseases of grapes. Two endochitinase (ech42 and ech33) genes and one N-acetyl-β-d-hexosaminidase (nag70) gene from biocontrol agents related to Trichoderma spp. were used to develop a set of 103 genetically modified (GM) ‘Thompson Seedless’ lines (568 plants) that were established in open field in 2004 and evaluated for fungal tolerance starting in 2006. Statistical analyses were carried out considering transgene, explant origin, and plant response to both fungi in the field and in detached leaf assays. The results allowed for the selection of the 19 consistently most tolerant lines through two consecutive years (2007–2008 and 2008–2009 seasons). Plants from these lines were grafted onto the rootstock Harmony and established in the field in 2009 for further characterization. Transgene status was shown in most of these lines by Southern blot, real-time PCR, ELISA, and immunostrips; the most tolerant candidates expressed the ech42–nag70 double gene construct and the ech33 gene from a local Hypocrea virens isolate. B. cinerea growth assays in Petri dishes supplemented with berry juices extracted from the most tolerant individuals of the selected population was inhibited. These results demonstrate that improved fungal tolerance can be attributed to transgene expression and support the iterative molecular and physiological phenotyping in order to define selected individuals from a population of GM grapevines.





Similar content being viewed by others
References
Bhat S, Srinivasan S (2002) Molecular and genetic analyses of transgenic plants: considerations and approaches. Plant Sci 164:673–681
Bolar J, Norelli J, Wong K, Hayes C, Harman G, Aldwinckle H (2000) Expression of endochitinase from Trichoderma harzianum in transgenic apple increases resistance to apple scab and reduces vigor. Phytopathology 90:72–77
Bolar J, Norelli J, Harman G, Brown S, Aldwinckle H (2001) Synergistic activity of endochitinase and exochitinase from Trichoderma atroviride (T. harzianum) against the pathogenic fungal (Venturia inaequalis) in transgenic apples plants. Transgenic Res 10:533–543
Butaye KJM, Goderis IJWM, Wouters PFJ, Pues JM-TG, Delauré SL, Broekaert WF, Depicker A, Cammue BPA, De Bolle MFC (2004) Stable high-level transgene expression in Arabidopsis thaliana using gene silencing mutants and matrix attachment regions. Plant J 39:440–449
Chye M, Zhao K, He Z, Ramalingam S, Fung K (2005) An agglutinating chitinase with two chitin-binding domains confers fungal protection in transgenic potato. Planta 220:717–730
Dalla Costa L, Vaccari I, Mandolini M, Martinelli L (2009) Elaboration of a reliable strategy based on real-time PCR to characterize genetically modified plantlets and to evaluate the efficiency of a marker gene removal in grape (Vitis spp.). J Agric Food Chem 57:2668–2677
Dana M, Pintor-Toro T, Cubero B (2006) Transgenic tobacco plants overexpressing chitinases of fungal origin show enhanced resistance to biotic and abiotic stress agents. Plant Physiol 142:722–730
De la Cruz J, Hidalgo-Gallego A, Lora J, Benítez T, Pintor-Toro T, Llobell A (1992) Isolation and characterization of three chitinases from Trichoderma harzianum. Eur J Biochem 206:859–867
Dhekney S, Li Z, Gray DJ (2011) Grapevines engineered to express cisgenic Vitis vinifera thaumatin-like protein exhibit fungal disease resistance. In Vitro Cell Dev Biol Plant 47:458–466
Di Genova A, Miyasaka A, Muñoz-Espinoza C, Vizoso P, Travisany D, Moraga C, Pinto M, Hinrichsen P, Orellana A, Maass A (2014) Whole genome comparison between table and wine grapes reveals a comprehensive catalog of structural variants. BMC Plant Biol 14:7. doi:10.1186/1471-2229-14-7
Distefano G, La Malfa S, Vitale A, Lorito M, Deng Z, Gentile A (2008) Defence-related gene expression in transgenic lemon plants producing an antimicrobial Trichoderma harzianum endochitinase during fungal infection. Transgenic Res 17:873–879
Domínguez A, Cervera M, Pérez R, Romero J, Fagoaga C, Cubero J, López M, Juárez J, Navarro L, Peña L (2004) Characterisation of regenerants obtained under selective conditions after Agrobacterium-mediated transformation of citrus explants reveals production of silenced and chimeric plants at unexpected high frequencies. Mol Breed 14:171–183
Driver JA, Kuniyuki AH (1984) In vitro propagation of paradox walnut rootstock. Hort Sci 19:507–509
Dutt M, Dhekney A, Gray D (2007) Transgenic plants from shoot apical meristems of Vitis vinifera L. ‘‘Thompson Seedless’’ via Agrobacterium-mediated transformation. Plant Cell Rep 26:2101–2110
Elad Y, Williamson B, Tudzynski P, Delen N (2004) Botrytis: biology, pathology and control. Kluwer, Dodrecht
Emani C, García J, Lopata-Finch E, Pozo M, Uribe P, Kim D, Sunilkumar G, Cook D, Kenerly C, Rathore K (2003) Enhanced fungal resistance in transgenic cotton expressing an endochitinase gene from Trichicoderma virens. Plant Biotechnol J 1:321–326
Esposito S, Colucci M, Frusciante L, Filippone E, Lorito M, Bresan R (2000) Antifungal transgenes expression in Petunia hybrida. Acta Hortic 508:157–161
Faize M, Malnoy M, Dupuis F, Chevalier M, Parisi L, Chevreau E (2003) Chitinases of Trichoderma atroviride induce scab resistance and some metabolic changes in two cultivars of apple. Phytopathology 12:1496–1504. doi:10.1094/PHYTO.2003.93.12.1496
Faize M, Faize L, Burgos L (2010) Using quantitative real-time PCR to detect chimeras in transgenic tobacco and apricot and to monitor their dissociation. BMC Biotechnol 10:53–60
Figueiredo A, Fortes A, Ferreira S, Sebastiana M, Sousa L, Acioli-Santos B, Pessoa F, Verpoorte R, Hae Choi Y, Pais M (2008) Transcriptional and metabolic profiling of grape (Vitis vinifera L.) leaves unravel possible innate resistance against pathogenic fungi. J Exp Bot 59:3371–3381
Flachowsky H, Riedel M, Reim S, Hanke V (2008) Evaluation of the uniformity and stability of T-DNA integration and gene expression in transgenic apple plants. Electron J Biotechnol 11:1–15
Gentile A, Deng Z, La Malfa S, Distefano G, Domina F, Vitale A, Polizzi G, Lorito M, Tribulato E (2007) Enhanced resistance to Phoma tracheiphila and Botrytis cinerea in transgenic lemon plants expressing a Trichoderma harzianum chitinase gene. Plant Breed 126:146–151
Gray DJ, Li Z, Dhekney S (2014) Precision breeding of grapevine (Vitis vinifera L.) for improved traits. Plant Sci. doi:10.1016/j.plantsci.2014.03.023
Haggag W (2008) Biotechnological aspects of plant resistant for fungal diseases management. American-Eurasian J Sustain Agric 2:1–18
Hayes CK, Klemsdal S, Lorito M, Di Pietro A, Peterbauer C, Nakas JP, Tronsmo A, Harman G (1994) Isolation and sequence of an endochitinase-encoding gene from a cDNA library of Trichoderma harzianum. Gene 138:143–148
Hily J-M, Scorza R, Webb K, Ravelonandro M (2005) Accumulation of the long class of siRNA is associated with resistance to Plum pox virus in a transgenic woody perennial plum tree. Mol Plant Microbe Interact 18:794–799. doi:10.1094/MPMI-18-0794
Hinrichsen P, Reyes MA, Castro A, Araya S, Garnier M, Prieto H, Reyes F, Muñoz C, Dell’Orto P, Moynihan MR (2005) Genetic transformation of grapevines with Trichoderma harzianum and antimicrobial peptide genes for improvement of fungal tolerance. Acta Hortic 689:469–474
Höenicka H, Fladung M (2006) Biosafety in Populus spp. and other forest trees: from non-native speciesto taxa derived from traditional breeding and genetic engineering. Trees Struct Funct 20:131–144
IPGRI, UPOV, OIV (1997) Descriptors for grapevine (Vitis spp.). International Union for the Protection of New Varieties of Plants, Geneva/Office International de la Vigne et du Vin, Paris/International Plant Genetic Resources Institute, Rome
Jaillon O, Aury J, Noel B, Policriti A, Clepet C, Casagrande A, Choisne N, Aubourg S, Vitulo N, Jubin C, Vezzi A, Legeai F, Hugueney P, Dasilva C, Horner D, Mica E, Jublot D, Poulain J, Bruyère C, Billault A, Segurens B, Gouyvenoux M, Ugarte E, Cattonaro F, Anthouard V, Vico V, Del Fabbro C, Alaux M, Di Gaspero G, Dumas V, Felice N, Paillard S, Juman I, Moroldo M, Scalabrin S, Canaguier A, Le Clainche I, Malacrida G, Durand E, Pesole G, Laucou V, Chatelet P, Merdinoglu D, Delledonne M, Pezzotti M, Lecharny A, Scarpelli C, Artiguenave F, Pè ME, Valle G, Morgante M, Caboche M, Adam-Blondon AF, Weissenbach J, Quétier F, Wincker P (2007) French-Italian public consortium for grapevine genome characterization: the grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–467
Keller M, Viret O, Cole F (2003) Botrytis cinerea infection in grape flowers: defense reaction, latency, and disease expression. Phytopathology 93:316–322
Kikkert JR, Ali GS, Wallace PG, Reisch B, Reustle GM (2000) Expression of a fungal chitinase in Vitis vinifera L. ‘Merlot’ and ‘Chardonnay’ plants produced by biolistic transformation. Acta Hortic 528:297–303
Li Z, Jayasankar S, Gray D (2001) Expression of a bifunctional green fluorescent protein (GFP) fusion marker under the control of three constitutive promoters and enhanced derivatives in transgenic grapes (Vitis vinifera). Plant Sci 160:877–887
Limón M, Lora J, García I, de la Cruz J, Llobell A, Benítez T, Pintor-Toro J (1995) Primary structure and expression pattern of the 33-kDa chitinase gene from the mycoparasitic fungus Trichoderma harzianum. Curr Genet 28:478–483
Liu M, Sun Z, Zhu J, Xu T, Harman G, Lorito M (2004) Enhancing rice resistance to pathogens by transformation with cell wall degrading enzyme genes from Trichoderma atroviride. J Zhejiang Univ Sci 5:133–136
Lorito M, Woo S, García-Fernandez I, Colicci G, Harman G, Pintor-Toro J, Filippone E, Muccifora S, Lawrence C, Zoina A, Tuzun S, Scala F (1998) Genes from mycoparasitic fungi as a source for improving plant resistance to fungal pathogens. Proc Natl Acad Sci USA 95:7860–7865
Martinelli L, Gribaudo I (2009) Strategies for effective somatic embryogenesis in grapevine: an appraisal. In: Roubelakis-Angelakis K (ed) Grapevine molecular physiology & biotechnology, 2nd edn. Springer, Netherlands, pp 461–493
Ming R, Hou S, Feng Y, Yu Q, Dionne-Laporte A, Saw J, Senin P, Wang W, Ly B, Lewis K, Salzberg S, Feng L, Jones M, Skelton R, Murray J, Chen C, Qian W, Shen J, Du P, Eustice M, Tong E, Tang H, Lyons E, Paull R, Michael T, Wall K, Rice D, Albert H, Wang M, Zhu Y, Schatz M, Nagarajan N, Acob R, Guan P, Blas A, Wai C, Ackerman C, Ren Y, Liu C, Wang J, Wang J, Na J, Shakirov E, Haas B, Thimmapuram J, Nelson D, Wang X, Bowers J, Gschwend A, Delcher A, Singh R, Suzuki J, Tripathi S, Neupane K, Wei K, Irikura B, Paidi M, Jiang N, Zhang W, Presting G, Windsor A, Navajas-Pérez R, Torres M, Feltus A, Porter B, Li Y, Burroughs M, Luo M, Liu L, Christopher D, Mount S, Moore P, Sugimura T, Jiang J, Schuler M, Friedman M, Mitchell-Olds T, Shippen D, dePamphilis C, Palmer J, Freeling M, Paterson A, Gonsalves D, Wang L, Alam M (2008) The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus). Nature 452:991–996. doi:10.1038/nature06856
Mora-Avilés A, Earle E (2004) Expression of pathogenesis-related genes in transgenic broccoplants expressing the Trichoderma harzianum-endochitinase gene. Rev Chapingo Ser Hortic 10:141–146
Müller F, Werner K, Kasai M, Francesconi A, Chanock S, Walsh T (1998) Rapid extraction of genomic DNA from medically important yeasts and filamentous fungi by high-speed cell disruption. J Clin Microbiol 36:1625–1629
Muñoz G, Hinrichsen P, Brygoo Y, Giraud T (2002) Genetic characterisation of Botryticinerea populations in Chile. Mycol Res 106:594–601
Peterlunger E, Di Gaspero G, Cipriani G, Sivilotti P, Zulini L, Marrazzo MT, Andreetta D, Testolin R (2003) Breeding strategy for the introgression of disease resistance genes into European grapevine. Acta Hortic 603:665–670
Pons E, Peris J, Peña L (2012) Field performance of transgenic citrus trees: assessment of the long-term expression of uidA and nptII transgenes and its impact on relevant agronomic and phenotypic characteristics. BMC Biotechnol 12:41. doi:10.1186/1472-6750-12-41
Quiroz-Figueroa F, Rojas-Herrera R, Galaz-Avalos R, Loyola-Vargas V (2006) Embryo production through somatic embryogenesis can be used to study cell differentiation in plants. Plant Cell Tissue Organ Cult 86:285–301
Ramming D, Gabler F, Smilanick J, Cadle-Davidson M, Barba P, Mahanil S, Cadle-Davidson L (2011) A single dominant locus, Ren4, confers rapid non-race-specific resistance to grapevine powdery mildew. Phytopahology 101:502–508
Reyes F, Reyes MA, Castro A, Araya S, Dell’Orto P, Moynihan MR, Muñoz C, Prieto H, Hinrichsen P (2005) A mid-scale platform for genetic transformation of different grapevine varieties: use of ‘Thompson Seedless’ as a model. In: International symposium on biotechnology of temperate fruit crops and tropical species, October 10–14, Daytona Beach, FL, USA
Rigotti S, Gindro K, Richter H, Viret O (2002) Characterization of molecular markers for specific and sensitive detection of Botrytis cinerea Pers.: Fr. in strawberry (Fragaria × ananassa Duch.) using PCR. FEMS Microbiol Lett 209:169–174
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Schuster A, Schmoll M (2010) Biology and biotechnology of Trichoderma. Appl Microbiol Biotechnol 87:787–799
Sharma P, Saravanan K, Ramesh R, Kumar V, Singh D, Sharma M, Henry M, Deep S (2012) Cloning and semi-quantitative expression of endochitinase (ech42) gene from Trichoderma spp. Afr J Biotechnol 11:12930–12938
Steenkamp J, Wiid I, Lourens A, van Helden P (1994) Improved method for DNA extraction from Vitis vinifera. Am J Enol Vitic 45:102–106
Tapia E, Sequeida A, Castro A, Montes C, Zamora P, Prieto H (2009) Development of grapevine somatic embryogenesis using an air-lift bioreactor as an efficient tool in the generation of transgenic plants. J Biotechnol 139:95–101
Urtubia C, Devia J, Castro A, Zamora P, Aguirre C, Tapia E, Barba P, Dell’Orto P, Moynihan MR, Petri C, Scorza R, Prieto H (2008) Agrobacterium-mediated genetic transformation of Prunus salicina. Plant Cell Rep 27:1333–1340
Vain P, James A, Worland B, Snape W (2002) Transgene behaviour across two generations in a large random population of transgenic rice plants produced by particle bombardment. Theor Appl Genet 105:878–889
Velasco R, Zharkikh A, Troggio M, Cartwright D, Cestaro A, Pruss D, Pindo M, Fitzgerald L, Vezzulli S, Reid J, Malacarne G, Iliev D, Coppola G, Wardell B, Micheletti D, Macalma T, Facci M, Mitchell J, Perazzolli M, Eldredge G, Gatto P, Oyzerski R, Moretto M, Gutin N, Stefanini M, Chen Y, Segala C, Davenport C, Demattè L, Mraz A, Battilana J, Stormo K, Costa F, Tao Q, Si-Ammour A, Harkins T, Lackey A, Perbost C, Taillon B, Stella A, Solovyev V, Fawcett JA, Sterck L, Vandepoele K, Grando SM, Toppo S, Moser C, Lanchbury J, Bogden R, Skolnick M, Sgaramella V, Bhatnagar SK, Fontana P, Gutin A, Van de Peer Y, Salamini F, Viola R (2007) A high quality draft consensus sequence of the genome of a heterozygous grapevine variety. PLoS One 2:e1326. doi:10.1371/journal.pone.0001326
Vellicce G, Díaz-Ricci J, Hernández L, Castagnaro A (2006) Enhanced resistance to Botrytis cinerea mediated by the transgenic expression of chitinase gene ch5B in strawberry. Transgenic Res 15:57–68
Wally O, Punja Z (2010) Genetic engineering for increasing fungal and bacterial disease resistance in crop plants. GM Crops 1(4):199–206
Acknowledgments
This work was funded by the BIOFRUTALES Consortium and the grant INNOVA CHILE 09PMG-7229. Authors are grateful to Carlos Muñoz, Patricio Hinrichsen, Paola Dell’Orto and Mike R. Moynihan by their participation in the founding works developing the GM ‘Thompson Seedless’ lines funded by the FONDEF CHILE D01I1064 grant.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rubio, J., Montes, C., Castro, Á. et al. Genetically engineered Thompson Seedless grapevine plants designed for fungal tolerance: selection and characterization of the best performing individuals in a field trial. Transgenic Res 24, 43–60 (2015). https://doi.org/10.1007/s11248-014-9811-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-014-9811-2