Abstract
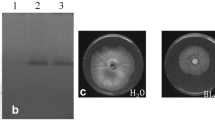
Plant chitinases are strong antifungal enzymes, and their genes can be utilized to engineer crop plants, including potatoes, with tolerance against fungal pathogens. The purified recombinant chitinase protein significantly inhibited the growth of phytopathogenic fungi, including Alternaria solani, in a qualitative invitro antifungal assay. Furthermore, transgenic potatoes (variety Desiree) over-expressing the endo-chitinase gene were successfully developed using the Agrobacterium-mediated transformation method. The transformation efficiency was 21% based on preliminary molecular and biochemical analyses, including histochemical GUS staining, PCR, and Southern blotting. Crude protein extracts from transgenic potato plants inhibited the hyphal growth of A. solani from 39.5 to 60.5% in a quantitative in vitro assay. The in-vitro bioassay showed that transgenic potato plants exhibit strong resistance against A. solani infection as these plants remains green and healthy, while non-transgenic control plants turned yellow and eventually died 3-week post infection. In the detached leaf assay, the leaves of transgenic plants challenged with A. Soltani showed a decrease in susceptibility compared to the control. The number of necrotic spots in the control leaves was significantly higher compared to the number in the transgenic leaves of potato plants infected with A. solani. Furthermore, A. solani infection triggered a gradual increase in mRNA expression of the transgene in transgenic plants compared to non-transgenic plants as revealed in the qRT-PCR assay. mRNA expression increased to 7.34-fold higher at 72-h post infection compared to the control, and mRNA expression was directly correlated with the abundance of specific transgene transcripts.










Similar content being viewed by others
Abbreviations
- SDS-PAGE:
-
Sodium dodecyl suplhate; poly acrylamide gel electrophoresis
- kDA:
-
Kilo Daltons
- bp:
-
Base pair
- GUS:
-
β-Glucuronidase
- qPCR:
-
Quantitative polymerase chain reaction
- mRNA:
-
Messenger RNA
References
Barrett CB (2010) Measuring food insecurity. Science 327(5967):825–828
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72(1–2):248–254
Chaerani R, Voorrips RE (2006) Tomato early blight (Alternaria solani): the pathogen, genetics, and breeding for resistance. J Gen Plant Pathol 72(6):335–347
Chhikara S, Chaudhury D, Dhankher OP, Jaiwal PK (2012) Combined expression of a barley class II chitinase and type I ribosome inactivating protein in transgenic Brassica juncea provides protection against Alternaria brassicae. Plant Cell Tissue Organ Cult (PCTOC) 108(1):83–89
Chopra R, Saini R (2014) Transformation of blackgram (Vigna mungo (L.) Hepper) by barley chitinase and ribosome-inactivating protein genes towards improving resistance to Corynespora leaf spot fungal disease. Appl Biochem Biotechnol 174(8):2791–2800
Collinge DB, Lund OS, Thordal-Christensen H (2008) What are the prospects for genetically engineered, disease resistant plants? Eur J Plant Pathol 121(3):217–231
De Alba AEM, Elvira-Matelot E, Vaucheret H (2013) Gene silencing in plants: a diversity of pathways. Biochim Biophys Acta (BBA) Gene Regul Mech 1829(12):1300–1308
de Cáceres González FFN, Davey MR, Sanchez EC, Wilson ZA (2015) Conferred resistance to Botrytis cinerea in Lilium by overexpression of the RCH10 chitinase gene. Plant Cell Rep 34(7):1201–1209
de la Vega LM, Barboza-Corona JE, Aguilar-Uscanga MG, Ramírez-Lepe M (2006) Purification and characterization of an exochitinase from Bacillus thuringiensis subsp. aizawai and its action against phytopathogenic fungi. Can J Microbiol 52(7):651–657
Ebadzad G, Cravador A (2014) Quantitative RT-PCR analysis of differentially expressed genes in Quercus suber in response to Phytophthora cinnamomi infection. SpringerPlus 3(1):1–13
Elhamid MA, Makboul H, Sedik M, Ismail I, Ibrahim M (2010) Cloning, expression and antifungal activity of an endochitinase gene derived from barley (Hordeum vulgare). Res J Agric Biol Sci 6(3):356–363
Ghosh S, Molla KA, Karmakar S, Datta SK, Datta K (2016) Enhanced resistance to late blight pathogen conferred by expression of rice oxalate oxidase 4 gene in transgenic potato. Plant Cell Tissue Organ Cult (PCTOC) 126(3):429–437 doi:10.1007/s11240-016-1011-8
Girhepuje P, Shinde G (2011) Transgenic tomato plants expressing a wheat endochitinase gene demonstrate enhanced resistance to Fusarium oxysporum f. sp. lycopersici. Plant Cell Tissue Organ Cult (PCTOC) 105(2):243–251
Iqbal MM, Nazir F, Ali S, Asif MA, Zafar Y, Iqbal J, Ali GM (2012) Over expression of rice chitinase gene in transgenic peanut (Arachis hypogaea L.) improves resistance against leaf spot. Mol Biotechnol 50(2):129–136
Islam R, Datta B (2015) Diversity of chitinases and their industrial potential. Int J Appl Res 1:55–60
Jabeen N, Chaudhary Z, Gulfraz M, Rashid H, Mirza B (2015) Expression of rice chitinase gene in genetically engineered tomato confers enhanced resistance to Fusarium wilt and early blight. Plant Pathol J 31(3):252
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6(13):3901
Jitonnom J, Lee VS, Nimmanpipug P, Rowlands HA, Mulholland AJ (2011) Quantum mechanics/molecular mechanics modeling of substrate-assisted catalysis in family 18 chitinases: conformational changes and the role of Asp142 in catalysis in ChiB. BioChemistry 50(21):4697–4711
Kalantidis K, Psaradakis S, Tabler M, Tsagris M (2002) The occurrence of CMV-specific short RNAs in transgenic tobacco expressing virus-derived double-stranded RNA is indicative of resistance to the virus. Mol Plant Microbe Interact 15(8):826–833
Kirubakaran SI, Sakthivel N (2007) Cloning and overexpression of antifungal barley chitinase gene in Escherichia coli. Protein Expr Purif 52(1):159–166
Kovács G, Sági L, Jacon G, Arinaitwe G, Busogoro J-P, Thiry E, Strosse H, Swennen R, Remy S (2013) Expression of a rice chitinase gene in transgenic banana (‘Gros Michel’, AAA genome group) confers resistance to black leaf streak disease. Transgenic Res 22(1):117–130
Lawrence CB, Singh NP, Qiu J, Gardner RG, Tuzun S (2000) Constitutive hydrolytic enzymes are associated with polygenic resistance of tomato to Alternaria solani and may function as an elicitor release mechanism. Physiol Mol Plant Pathol 57(5):211–220
Mondal KK, Bhattacharya R, Koundal K, Chatterjee S (2007) Transgenic Indian mustard (Brassica juncea) expressing tomato glucanase leads to arrested growth of Alternaria brassicae. Plant Cell Rep 26(2):247–252
Narayanasamy P (2013) Mechanisms of action of fungal biological control agents biological management of diseases of crops. Springer, New York, pp 99–200
Nash A, Gardner R (1988) Tomato early blight resistance in a breeding line derived from Lycopersicon hirsutum PI 126445. Plant Dis 72(3):206–209
Naz FARAH, Rauf CA, Abbasi NA, Haque I, Ahmad I (2008) Influence of inoculum levels of Rhizoctonia solani and susceptibility on new potato germplasm. Pak J Bot 40(5):2199–2209
Nonhebel S (2012) Global food supply and the impacts of increased use of biofuels. Energy 37(1):115–121
Patil RS, Ghormade V, Deshpande MV (2000) Chitinolytic enzymes: an exploration. Enzym Microb Technol 26(7):473–483
Prasad K, Bhatnagar-Mathur P, Waliyar F, Sharma KK (2013) Overexpression of a chitinase gene in transgenic peanut confers enhanced resistance to major soil borne and foliar fungal pathogens. J Plant Biochem Biotechnol 22(2):222–233
Sharma N, Sharma K, Gaur R, Gupta V (2011) Role of chitinase in plant defense. Asian J Biochem 6:29–37
Sherf AF, MacNab AA (1986) Vegetable diseases and their control. Wiley, New York
Singh A, Kirubakaran SI, Sakthivel N (2007) Heterologous expression of new antifungal chitinase from wheat. Protein Expr Purif 56(1):100–109
Solgi T, Moradyar M, Zamani MR, Motallebi M (2015) Transformation of canola by chit33 gene towards improving resistance to Sclerotinia sclerotiorum. Plant Prot Sci 51(1):6–12
Su Y, Xu L, Fu Z, Yang Y, Guo J, Wang S, Que Y (2014) ScChi, encoding an acidic class III chitinase of sugarcane, confers positive responses to biotic and abiotic stresses in sugarcane. Int J Mol Sci 15(2):2738–2760
Tang J, Scarth R, Fristensky B (2003) Effects of genomic position and copy number of Acyl-ACP thioesterase transgenes on the level of the target fatty acids in Brassica napus L. Mol Breed 12(1):71–81
Wang Y, Xiong G, Hu J, Jiang L, Yu H, Xu J, Fang Y, Zeng L, Xu E, Xu J (2015) Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat Genet 47(8):944–948
Xayphakatsa K, Tsukiyama T, Inouye K, Okumoto Y, Nakazaki T, Tanisaka T (2008) Gene cloning, expression, purification and characterization of rice (Oryza sativa L.) class II chitinase CHT11. Enzym Microb Technol 43(1):19–24
Author contributions
AK: All the research work/experiments were carried out by this author. IN: The study was planned and designed by this author. He is the advisor of Mr. Anwar Khan. BT: This author helped in the experiments including realtime assays and cloning, expression studies. Also research article is being edited by this author. KA: SDS–PAGE has been performed by this author. MT: This author carried out protein purification and chitinase activity assay. AQR: The author helped with experiments.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Khan, A., Nasir, I.A., Tabassum, B. et al. Expression studies of chitinase gene in transgenic potato against Alternaria solani . Plant Cell Tiss Organ Cult 128, 563–576 (2017). https://doi.org/10.1007/s11240-016-1134-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-016-1134-y