Abstract
Polyethylene terephthalate (PET) is one of the most marketed aromatic polyesters in the world with an annual demand in 2022 of approximately 29 million metric tons, expected to increase by 40% by 2030. The escalating volume of PET waste and the current inadequacy of recycling methods have led to an accumulation of PET in the terrestrial ecosystem, thereby posing significant global health risks. The pressing global energy and environmental issues associated with PET underscore the urgent need for “upcycling” technologies. These technologies aim to transform reclaimed PET into higher-value products, addressing both energy concerns and environmental sustainability. Enzyme-mediated biocatalytic depolymerization has emerged as a potentially bio-sustainable method for treating and recycling plastics. Numerous plastic-degrading enzymes have been identified from microbial origins, and advancements in protein engineering have been employed to modify and enhance these enzymes. Microbial metabolic engineering allows for the development of modified microbial chassis capable of degrading PET substrates and converting their derived monomers into industrial relevant products. In this review, we describe several engineering approaches aiming at enhancing the performances of PET-degrading enzymes and we present the current metabolic engineering strategies adopted to bio-upcycle PET into high-value molecules.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Plastics, synthetic polymers once hailed as a revolutionary material with limitless applications, have undeniably become a ubiquitous part of our modern lives. Their durability, versatility, and cost-effectiveness have led to their pervasive use across various industries and in everyday consumer products. However, the exponential growth in plastic production and consumption has given rise to a pressing global challenge—plastic waste and its subsequent disposal (Van Geem 2023). Over the last 50 years, plastic production has grown by 680%, with an estimated production of approximately 390 million metric tons (Mt) (OECD 2022). The production of plastic, accompanied by improper disposal, has contributed to an accumulation of plastic debris, such as microplastics, in aquatic and soil environments, posing a real threat to natural ecosystems and human well-being (Boots et al. 2019; Koelmans et al. 2019; Redondo-Hasselerharm et al. 2020; Seeley et al. 2020). One of the most common petroleum-derived plastics for packaging materials is the semi-aromatic plastic poly (ethylene terephthalate) (PET). PET is a heteroatomic synthetic polymer composed of terephthalic acid (TPA) and ethylene glycol (EG) linked by ester bonds (Sinha et al. 2010). Most PET plastic is landfilled or incinerated, processes that do not create economic value and contribute to releasing CO2 into the atmosphere, posing an additional environmental issue (Lazarevic et al. 2010; Antelava et al. 2019; Vollmer et al. 2020). Mismanagement of PET waste exacerbates the problem of plastic pollution, necessitating the exploration of economically viable solutions for its end-of-life management and embracing circular economy principles (Meys et al. 2020). One such solution is the recycling of post-consumer PET, aiming to transform it back into its original material form (closed-loop recycling) (Rahimi and García, 2017). Currently, only 9% of plastic materials, including PET, are mechanically recycled (Jehanno et al. 2022). However, this process often leads to downcycled material due to the poor mechanical and structural quality of the resulting recycled materials (Jehanno et al. 2022).
PET can undergo recycling processes to be re-polymerized into high-quality PET, provided that its monomers are available at adequate (very high) purity. This can be achieved using several thermal processes, including thermal oxidation and catalytic pyrolysis, as well as chemical processes, such as alkaline and acidic hydrolysis, glycolysis, and alcoholysis (Rahimi and García, 2017; Kim et al. 2021; Gluth et al. 2022). These methodologies may pose environmental issues due to the use of toxic chemicals and potential cost issues, as a result of the elevated energy-demand necessary for completing the processes, rendering them economically less profitable (Gluth et al. 2022; Jehanno et al. 2022). The use of biotechnological microbial agents for achieving PET depolymerization would circumvent toxicity issues and potentially increase the economic biodegradation feasibility (Benavides Fernández et al. 2022). The recent discovery of a soil bacterium, Ideonella sakaiensis, capable of secreting PET depolymerizing enzymes (Fig. 1a, b) that can cleave the ester bonds of PET, has truly revolutionized the field, opening up new possibilities for the development of economically viable bio-based recycling processes for PET (Yoshida et al. 2016). Advances in bioinformatics and omics approaches, such as multi-omics-based systems biology, metagenomics, and metabolic pathway reconstruction techniques, have enabled researchers to discover new classes of PET-degrading enzymes from different microbial ecosystems (plastisphere) and delineate the biochemical paths underpinning the metabolism and bioconversion of PET monomers in microbes (Erickson et al. 2022; Zhu et al. 2022a; Ji et al. 2023a, b). Despite the potentially great advantages of enzymatic PET biodegradation, several drawbacks regarding the catalytic efficiency and thermal stability of native PET hydrolases remain. These limitations have resulted in PET hydrolases having an insufficient turnover rate for current industrial applications (Wei et al. 2022; Xu et al. 2023). However, the elucidation of crystal structures and catalytic mechanisms of several classes of PET hydrolases facilitated the exploration of structural modifications through different protein engineering designs to increase enzyme performances. Advanced synthetic biology tools and optimization of microbial cell factory designs have promoted the rapid development of various models and non-model microbial systems for decomposing and assimilating carbon from PET substrates (Blank et al. 2020). These systems employ synthetic microbe-based biological reactions or integrated biological and chemical reactions to create an open-loop recycling system for producing, from PET depolymerization intermediates, TPA and EG, higher-value compounds (upcycling) (Lee et al. 2023; Thakur et al. 2023).
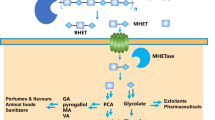
Schematic representation of the enzymatic degradation of PET, including a timeline highlighting key landmarks of identified hydrolytic enzymes and the structure of selected model enzymes. a: enzymatic degradation of PET into the monomers BHET, MHET, TPA and EG. b: timeline showing a selection of important PET hydrolases discovered. Cutinase, esterase, PETase, MHETase, and lipase enzymes are highlighted in purple, blue, aquamarine, orange, and yellow, respectively. Key enzymes include: CcC: C. cladosporioides cutinase, TfF: T. fusca cutinase, HiC: H. insolens cutinase, ThcCut1/ThcCut2: T. cellulosilytica cutinases, Cut190: S. viridis cutinase, LCC: leaf-branch-compost cutinase, PHL7: thermophilic cutinase-like enzyme; ThfCa: T. fusca carboxylesterase, BsEstB: B. subtilis nitrobenzylesterase, ChryBHETase: carboxylesterase from Chryseobacterium sp. PET-29, IsPETase: I. sakaiensis PETase, RpPETase: membrane anchoring PETase-like enzyme of R. pyridinivorans, CaLb: C. antarctica lipase B. c: cartoon representation of the side view of the LCC cutinase (PDB ID: 7VVE) colored in magenta and bound to BHET (colored in black and red). The amino acid residues forming disulfide bonds are enclosed in an oval shape, while the amino acids composing the catalytic triad are displayed in the panel. d: cartoon representation of the side view of the IsPETase (PDB ID: 61LW) colored in aquamarine and bound to BHET (colored in black and red). The extended loop of the enzyme is highlighted in blue. The panel shows the BHET substrate in the active pocket of IsPETase surrounded by the amino acids composing the catalytic triad. e: cartoon representation of the top view of the IsMHETase (PDB ID: 6QGA) colored in orange and bound to the MHET analogue MHETA (grey and red). The panel shows the catalytic triad of IsMHETase (S225, H528, D492) and the amino acids composing the lid domain (colored in density blue). f: cartoon representation of the top view of the ThfCa (PDB ID: 7W1J) colored in blue and bound to MHET (colored in red and green). The panel displays the amino acids composition of the catalytic triad (S185, E319, and H415) and the G-X-S-X-G motif (G184, E185, S186, A187, G188). g: cartoon representation of the top view of the CaLb (PDB ID: 1TCA) colored in yellow and bound to MHET (colored in red and green). The panel shows the amino acids composing the catalytic triad and those responsible for pH-based substrate recognition (colored in deep purple). Protein structures were drawn with the Pymol software (DeLano 2002). (Color figure online)
In this review, we will first overview several classes of PET-degrading enzymes with an eye on their mechanistic activity on PET. Secondly, we will focus on the engineering tools implemented for incrementing PET-degrading enzymes’ catalytic performance along with several studies concerning their microbial production and secretion. Finally, we will take a closer look at the new metabolic engineering approaches employed thus far to rewire microbial metabolism toward the generation of microbial-based PET upcycling systems.
2 Enzymatic degradation of PET
Enzymatic degradation of PET involves the hydrolysis of ester bonds to generate its hydrolysis products: BHET (bis(2-hydroxyethyl) terephthalate), MHET (mono-(2-hydroxyethyl) terephthalate), TPA and EG (Fig. 1a) (Mohanan et al. 2020).
Several classes of hydrolases (EC 3.1.1), specifically, cutinases (EC 3.1.1.74), poly (ethylene terephthalate) hydrolase (EC 3.1.1.101), mono (ethylene terephthalate) hydrolase (EC 3.1.1.102), carboxylesterases (EC 3.1.1.1), and lipases (EC 3.1.1.3), have previously been identified as possessing PET and/or PET degradation intermediates' hydrolytic capabilities (Fig. 1b) (Mohanan et al. 2020). Structurally, all PET hydrolases can be classified into two types: type-I, mostly from fungal origins (lacking the additional disulfide bond (Fig. 1b) and the typical G-X-S-XX-G extended loop), and type-II enzymes, mainly from bacterial sources, characterized by the presence of an additional disulfide bond (Fig. 1c) and an extended loop (Fig. 1d) (Han et al. 2017; Joo et al. 2018). The presence of these additional structures is suggested to confer type-II enzymes better biodegradation capabilities (Wei et al. 2022).
2.1 PET-degrading enzymes
Cutinases are a class of hydrolases belonging to the α/β superfamily. They naturally catalyze the hydrolysis of cutin, an insoluble polyester polymer that forms the plant cuticle through interlinked ester bonds of omega-hydroxy acids (Liang and Zou 2023). Thus, they are prevalently found in saprophytes and phytopathogenic microorganisms, such as fungi and bacteria, but also in green plants. Fungal cutinases have a smaller molecular weight (ranging from 22 to 26 kDa for fungi and 30 to 40 kDa for bacteria), possess two disulfide bonds instead of one, undergo several post-translational modifications, have an optimum temperature ranging between 30 and 40 °C (40–75 °C for bacterial cutinases), and an optimal pH between 7.5 and 10 (7–8 for bacterial cutinases) (Liang and Zou 2023). They share a catalytic site composed of the amino acid triad consisting of serine, histidine, and aspartate (Fig. 1c) and lack the typical hydrophobic lid present in most lipases and esterases, which covers the active site and serves for interfacial substrate activation (Khairul Anuar et al. 2022). The exposed active site makes them more available to accept a broad range of hydrophobic substrates, including PET, and therefore they are powerful catalysts to be exploited in biotechnology.
The first reported cutinase active on PET substrates dates back to 1983 (Fig. 1b) and was found in the darkly pigmented fungal plant-saprotroph Cladosporium cladosporioides (CcC) (Fig. 1b) (Kleeberg et al. 1998). However, the breakthrough achievement was the characterization of a cutinase from the thermophilic actinomycete Thermobifida fusca (TfH) (Fig. 1b), whose depolymerization activity caused the loss of 50% weight of PET bottles with 10% crystallinity (Müller et al. 2005). Follow-up studies showed that the degradation activity of the industrially commercialized HiC (Fig. 1b), from the aerobic cellulolytic fungus Humicola insolens, over PET films is not inhibited by PET degradation intermediates, and the enzyme performed well under immobilized conditions (Ronkvist et al. 2009). Structure-activity studies of the two Thermobifida cellulosilytica cutinases (ThcCut1 and ThcCut2) (Fig. 1b) served for elucidating several structural constraints necessary for interacting with the hydrophobic PET surfaces (Herrero Acero et al. 2011). Later studies on the structure of Cut190 from Saccharomonospora viridis (Fig. 1b) were fundamental for shedding light on the role of Ca2+ in controlling the opening and closing mechanisms of the active site for substrate binding (Oda et al. 2018). This mechanism was later exploited for tailoring enzymes with improved midpoints of thermal denaturation (Tm) and optimal temperatures (Topt) for PET degradation (Wei et al. 2022). The same year, a metagenomics-based approach identified a leaf-branch-compost cutinase (LCC) (Fig. 1b, c) highly active on PET substrates, which has then become one important PET-hydrolase model for enzyme engineering studies (see below) (Sulaiman et al. 2014). Recently, a metagenomic-derived cutinase-like enzyme, the polyester hydrolase PHL7 (Fig. 1b), was shown to possess a substrate-binding mode similar to that of LCC and to depolymerize PET films with higher efficiency, outperforming previous discovered PET-degrading enzymes (Sonnendecker et al. 2022; Richter et al. 2023).
I. sakaiensis secretes two types of hydrolases, an IsPETase (Fig. 1b, d) and an IsMHETase (Fig. 1b, e), that synergistically depolymerize PET into their monomeric products. IsPETase can establish a contact mainly with amorphous PET and hydrolyzes it into MHET as the major product, along with TPA, and BHET (Yoshida et al. 2016; Han et al. 2017; Joo et al. 2018). IsMHETase takes part in the final degradation process by exclusively hydrolyzing MHET into TPA and EG (Fig. 1a) (Yoshida et al. 2016). The biodegradation proceeds via the initial cleavage of the PETn polymer ester bond, producing a molecule of MHET and one of PETn−1, which is subsequently decomposed into another molecule of MHET and PETn−2. Depending on the binding site, BHET and TPA may also be generated in addition to MHET. The insoluble BHET is then further decomposed into MHET, while TPA and EG can be assimilated and used as carbon source by the microorganisms (see below) (Jehanno et al. 2022). PETases can be classified as type-II PET hydrolases, as they possess an extended loop that yields a subsite to bind PET (Fig. 1d) (Joo et al. 2018). Their catalytic active site is wider than that of cutinases, and the presence of an elongated cleft provides greater specificity over bulkier substrates like PET. The high native depolymerization activity, mainly against amorphous PET, and its preference for mild temperature regime (30 °C), make IsPETase the most studied and used PET-degrading enzyme. Recently, a new PETase-like enzyme was surprisingly discovered from the metagenome of human saliva. It possesses a Tm of 54 °C, 12° higher than that of IsPETase, and outperformed it in degrading PET powder by 68-fold (Eiamthong et al. 2022).
IsMHETase (Fig. 1b, e), characterized only in three organisms so far (Palm et al. 2019; Meyer-Cifuentes and Öztürk 2021; Perez-Garcia et al. 2023), can be classified within the tannase family, belonging to Block X of the α/β-hydrolase fold enzymes (Palm et al. 2019). In comparison to feruloyl esterase, the structurally closest enzyme, MHETases possess several additional loops defining the lid domain covering the active site (Fig. 1e). Important differences in enzyme function may be related to events of insertion, deletion, or recombination throughout evolution; hence, the presence of these supplementary structures could play a role in differentiating IsMHETase functions from other tannase enzymes (Palm et al. 2019).
Notably, the aerobic, rod-shaped bacterium Rhodococcus pyridinivorans possesses an unprecedented PET-degrading enzymatic apparatus composed of hydrolases exposed on the cell surface (Fig. 1b) (Guo et al. 2023a; b). Studies of this peculiar anchor structure could help in designing whole-cell bacterial systems for PET depolymerization (see below).
Carboxylesterases are lipolytic hydrolases that catalyze the formation and cleavage of short-chain carboxylic esters in a vast array of substrates (Johan et al. 2021). Interesting properties, such as a broad range of substrate specificity and a high degree of stability in inorganic solvents, make them promising biocatalysts for industrial purposes (Castilla et al. 2022). They possess the consensus motif, G-X-S-X-G, typical of serine hydrolases and cutinases (see above), and a catalytic triad composed of S-E-H (Fig. 1f). The polyesterases from the thermophilic actinomycete Thermobifida fusca (ThfCa) and Thermobifida halotolerans (ThhCa) displayed hydrolytic activity against BHET and MHET (Fig. 1a, b, f) (Billig et al. 2010; Ribitsch et al. 2012). This activity was exploited for generating a dual enzyme system (ThfCA-IsPETase) that outperformed the IsPETase enzyme alone (von Haugwitz et al. 2022). Recently, the nitrobenzylesterases BsEstB from B. subtilis and ChryBHETase from Chryseobacterium sp. PET-29 (Fig. 1a, b) were shown to convert BHET into TPA (Li et al. 2023). Thus, they can be applied in tandem with other PET-degrading enzymes for consuming BHET formed by PET degradation.
Lipases are hydrolytic enzymes active against short-chain carboxylic esters (Chandra et al. 2020). Unlike cutinases, they possess a lid covering the active site, which potentially reduces their activity against PET substrates (Castilla et al. 2022). Thus, they require interfacial activation to start the catalytic process. Lipases have been shown to have promiscuous activity against a broad range of different polyester substrates (Castilla et al. 2022). As early as 2005, the purified lipase from the green plant Triticum aestivum demonstrated partial hydrolysis activity on PET (Nechwatal et al. 2006). Since then, several other lipases from fungal (Eberl et al. 2009), and bacterial origins (Danso et al. 2018; Blázquez-Sánchez et al. 2022; Nikolaivits et al. 2022), have been reported to possess PET-degrading capabilities. Noteworthy, the specificity of Candida antarctica lipase B (CaLb) (Fig. 1g) over PET substrates can be modulated by changing the pH regime, with high-alkaline conditions favoring the partial hydrolysis of BHET into MHET and EG, while mild-alkaline to slightly acidic pH regimes favor full hydrolysis into TPA and EG (Fig. 1g) (Świderek et al. 2023). These are interesting features to be exploited for generating pH-controlled biotransformation systems.
Although the number of PET-degrading enzymes discovered has considerably increased over the years, the naturally occurring activity of these enzymes is not particularly suited for their industrial application as plastic biocatalysts. Hence, protein engineering strategies can be envisaged to both improve their efficiency and stability.
2.2 Improving catalytic efficiency of PET hydrolases
Improper balance between enzyme and substrate, low thermostability, limited enzyme-substrate affinity and inhibition due to oligomeric intermediates affect the catalytic efficiency of PET-degrading enzymes. For instance, the high degree of crystallinity of several PET polymers accounts for more rigid ester linkages, rendering them more resistant to enzymatic cracking (Thakur et al. 2023). Since PET tends to flex above glass transition temperature (around 70–80 °C), improving PET hydrolases’ Tm and Topt around these temperatures should also favor their hydrolytic activity on the more exposed target ester linkage (Thakur et al. 2023).
Enzyme engineering can tailor natural enzymes to desired activities through the modification of their amino acid sequences (Jemli et al. 2016). The goal of this process is to generate enzymatic variants with improved functional properties, such as catalytic performance, thermostability, reaction by-product inhibition, and substrate specificity (Bell et al. 2021).
Numerous new-to-nature enzymes, with potential industrial applications, have been efficiently produced over the last decade through several approaches to enzyme engineering guided by computational design (Bell et al. 2021). Here, we take a look at the most recent cutting-edge approaches for increasing PET-degrading enzymes' properties with an eye on some of the most notable enzymatic variants developed so far (Table 1).
2.2.1 Optimization of reaction conditions
Enhancement of enzyme performance can be achieved by optimizing general enzymatic catalytic conditions, including reaction temperature, enzyme concentration, pH values, and buffer concentrations. The exploration of the above-mentioned parameters was used to screen the best conditions favoring the activity of the cutinase HiC (Castro et al. 2019). To determine their optimal values for TPA production, multiple experimental designs and a path of steepest ascent strategy were applied sequentially, identifying pH and PET concentration as the most influential parameters. Under the newly optimized parameters, HiC dramatically enhanced its catalytic activity, achieving a tenfold improvement of TPA production from post-consumer PET (pc-PET) (Table 1). This approach represents one of the few examples of efficiently improving PET catalytic hydrolysis without appealing to enzyme engineering.
2.2.2 Rational and semi-rational approaches
The elucidation of the molecular mechanisms and amino acid architecture of several PET hydrolases (Tournier et al. 2020; Bååth et al. 2021) has opened the door to the application of semi-rational and rational approaches for enzyme engineering. IsPETase performs its activity only under mild conditions (30 °C) and extracellularly. Indeed, the lack of thermal stability in a potentially harsh environment can be detrimental to its durability and enzymatic performance over time. A comparison between the IsPETase structure and the structure of the thermostable cutinase from T. fusca (TfCUT2) allowed the identification of structural hot spots for increasing thermostability (Son et al. 2019). A double mutation (S121E and N172) applied in the flexible β6–β7 connecting loop region of IsPETase generated a new water-mediated hydrogen bond, able to stabilize thermal resistance (Tm value increased by 8.81 °C). The resulting enzymatic variant exhibited prolonged maintenance of PET degradation activity at both 30 °C and 40 °C temperatures (Son et al. 2019).
Similarly, rational studies on the structure of IsPETase helped in the identification of two key amino acid residues (S214 and I218, and histidine and phenylalanine in other homologous enzymes) associated with enzyme flexibility. A double mutation of these two residues redesigned the substrate binding-pocket of the enzyme, thus modifying the enzyme's preference for bulkier substrates but with a decrease of Tm values by around 10 °C (Chen et al. 2021). In combination with molecular docking, the enzyme contact-surface was used for detecting several amino acids located in the hydrophobic groove of the thermostable cutinase LCC as potential mutagenic targets for improving thermostability (Tournier et al. 2020). Among the enzyme’s variants derived from the targets, the LCCICCG variant showed a conspicuous 60% depolymerization yield improvement on post-consumer colored-flake PET waste (pcw-PET) in comparison to the LCC wild-type enzyme. Furthermore, the depolymerization yield on pcw-PET, calculated as grams of TPA produced per hour per gram of enzyme (corresponding to a final yield of 42.1 gTPA l−1 h−1), resulted 98-fold higher than that of the previously reported enzyme, the TfCut2 (Fig. 1a), achieved on easily degradable amorphous PET. The further introduction of several new mutations for increasing hydrophilic interactions on the protein surface and internal hydrophobic interactions stabilized the LCCICCG variant structure, conferring improved thermostability and depolymerization activity on both amorphous and crystalline PET (Table 1) (Zeng et al. 2022). The new variant (LCCICCG_RIP) showed approximately 1.6-fold higher formation of PET hydrolytic products MHET and TPA in comparison to LCCICCG at 74 °C using amorphous PET as a substrate (Zeng et al. 2022). In a similar fashion, molecular dynamics (MD) simulation and analysis were conducted on ChryBHETase and BsEstB to explore the interaction between PET hydrolase and its substrate in the presence or absence of the BHET substrate. The study revealed the presence of barrier structures, at the level of the lid covering the active site, that impedes the BHET to reach the active site. The identified barrier regions were repositioned by introducing a flexible linker (-GG-), and the resulting variants (ΔBsEst and ΔChryBHETase) displayed a 2- to threefold improvement over the respective wild-type enzyme in converting BHET into TPA (Li et al. 2023).
Properties of catalysts are generally correlated to substrate specificity, which in turn is often controlled by steric hindrance in the active site (Korendovych 2018). Thus, the structural visualization of the amino acid residues in the active site generally assists in finding targets for mutagenesis. A semi-rational protein engineering strategy, consisting of site-saturation mutagenesis (SSM) coupled with an expressly designed colorimetric assay for high-throughput screening of PET nanoparticles degradation, was efficiently used to screen different variants of IsPETase and the thermostable LCC (Pirillo et al. 2022a, b). The resulting variants, TS-ΔIsPET and ΔLCC, showed a 175% and 130% higher catalytic activity on PET substrates in comparison to their relative native enzymes (IsPETase and LCC) (Table 1).
Laborious and costly protein purification steps can limit the detection and production of PET-degrading enzymes' variants toward industrial application. Recently, a semi-rational approach that integrates expression and cell secretion (see below) and fluorescent product detection was created to enable high-throughput screening of IsPETase variants (Zurier and Goddard 2023). Detection of both monomeric (bis-(2-hydroxyethyl) terephthalate, BHET-OH, mono-(2-hydroxyethyl) terephthalate, MHET-OH, and terephthalic acid, TPA-OH), and oligomeric (ethylene terephthalate, OET) degradation products by fluorescence spectroscopy, enabled the identification of 12 (out of a library of 57) double mutant variants having a significantly higher depolymerase activity of the secreted proteins than the wild-type. This approach consents a quicker product detection in comparison to chromatographic methods, nonetheless it does not guarantee a linear correlation between enzyme concentration and depolymerization rate, thus it limits the determination of the enzyme catalytic efficiency.
2.2.3 Directed evolution
Directed evolution approaches can, in principle, be successfully applied with no structural knowledge of a protein (Markel et al. 2020). This method enables the identification of even remote amino acid mutations that can confer the protein new beneficial properties (Correddu et al. 2021).
Following this concept, IsPETase was utilized for a high-throughput directed evolution strategy, employing degenerate NNG/T codons for the mutagenesis process (Bell et al. 2022). The best-evolved variant, named HotPETase (Table 1), accounted for 21 mutations and displayed a sensational improvement in Tm, up to 82.5 °C, and in the rate of depolymerization over semicrystalline PET material (cry-PET) higher than that of other engineered enzymes, albeit with decreased longevity. HotPETase also showed conserved catalytic efficiency over a wide range of temperature regimes (40–70 °C) and the ability to depolymerize different plastic sources with diverse grades of crystallinity (Bell et al. 2022). A similar approach was very efficiently applied to create a synthetic variant of IsPETase with vastly improved catalytic performances (Shi et al. 2023). They utilized a fluorometric screening assay based on the cleavage activity of the enzyme over the synthetic compound bis BHET-OH, an analogue of BHET, toward the formation of the fluorescent compound TPA–OH. The catalytic efficiency of the evolved PET hydrolase variants was easily monitored by following the release of the fluorescent product TPA–OH. The most improved variant, called DepoPETase (Table 1), was used to achieve complete depolymerization of PC-PET flakes in 96 h under mild temperature conditions (Shi et al. 2023).
2.2.4 Machine learning aided approaches
Machine learning allows for the data-driven identification of existing but unseen protein patterns and correlates them to enzyme properties, thus proposing new enzymatic variants with likely improved features (Yang et al. 2019). This approach, coupled with classic molecular docking and/or directed evolution, offers the possibility to identify multiple enzymatic variants considerably improved in fitness and activity.
The machine learning optimal temperature prediction tool (Preoptem (Zhang et al. 2022)) and directed-evolution strategies were applied for redesigning the PET hydrolase, LCCICCG (Ding et al. 2023). The new six-point mutation variant exhibited increased Tm by 15 °C and 4.63-fold and 1.29-fold production improvement at 80 °C compared with LCCICCG and LCCICCG_RIP, respectively.
Using a computational approach based on the design and screening of about 200 variants of IsPETase, Cui et al. (2021b) were able to conspicuously increase IsPETase thermostability. The strategy utilized, named GRAPE (GReedy Accumulated strategy for Protein Engineering), makes use of four complementary algorithms to redesign stabilizing mutations and minimize the potential destabilizing ones. The best-generated variant (DuraPETase) displayed a substantial 300-fold improved activity on semicrystalline PET films (Table 1) in comparison to the native IsPETase (Cui et al. 2021b). The same strategy was adopted to maximize the activity of the highly active thermostable BhrPETase from the Bacterium HR29 (under review, Wu et al. 2023). The mutations designed by utilizing the GRAPE algorithm were then validated experimentally. The best variant, TurboPETase (Table 1), exhibited much-improved substrate binding flexibility, optimized protein surface charge–charge interactions, and augmented thermostability due to the insertion of additional disulfide bonds compared to the wild-type enzymatic form. Of particular interest is the abolishing of the “physical aging,” an issue affecting the activity of the LCCICCG, thus reducing the PET to TPA conversion loss.
A deep learning algorithm followed by a rational protein engineering approach was used to generate an improved variant of PET hydrolases, using IsPETase, DuraPETase, and ThermoPETase as protein scaffolds (Lu et al. 2022). Among the created variants, the Functional, Active, Stable, and Tolerant PETase (FAST-PETase) (Table 1) exhibited far superior PET-degrading capabilities in comparison to the best to date engineered PET hydrolases (DuraPETase, ThermoPETase, LCCICCG and LCC) (Table 1). Across a various range of environmental parameters (pH 6.5–8.0 and temperature 40–50 °C), FAST-PETase showed a substantial catalytic improvement (from 3.2- to 141.6-fold) and higher activity (from 2.4- to 38-fold) on model amorphous PET. Furthermore, the new variant was capable of fully depolymerizing various post-consumer PET substrates with different crystallinity grades (Table 1).
While data-driven approaches were cardinal for engineering the most performant PET-degrading enzymes, the depolymerization rate of high-crystalline PET is still very low and the reaction difficult to be accomplished by relying only on enzymatic catalysis. The creation of enzymatic systems based on temperature above 75 °C can have detrimental effects on the enzymatic performances. At these temperatures, PET transition from an amorphous to a crystalline state occurs quicker. Therefore, future studies for enzyme improvement may need to focus more on increasing efficiency rather than improving thermostability. The catalytic activity of various PET hydrolases is likely due to naturally occurring promiscuity over PET as well as similar hydrolysable substrates (e.g., chitin). Hence, the possibility of directly evolving an ancestor, promiscuous enzyme, among known hydrolases, to a new specific and efficient PET-degrading enzyme faces a infinite source of templates in the whole environment (Erickson et al. 2022; Giangeri et al. 2022; Karunatillaka et al. 2022; Ji et al. 2023c). However, enzyme engineering studies have been conducted almost exclusively on two model enzymes, the LCC and IsPETase, slowing down the investigation over new classes of PET hydrolases. Recent advancements in protein structure prediction techniques, exemplified by prediction programs such as AlphaFold2 and RoseTTAFold, and the continuous development of automating protein engineering processes (Beal and Rogers 2020; Martin et al. 2023; Rapp et al. 2024), have the potential to aid in the design of novel PET hydrolases discovered through database searches and accelerate their optimization (Jumper et al. 2021; Baek et al. 2023).
Thus, exploiting the genetic variability of new natural occurring enzymes will provide additional paths for protein engineering.
2.3 Improving PET degradation by enzyme co-localization
Enzymatic surface erosion of PET substrates involves the initial degradation of the exterior surface to expose the inner side of the substrate. However, only a few areas of the surface can be accessed by enzymatic activity. Spatial clustering of enzymes acting sequentially is an elegant approach, mainly used to improve synthetic metabolic pathways (Qiu et al. 2018). This concept relies on engineering enzymes involved in sequential catalysis reactions to collocate them in proximity through chemical and/or structural interactions. In this regard, several synthetic strategies can be adopted to improve enzyme/substrate interaction and minimize the long-range diffusion of intermediates, including creating enzyme fusion and multi-enzyme complexes (Ma et al. 2022; Monterrey et al. 2022).
2.3.1 Synergistic enzymatic reactions
Oligomeric intermediates produced during PET hydrolysis, including MHET and BHET, can inhibit the depolymerization of PET hydrolases (Wei et al. 2022). IsPETase cannot hydrolyze MHET and has low activity over BHET; thus, their accumulation during PET degradation can obstruct enzyme activity. Enzymatic cooperation represents an alternative way to evade inhibition effects caused by oligomeric intermediates (Ma et al. 2022). Recently, biochemical, structural, and bioinformatics studies were applied to better understand the mechanism of action of both extracellularly secreted PETase and MHETase from I. sakaiensis (Knott et al. 2020). The wild activity of IsPETase and IsMHETase toward PET was exploited by creating a chimeric protein composed of IsPETase and IsMHETase fused by flexible glycine-serine linkers of 12–20 amino acids (Knott et al. 2020). The activity of the new two-enzyme system was assessed against amorphous PET. The degradation activity was three times higher than the activity of the two equimolarly cocktailed enzymes (Knott et al. 2020). Thus, they proved the synergistic effect of IsPETase and IsMHETase on PET degradation and the applicability of this approach for minimizing the decline in activity due to product inhibition. Similarly, combining ΔBHETase (see above) with different PET hydrolases in a dual-enzyme system led to a significant rise in TPA production compared to using the hydrolase alone in a single-enzyme system (Li et al. 2023). The identification of new MHETase and BHETase-like enzymes (Knott et al. 2020; Meyer-Cifuentes and Öztürk 2021; Li et al. 2023; Perez-Garcia et al. 2023), possessing distinctive specificity toward MHET and BHET substrates, may expand the range of candidates for generating new chimeric proteins.
One of the limiting steps likely impairing the efficiency of hydrolases in depolymerizing PET relates to their binding to its surface. Multi-domain architectures are common structures often adopted in nature by enzymes involved in polymer deconstruction (Graham et al. 2022). These modules are generally composed of a catalytic domain and a non-catalytic one. Several non-catalytic binding domains, including carbohydrate-binding modules (CBMs), hydrophobic proteins, and anchor peptides (AP), have been applied to enhance the binding activity of PET-degrading enzymes over PET substrates (Liu et al. 2022b).
2.3.2 Carbohydrate binding modules (CBMs)
CBMs, first discovered in microorganisms degrading chitin polymer chains, promote substrate accessibility of hydrolytic enzymes (Vaaje-Kolstad et al. 2010). Based on the substrate specificity and mode of binding, they can be grouped into different types: A (planar-type), B (endo-type), C (exo-type) (Graham et al. 2022). Type A recognizes the hydrophobic surface of polymers and has thus been utilized as an enhancer for PET hydrolysis. Two chimeric versions of the ThcCut1 cutinase (Fig. 1b) were constructed by fusing the CBMs from the cellobiohydrolase I from the fungus Trichoderma reseei and from a polyhydroxyalkanoate depolymerase from the gram-negative Alcaligenes faecalis (Ribitsch et al. 2013). Both CBM-fused enzymes displayed augmented PET-degrading activity but also diminished activity on soluble subproducts due to steric hindrance.
More recently, another CBM from the chitin-binding domain (ChBD) from Chitinolyticbacter meiyuanensis was fused, via a linker region, to the cutinase variant LCCICCG (Table 1) (Xue et al. 2021). ChBD-LCCICCG increased cutinase performance on high-crystalline PET (40%), realizing 11.6 times more degradation products, but showed performances decreasing over time likely due to steric inhibition by the PET subproducts. Site-directed mutagenesis targeting the binding site of the CBM from the endo-beta-1, 4-glucanase A (CenA) from Cellulomonas fimi was applied to increase the activity of the T. fusca cutinase–fusion protein (Thsf-CenA) (Zhang et al. 2013). The mutation enhanced the hydrophobic interaction with PET fiber, leading to an increment in PET fiber hydrolysis. In this context, several aromatic amino acids within the CBM-binding modules were identified as crucial for tuning the affinity of the dimeric CBM-PET hydrolase fusion protein to the substrate (Rennison et al. 2023).
2.3.3 Hydrophobin proteins
Hydrophobins are low molecular weight self-assembled proteins extracellularly secreted by fungi that can promote interaction with hydrophobic leaf surfaces (Bayry et al. 2012). Thus, hydrophobic binding modules (HBM) may assist PET hydrolytic enzymes in targeting the highly hydrophobic PET surfaces. The covalent fusion of HBM to PET hydrolytic enzymes should increase the enzyme concentration on the PET surface and, therefore, enhance hydrolysis. The hydrophobins HFB4 and HFB7 from the saprophytic fungus Trichoderma sp. were C-terminally fused to the cutinase Cut1 from T. cellulosilytica (Ribitsch et al. 2015b). The resulting chimeric protein exhibited enhanced adsorption activity on the surface of PET film. Moreover, the presence of the fungal hydrophobin RolA from Aspergillus orizae could stimulate IsPETase hydrolysis when the PET samples were preincubated with the hydrophobic protein (Puspitasari et al. 2021). The activity of RolA on the PET surface may decrease the PET surface hydrophobicity, thus empowering IsPETase to effectively contact the PET surface.
2.3.4 Anchor peptides (APs)
APs are small molecules with size ranging from 20 to 1000 amino acids. Depending on their structures and physical properties, they can promote the adhesion and interaction with several macromolecules (Dittrich et al. 2022). Because of their adhesive properties, they have been tested as facilitators of adhesion to PET substrates. Thus, surface-specific anchor peptides can be genetically fused to the target protein. The dermaseptins DSI, host defense peptide produced by the skin of frogs, were shown to independently increase the adhesion of the biocatalyst phenolic acid decarboxylase to the PET surface (Büscher et al. 2019). This property was then exploited for increasing the PET-degrading capabilities of the cutinase ThfCut2 from T. fusca (Liu et al. 2022b). The DSI-ThfCut2 chimeric protein displayed a 22.7-fold improvement in depolymerizing PET amorphous particles; it also outperformed other dimeric modules constructed using different binding domains, including Bacillus anthracis CBM (BaCBM2) (Weber et al. 2019), the anchoring peptide Tachystain A2 (TA2) (Islam et al. 2019) and the hydrophobins HFB4 and HFB7 (Ribitsch et al. 2015b). Very recently, this approach was readapted to the LCCICCG variant (Table 1), to increase its PET depolymerization efficiency. The α-synuclein (αS), a mammalian protein abundant in presynaptic terminals in both the central and peripheral nervous systems, can promote stable contact with hydrophobic surfaces (Bhak et al. 2020). The αS C-terminal sequence (YEMPSEEGYQDYEPEA) was genetically fused to the C-terminal of ICCG using a flexible linker peptide (GGGGS)2 as a separator. The resulting fusion protein (LCCICCG-αSP) exhibited enhanced degradation stability over time and vastly improved catalytic efficiency compared to the LCCICCG variant (up to 3.7-fold) on p-nitrophenyl butyrate (p-NPB) (Su et al. 2022).
Fusion strategies applied so far have shown to increase enzyme access to the PET surface. This effect, however, seems to be enzyme-dependent; thus, the fusion strategy that works with one PET-hydrolase may not function with a different biocatalyst (Graham et al. 2022), making the detection of a suitable fusion partner more challenging. The high hydrophobicity of the PET surface stands as a challenging obstacle to overcome only relying on enzymatic activity. Additional thermal pretreatments are still necessary for modifying the PET hydrophobic surface and reducing its crystallinity rate. However, similar to the cellulose degradation enzymatic systems (Anandharaj et al. 2020), the construction of synthetic scaffolds comprising multi-modular PET-degrading hydrolases may offer alternative solutions for synergistically increasing enzymatic activity, minimizing product inhibition, and improving substrate adhesion.
3 Optimization of microbial systems for the production of PET-degrading enzymes
Industrial-scale enzymatic degradation requires the production of large amounts of functional enzymes, which are laboriously purified from microbial cultures grown in fed-batch or continuous fermenters (Xu et al. 2023). Thus, enzyme production at the industrial scale can be simplified by selecting a suitable microbial source equipped with either a wild or synthetic efficient protein secretion system. PET-degrading enzymes have been efficiently expressed in various microorganisms, and several metabolic engineering approaches have been adopted to increase their production and catalytic efficiency (Mudondo et al. 2023). Below we describe the approaches aimed at increasing enzyme secretion from the host cell factory, enhancing proteins’ three-dimensional shapes and creating microbial-based whole-cell PET biocatalysts.
3.1 Optimization of microbial protein-secretion system
The high molecular weight of PET polymers physically impedes their cellular internalization. Although heterologous PET hydrolase expression and secretion have been achieved in several microorganisms, their secretion has been too inefficient to achieve PET degradation. This limit has been tackled by both tailoring PET hydrolases’ endogenous signal peptide (SP) and improving the host’s protein secretion systems.
The improved engineered version of I. sakaiensis PETase (PETaseR280A) (Joo et al. 2018) was expressed in the photosynthetic diatom Phaeodactylum tricornutum (Moog et al. 2019). The replacement of the endogenous I. sakaiensis SP with the SP of P. tricornutum alkaline phosphatase dramatically increased PETaseR280A secretion. The recombinant secreted protein retained its PET hydrolytic activity even in saltwater-based environments. Conversely, the endogenous I. sakaiensis SP failed to promote IsPETase secretion when expressed in the green algae Chlamydomonas reinhardtii (Kim et al. 2020). While the protein secretion system of micro and macroalgae has not been well elucidated yet, these works highlight the suitability of photosynthetic organisms as potential photo-biodegradable plastics agents.
Several known sec-dependent signal peptides (SDSP) were tested in Escherichia coli for their capability to better promote IsMHETase extracellular translocation (Sagong et al. 2020). Among the signal peptides tested, the maltose/maltodextrin-binding periplasmic protein (SPMalE), thiol/disulfide interchange protein (SPDsbA), TolPal system protein (SPTolB) and maltoporin (SPLamB) supported protein secretion, with the latter resulting to be the most efficient (2.5 mg L−1). On the contrary, the periplasmic molecular chaperone SurA (SPSurA) and the heterologous SP in pectate lyase II from Pectobacterium carotovorum (SPPelB) failed in promoting IsMHETase secretion (Sagong et al. 2020). SPLamB accounted for the best SP also for the extracellular exportation of IsPETase (6.2 mg L−1) (Seo et al. 2019). IsPETase excretion was also improved by genetically modifying the SPLamB sequence via random mutagenesis. The evolved version of E. coli endogenous SPPelB led to a 1.7-fold increase in IsPETase secretion (Shi et al. 2021). Follow-up studies showed how synthetic biology could favor increasing SP protein translocation efficiency. SP enhancers, derived from the native SP of β-fructofuranosidase from Arthrobacter arilaitensis, were shown to be strong protein secretory signals (He et al. 2014). The N-terminal fusion of SP enhancers (B1) with SPOmpA, SPMalE and SPPelB dramatically boosted their ability to mediate protein secretion in E. coli (Cui et al. 2021a). This increment was particularly appreciable in the SPPelB: the yield of IsPETase excreted into the external medium reached 650 mg L−1, about 60-fold higher than that obtained with the wild-type SPPelB (Cui et al. 2021a).
Microorganisms naturally equipped with a sophisticated protein secretion machinery, such as Yarrowia lipolytica and B. subtilis, were also utilized as hosts for extracellular PETase production (Huang et al. 2018; Kosiorowska et al. 2022). Compared to the E. coli expression system, they exhibit a lower tendency to form inclusion bodies, much higher secretion capacity, and advantageous conditions for protein folding (Guo et al. 2022). Recently, both E. coli and B. subtilis expression systems were compared in the production of the cutinase ThfCut2 from T. fusca. B. subtilis showed superior extracellular secretion capability, and the secreted cutinase was more active and thermostable (Wei et al. 2019). Several different endogenous SPs, together with SPs from E. coli, were tested in B. subtilis (Huang et al. 2018): curiously, the native SP from IsPETase was more efficient in guiding the enzyme into the extracellular milieu, reaching the best extracellular yield (15 mg L−1) when the twin-arginine translocase (TAT) system was inactivated. Moreover, the yield of secreted heterologous proteins often suffers the degrading activity of extracellular exoproteases (Wu et al. 2016; Hughes et al. 2022). This could be circumvented by utilizing a B. subtilis strain mutant, with a depleted proteolytic apparatus (B. subtilis WB600). Using this strain in combination with a weak promoter (P43), for reducing the metabolic burden of expression, the translation and the extracellular secretion of IsPETase were improved, but the activity on PET-film was hampered (Wang et al. 2020). Indeed, despite major advances accomplished in the modification of the microbial secretory apparatuses for enhancing recombinant protein recovery, the metabolic mechanisms underlying protein translocation and secretion remain mostly unknown, which clamp down further metabolic engineering progresses. Since protein secretion is a mechanism heavily relying on the correct protein folding, and secreted proteins fold slower than the non-secreted ones (McCaul and Braakman 2022), optimization of intracellular chaperones is another strategy already used for improving production of PET-degrading enzymes.
3.1.1 Metabolic engineering for enhancing protein folding
While correct protein folding is directly linked to extracellular exportation, incorrect folding also causes formation of intracellular inclusion bodies in E. coli. Co-expression of chaperone proteins is a strategy often utilized in E. coli to attenuate this event. The co-expression of GroEL/ES chaperone proteins along with ThermoPETase resulted in a 12.5-fold higher intracellular expression than that of ThermoPETase alone. Additionally, the C-terminal fusion of the antiterminator NusA promoted higher ThermoPETase solubility (final yield 80 mg L−1) and conferred improved long-term catalytic activity on PET with low crystallinity (Aer et al. 2022). An analogous approach was utilized to create a chimeric protein composed of IsPETase and the carbohydrate-binding module (CBM66) from the exo-levanase of B. subtilis (SacC). As said above, CBMs are parts of carbohydrate-active enzymes with carbohydrate affinity; they are also extensively used to increase protein solubility. The N-terminal fusion of the CBM66 tag increased IsPETase solubility and led to a 3.7-fold higher expression amount of protein (approximately 370 mg L−1) compared to the wild-type IsPETase (Ko et al. 2021). Most proteins secreted by eukaryotic organisms are subjected to post-translational modifications (PTM) of one or more amino acids (Ramazi and Zahiri 2021). Although much less abundant and more diverse, PTMs are also present in bacteria (Macek et al. 2019), where they generally serve to modulate protein properties, rendering them more stable against degradation and fluctuation of temperatures and pH (Prabakaran et al. 2012). The heterologous expression of IsPETase in the methylotrophic yeast Komagataella phaffa (Pichia pastoris) resulted in a high degree of N- and O-glycosylation. As a result, the protein displayed augmented thermostability and higher specific activity over post-consumer PET trays than IsPETase expressed in E. coli (Deng et al. 2023). Furthermore, the covalently attached glycans on the protein surface increased enzyme solubility and its resistance to proteolysis.
Employing microbial expression systems for the production and secretion of PET hydrolases gives great benefits in terms of productivity, i.e., potential high yield of production in a relatively short period of time. However, it also entails several drawbacks. Secreted enzymes often exhibit a short catalytic lifespan and non-reusability, which may require the adoption of costly purification processes that could restrain their industrial applications. Another strategy used for circumventing these problems is the immobilization of the biocatalysts (Grunwald 2018).
3.1.2 Extracellular enzymatic decoration
Recently, bacterial cells were exploited as whole-cell living recycler platforms (Jia et al. 2022; Zhu et al. 2022b). A synthetic three-component enzymatic complex comprising the IsPETase fused to the class II hydrophobin HFBII from the mesophilic filamentous fungus Trichoderma reesei and the outer membrane-bound fatty acid transporter FadL was created in E. coli (Jia et al. 2022). FadL ensured correct IsPETase-HFBII cell surface display, while HFBII facilitated microbial PET substrate colonization and biodegradation (approximately 0.32 g L−1 of TPA in 7 days, 21 and 37% higher than that generated using E. coli displaying only the IsPETase and the free IsPETase, respectively) (Jia et al. 2022). In a similar fashion, the yeast K. phaffa was engineered to simultaneously display the IsPETases and hydrophobin proteins on the cell surface (see above) (Macindoe et al. 2012; Chen et al. 2022). The concomitant cell surface decoration with IsPETase and the T. reesei (HFBI) led to a 300-fold increase in degradation efficiency compared to that of the native IsPETase toward PET with 45% of crystallinity at 30 °C, and a conversion rate of 10% in ten days (Chen et al. 2022). Moreover, the applicability of binding domains and anchor peptides as hydrophobic enhancers for surface adsorption was successfully tested in vitro (see above) using various modules from different organisms (Ribitsch et al. 2013, 2015a; Puspitasari et al. 2021; Xue et al. 2021; Liu et al. 2022b). Recently, a platform termed biofilm-integrated nanofiber display (BIND), permitting the efficient anchorage of proteins on the curli (proteinaceous amyloid nanofibers) of E. coli has been proposed (Nguyen et al. 2014). This strategy was used to immobilize IsPETase and create a BIND-PETase system whose degradation efficiency after 7 days ranged between 5 and 7%, depending on the crystallinity nature of the post-consumer PET used (Zhu et al. 2022b). Although the efficiency of the system needs improvement, this work once again demonstrated the applicability of synthetic biology tools to the plastic recycling industry.
The constitution of whole-cell PET degradation systems is hampered by the scarce knowledge of the catalytic properties of the diverse useful enzymes and of the detailed degradation pathways. The recent detection of a bacterium naturally displaying a PETase-like enzyme on its cell surface (Guo et al. 2023a; b) may serve as a model for better understanding enzyme-membrane anchoring mechanisms toward the generation of a self-sufficient whole-cell biocatalyst.
4 Strategies for PET upcycling
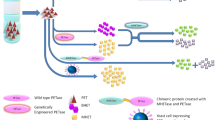
The microbial biodegradation of PET plastics is achieved by three consecutive steps, which may be performed by a whole bacterial community: bio-fragmentation, assimilation, and mineralization (Fig. 2). Bio-fragmentation is an enzymatic process carried out by the activity of PET hydrolase enzymes. Microbes then assimilate the PET-derived monomers and direct them into the central carbon metabolism, in a process of molecular recycling that can be enhanced via metabolic engineering. In the following paragraph, we explore some strategies applied for the utilization of EG and TPA as metabolic feedstocks and their sequential valorization (Fig. 2). Furthermore, we also describe several attempts for harnessing microorganisms for creating self-sufficient whole-cell biocatalysts.
4.1 Valorization of the PET-derived monomer ethylene glycol (EG)
EG, a C2 compound formed during PET hydrolysis (Figs. 1, 2), holds promise as a substrate for the orthogonal production of various chemicals. In contrast to glucose, it minimally interferes with pathways for biomass and target chemical production, including dicarboxylic acids, alcohols, diols, and terpenoids (Pandit et al. 2017; Panda et al. 2023; Wagner et al. 2023a). So far, only a few bacterial species are known to possess the metabolic arsenal for utilizing EG as a carbon source. In anaerobic microorganisms, such as Clostridium spp. and Acinetobacter spp., the oxygen sensitive diol-dehydratase dehydrates EG to acetaldehyde. Acetaldehyde is then disproportionated to ethanol and acetyl coenzyme A (Ac-CoA) by a CoA-dependent acetaldehyde dehydrogenase (PduP) and an NADH-dependent alcohol dehydrogenase (Fig. 3) (Gaston and Stadtman 1963; Trifunović et al. 2016). In aerobic catabolism, conserved in several Pseudomonadaceae spp. and Flavobacteriaceae spp., EG is sequentially oxidized to glyoxylate (Fig. 3) (Child and Willetts 1978; Mückschel et al. 2012; Li et al. 2019). Moreover, this natural pathway was recently demonstrated in the yeast Y. lipolytica (da Costa et al. 2020; Carniel et al. 2023). Glyoxylate is a gluconeogenic substrate common in most prokaryotic and eukaryotic organisms (Chew et al. 2019), serving as a hub metabolite in the central carbon metabolism (Fig. 3) (Yang et al. 2022). Thus, EG can potentially be converted, in only three enzymatic steps, into a valuable metabolite and/or be potentially pushed towards the generation of a desired chemical compound for completing PET upcycling.
Schematic representation of the metabolic routes for EG and TPA cellular internalization. The pathways depicted encompass both native and novel biosynthetic routes sourced from various organisms, each associated with the production of specific chemical compounds. Enzymes, highlighted in light blue, facilitate these processes. Different pathways are enclosed in various colored shapes, each described in the legends provided in the upper part of the figure. The back-flipped “T” symbol, accompanied by the enzyme name colored in red, signifies the transcriptional regulator controlling the pathway. Consecutive arrows denote multiple enzymatic steps. Dashed lines indicate connections between different metabolic pathways. Dark blue lines represent the cellular uptake of compounds (e.g., EG, TPA, polyolefins) from the extracellular environment. List of proteins with relative UniProt (https://www.uniprot.org/) (Bateman 2019) IDs: FucOMut: P0A9S1; Gox0313: Q5FU50; AldA: P25553; EcFsaA: P32669; EcKdsD: P45395; EcRpe: P0AG07; CaPkt: C6PNC7; RpPduP: Q21A49; GCCMut: H1KHB8; TCR: A9WIU3; EcFsaATA: P32669; PcTadH: A0A1X7DDC2; TtLac11: A0A286RDQ9; AaAraD: F0Q4R8; EcMDH5Q: P61889; GlcDEFG: Q88GH8, Q88GH7, Q88GH6, Q88GH5; PP_3749: Q88GH4; GlxC: Q88F03; GclR: Q88F16; PhaA: P14611; PhaB: P14697; PhaC: P23608; PhaJ: Q88KS2; Dxs: Q8NPB2; Idi: 4QG13; CrtE: Q93QX8; Phefbr: P0A9J8; TyrA: P07023; SeTal: K0K4B0; TyrR: P07604; AroGMut: P0AB91; TphC: A0A5N7XFM8; TpaK: Q0RWE8; TphA1II: Q3C1D2; TphA2II: Q3C1D5; TphA3II: Q3C1D4; TphB: Q3C1D3; RnOmtMut: B6CZ62; HsOmt: B8XPJ7; NiCAR: Q6RKB1; AroY: B2DCZ6; PhKLMNOPQ: Q93JV2, Q93JV2, Q84AQ3, Q84AQ2, Q84AQ1, Q84AQ0, A0A0S2UPA7; CatA: Q6XUN7; PcaGH: P00437, P00436; PcaB: Q88N37; PcaC: Q88N35; PcaD: Q88N36; PcaJ: P0A101; PobAMut: Q88H28; LigA: P22635; LigB: P22636; LigC: Q9KWL3; DcaPK: Q937S8, Q937T1; DcaIJ: Q937S9; DcaA,: Q937T2; PsrA: Q88KZ5; PaaX: Q88HR7. (Color figure online)
The application of synthetic non-natural pathways for substrate utilization toward chemical production allows a high degree of orthogonality at less energetic/cellular costs (Pandit et al. 2017). Hence, an E. coli strain encoding a synthetic EG assimilation pathway was used for evaluating EG utilization as feedstock and for the production of glycolate (Pandit et al. 2021). The oxygen-sensitive E. coli enzyme 1, 2-propanediol oxidoreductase (FucO) was mutated by the introduction of two-point mutations (I7L, L8V), to create an O2-tolerant variant (FucOMut) with augmented specificity on EG. An E. coli strain expressing this genetic variant converted EG to glycolaldehyde (GlAl), while the overexpression of the native glycolaldehyde dehydrogenase (aldA) ensured efficient oxidation to glycolate (Fig. 3 and Table 2). Further metabolic modelling analysis identified oxygen as a key parameter for optimizing glycolate yield and ensuring dynamic metabolic control (Hartline et al. 2021; Pandit et al. 2021). The E. coli strain carrying the same EG entry point pathway (fucOMut_aldA) was also utilized as a basis for further genetic modifications to convert EG into aromatic chemicals (Panda et al. 2023). Firstly, the HTH-type transcriptional regulatory protein (tyrR) and the prephenate dehydratase (pheA) genes were genetically knocked out to diminish repression on L-tyrosine production and stop L-phenylalanine formation, respectively. L-tyrosine production from EG was then achieved by overexpressing the L-tyrosine operon tyrA_aroG, encoding an aminomethyltransferase and a deoxyheptonate aldolase (Fig. 3). Secondly, additional expression of the setal gene, encoding a tyrosine ammonia lyase from Saccharothrix espanaensis, directed the metabolic flux towards p-coumaric acid (Fig. 3 and Table 2). Conversely, genetic strategies involving the double knockout of tyrR and tyrA genes, coupled with the overexpression of mutated pheA (phefbr, T326P) and aroG (D146N) genes (Zhou et al. 2010)), were employed to convert EG to L-phenylalanine (Fig. 3 and Table 2).
Using a series of structure-guided rational design and directed evolution techniques, enzymatic variants with new-to-nature activities and substrate specificities were generated to create a synthetic pathway for efficient EG conversion (Fig. 3) (Scheffen et al. 2021). The new enzymatic variant of the biotin-dependent propionyl-CoA carboxylase (GCCMut), carrying five amino acid substitutions (D407I, Y143H, L100S, I450V, W502R), from Methylorubrum extorquens, exhibited a 30-fold increase in catalytic efficiency for glycolyl-CoA carboxylation and decreased affinity toward its natural substrate propionyl-CoA. In parallel, the bifunctional malonyl-CoA reductase (TCR) from Chloroflexus aurantiacus converted tartronyl-CoA directly to glycerate. This new enzymatic route (tartronyl-CoA (TaCo) pathway (Scheffen et al. 2021)) forms a carboxylation module for the conversion of glycolate (C2) to glycerate (C3) (Fig. 3). This carboxylation module was then combined with a new EG-utilization module. The NAD-dependent alcohol dehydrogenase (Gox0313) from the acetic acid bacterium Gluconobacter oxydans oxidizes EG into glycolaldehyde, while the aldehyde dehydrogenase from Rhodopseudomonas palustris (RpPduP) converts glycolaldehyde into glycolyl-CoA. The combination of this EG module with the TaCo pathway, along with the integration of an ATP regeneration system, ensured direct glycerate production from EG (Fig. 3).
After an extensive search, leading to the identification and characterization of potential candidate enzymes with the desired catalytic activities, a five-step synthetic pathway was constructed for the carbon-conserving biosynthesis of 2,4-dihydroxybutyric acid (DHB) from EG (Fig. 3) (Frazão et al. 2023). After the first conversion of EG to glycolaldehyde by Gox0313 (Fig. 3), the modified version of the D-fructose-6P aldolase from E. coli (EcFsaATA, carrying two amino acid substitutions (L107Y, A129G)) catalyzed the reversible condensation of two molecules of glycolaldehyde to yield D-threose. Then, the D-threo-aldose 1-dehydrogenase from Paraburkholderia caryophylli (PcTadH) was employed for the oxidation of the four-carbon sugar into D-threono-1,4-lactone. The consecutive activities of the gluconolactonase from Thermogutta terrifontis (TtLac11) and the D-arabinonate dehydratases from Acidovorax avenae (AaAraD) ensure the synthesis of 2-oxo-4- hydroxybutyrate (OHB) (Fig. 3). The malate dehydrogenase mutant EcMdh5Q, carrying five-point mutations (I12V, R81A, M85Q, D86S, G179D), catalyzed the final reduction to yield DHB (Fig. 3). An E. coli strain engineered with this new enzymatic route produced DHB from EG. The pathway exhibited a 51% carbon yield improvement compared to the previous metabolic routes tested in E. coli for producing DHB (Walther et al. 2017). This underscores the effectiveness of designing stoichiometrically favored enzymatic reactions as a potent strategy to generate valuable carbon-efficient metabolic routes.
Recently, the cellular applicability of the synthetic Ara5P-dependent glycolaldehyde assimilation (GAA) pathway (Trudeau et al. 2018) was showcased in E. coli for evaluating the conversion of EG to Ac-CoA (Wagner et al. 2023b). The synthetic pathway, extrachromosomally expressed, departed with two enzymatic reactions catalyzed by FucOMut and FsaA to convert EG into arabinose 5-phosphate (Fig. 3). Then, the native E. coli enzymes, D-arabinose 5-phosphate isomerase (EcKdsD) and D-ribulose 5-phosphate 3-epimerase (EcRpE) produced the ketose sugar xylulose-5-Phopshate (Xu5P). Finally, the phosphoketolase from Clostridium acetobutylicum (CaPkt) cleaved Xu5P into glyceraldehyde 3-phosphate (GA3P) and acetyl phosphate (AcP), which is ultimately converted to Ac-CoA by the endogenous phosphate acetyltransferase (Pta) (Fig. 3). The poor performance of the pathway was attributed to two major bottlenecks: the very low affinity of FsaA toward GA and the thermodynamically unfavorable oxidative reaction to convert EG into GlAl. Future genetic implementations may, therefore, focus on engineering more efficient enzymes with higher substrate specificity and on creating stable microbial strains with genomically integrated synthetic pathways to minimize the metabolic burden and avoid plasmid instability (Saleski et al. 2021).
4.2 Valorization of the PET-derived monomer terephthalic acid (TPA)
The necessary pathway for the turnover of PET-derived aromatic compounds has been identified in few bacterial genera, including Ideonella (Yoshida et al. 2016), Rhodococcus (Choi et al. 2005; Patrauchan et al. 2005), Comamonas (Sasoh et al. 2006), Delftia (Shigematsu et al. 2003), Paraburkholderia (Chain et al. 2006) and Pseudomonas (Narancic et al. 2021). TPA is converted into protocatechuic acid (PCA) via a pathway encoded by the tphABC operon (Fig. 3) (Sasoh et al. 2006). The operon regulator TphR senses TPA and, in turn, activates the enzymatic routes for its assimilation and catabolism. The periplasmic protein TphC, a member of the tripartite tricarboxylate transporters (TTT), facilitates the cellular uptake of TPA (Gautom et al. 2021). In the cytoplasm, TPA is converted into 1,6-dihydroxycyclohexa-2,4-diene-dicarboxylate (DCD) by the O2-dependent terephthalate dioxygenase (TPADO), a three-complex metalloenzyme member of the family of Rieske oxygenases (ROs) (Kincannon et al. 2022). Then, the zinc-dependent dehydrogenase TphB (1,2-dihydroxy-3,5-cyclohexadiene-1, 4-dicarboxylate dehydrogenase) converts DCD into PCA, a valuable metabolite that can easily be funneled into the central carbon metabolism (Fig. 3). This complex system represents the entry point of plastic recycling and offers an interesting starting point for the bioconversion of TPA into industrially relevant compounds.
Inspired by the native biosynthetic path for TPA assimilation, several metabolic engineering studies delineated the applicability of TPA as a feedstock for microbial growth on the route to complete cell-based PET valorization. Several bacterial-based systems have been efficiently applied for the bioconversion of TPA into different valuable aromatic and non/aromatic compounds (Fig. 3) (Kim et al. 2019). Firstly, the expression of the tphABC operon genes, along with the tphB gene in E. coli, allowed the new genetically modified strain (PCA-1) to correctly internalize TPA and efficiently convert it into PCA. The PCA-1 strain was further engineered with a protocatechuate decarboxylase (AroY) from Enterobacter cloacae, for the conversion of PCA into catechol (strain PCA-2) (Fig. 3). These two strains were then used as the basis for further genetic modifications. The PCA-2 strain, expressing a catechol 1, 2-dioxygenase (catA) gene from Pseudomonas putida KT2440, and the PCA-1 strain, expressing an O-methyltransferase (omt) gene from Homo sapiens, efficiently converted TPA into muconic acid (MA) and vanillic acid (VA), respectively (Fig. 3, Table 2) (Kim et al. 2019). Follow-up studies on the conversion of TPA into the reduced VA form, vanillin, further optimized the final yield by combining metabolic engineering and classic multiparametric experimental design (Sadler and Wallace 2021). These experiments were carried out on E. coli strain with reduced aromatic aldehyde reduction (RARE), due to the deletion of genes encoding enzymes active on aldehyde substrates (Kunjapur et al. 2014). E. coliRARE was engineered for the heterologous synthesis of a carboxylic acid reductase from Nocardia iowensis (NiCAR) and a single-point mutant of the Omt from Rattus norvegicus (RnOmt, Y200L) (Fig. 3). Additionally, a phosphopantetheinyl transferase (psfP) from Bacillus subtilis was also expressed since it is required for post-translational modification of apoCAR to yield the functional holo-CAR enzyme (Horvat and Winkler 2020). Negligible production of vanillin was improved by modulating culture conditions, such as pH (5.5), in situ product removal for minimizing vanillic acid cell toxicity, and the use of oleyl alcohol for increasing intracellular TPA uptake (Table 2) (Sadler and Wallace 2021). However, the efficiency of the heterologous metabolic architecture was limited by the modest membrane permeability to TPA. This highlights the need to consider the expression of transporters specific for or adapted to the host membrane when designing microbial systems for PET upcycling. A two-strain system approach, composed of the E. coli strain PCA-1 and another E. coli strain expressing a double mutant version of p-hydroxybenzoate hydroxylase (pobAMut, T294A, Y385F) from P. putida KT2440, was used to convert TPA into gallic acid (GA) (Fig. 3, Tables 2). Moreover, a mixed co-culture of PCA-1 and the new E. coli strain expressing the phKLMNOPQ operon, encoding a phenol hydroxylase from Pseudomonas stutzeri OX1, efficiently produced the antioxidant compound pyrogallol (PG) from TPA (Fig. 3, Table 2).
Similarly, Kang et al. (2020) employed a hybrid chemo-microbial methodology, wherein PET underwent initial chemo-depolymerization into TPA, followed by its microbial valorization into 2-pyrone-4, 6-dicarboxylic acid (PDC) (Fig. 3 and Table 2). TPA conversion into PDC was achieved by a two-microbial strain system approach. The authors made use of the PCA-1 strain (see above) and an E. coli strain expressing the ligAB genes encoding a 3, 4-dihydroxybenzoic acid-4, 5-deoxygenase and the ligC gene encoding a carboxymethyl muconate semialdehyde dehydrogenase (CHMS) from Sphingomonas paucimobilis (Fig. 3). The co-culture system exhibited almost complete conversion of TPA into PDC (Table 2).
This work further highlights how the co-colture of two (or more) strains may be advantageous for simultaneously expressing large multigene systems. The high expression level of multi-gene heterologous pathways may reduce ribosome availability for the translation of native as well as exogenous proteins and, in turn, the concentration of metabolic precursors, causing cell growth defects and reduced productivity. The utilization of microbial consortia, in which reactions of long biosynthetic pathways are distributed among different strains working synergically, can reduce the metabolic burden on a single strain and may be more advantageous for generating productive microbial platforms.
4.3 PET depolymerization and simultaneous valorization of its hydrolyzates EG and TPA
While microbial-based approaches for PET valorization described so far focused solely on either TPA or EG conversion, other microbial systems were developed for concurrently converting terephthalic acid and ethylene glycol into valuable products. These integrated strategies potentially present multiple benefits compared to individual conversions, such as maximization of resource utilization and simplification of the overall process, potentially leading to cost savings and streamlined production (Liu et al. 2022a; Diao et al. 2023). To this end, several strategies were employed, leveraging cross-disciplinary approaches that combine chemical depolymerization with microbial fermentation, and utilize whole-cell biocatalysts.
4.3.1 Hybrid chemo-enzymatic/microbial processes for PET degradation and valorization
Werner et al. (Werner et al. 2021) demonstrated β-ketoadipic acid (βKA) biosynthesis through a method that integrates chemical breakdown with biological upcycling of PET. The BHET generated from PET by chemocatalytic glycolysis was further metabolized via TPA catabolism, facilitated by an engineered strain of P. putida KT2440. The hydrolysis of BHET into TPA was guaranteed by the heterologous expression of IsPETase and IsMHETase. Correct TPA catabolism was ensured by a heterologous pathway composed of the tphAB operon from C. camamonas and tpaK gene encoding a terephthalate transporter from Rhodococcus jostii (Fig. 3). The genetic knockout of the pcaiJ gene, encoding a 3-oxoadipate CoA-transferase, promoted β-ketoadipic acid (βKA) production. In parallel, EG utilization was incremented by genetic knockout of the glyoxylate transcriptional regulator (gclR) and overexpression of the endogenous glycolate oxidase operon (glcDEFG_PP_3749) (Fig. 3). The new strain (RC038) showed efficient BHET to βKA conversion (Fig. 3 and Table 2); however, βKA repressed EG utilization. This highlights the necessity of improving simultaneous utilization of substrates for designing an effective bioconversion process (Werner et al. 2021).
This effort demonstrated the applicability of open-loop recycling systems (i.e., systems in which the output is exclusively controlled by the substrate intake). However, an additional carbon source, in the form of glucose, was necessary to support cell growth, making the process economically inconvenient. This limitation was overcome by additional engineering efforts on the strain RC038 (see above). The new strain could selectively grow on plastic-derived monomers generated via metal-promoted autoxidation (Sullivan et al. 2022). Specifically, the strain was modified by introducing the dcaAKIJP operon from Acinetobacter baylyi, which encodes enzymes for dicarboxylate uptake, and by deleting genes paaX and psrA, which encode the repressors of phenylacetate catabolism and β-oxidation, respectively (Fig. 3) (Ackermann et al. 2021). The resulting strain efficiently produced βKA (Table 2), growing on C4 to C10 polyolefins as sole carbon sources and, in nitrogen-depleted conditions, accumulated the biopolymers polyhydroxyalkanoates (PHA).
Similarly, an engineered strain of the carotenogenic bacteria R. jostii RHA1, which possesses the TPA and EG assimilation pathways, could efficiently utilize PET hydrolyzates (TPA and EG derived from PET alkaline hydrolysis) as its sole carbon sources (Diao et al. 2023). Heterologous expression of genes responsible for the key steps of the methylerythritol phosphate pathway (MEP)—namely 1-deoxy-D-xylulose-5-phosphate synthase (DXS) and isopentenyl pyrophosphate isomerase (IDI), sourced from Corynebacterium glutamicum, ensured increasing metabolic flux towards the production of the tetraterpenoid lycopene. This approach resulted in a remarkable 348-fold increase in production compared to that of the wild-type strain (Fig. 3, Table 2) (Diao et al. 2023).
While these PET-recycling microbial systems still require further optimization to reach economic feasibility, these examples validate the convenience of coupling chemical and biological catalysis to generate industrially relevant products from plastic materials.
4.3.2 Whole-cell microbial biocatalysts for PET degradation and valorization
As described above, enzyme engineering strategies enable the generation of new PET-degrading enzymatic variants with improved catalytic activities. In turn, metabolic engineering enables microorganisms to produce and secrete them extracellularly (see above), and eventually utilize their catalytic product for growth. That is, the generation of microbe-based biological systems to simultaneously degrade PET substrates, by producing and secreting PET hydrolases, and growing on TPA and EG, would offer an economic and self-sustaining solution toward PET upcycling.
Two attempts to create a whole-cell biocatalysts using P. putida KT2440 as a microbial host were recently reported. In the first study, P. putida KT2440 was adaptively evolved to use TPA and EG as sole carbon sources. The authors also screened and identified a 30-mer signal peptide allowing efficient secretion of the HiC cutinase (Fig. 1b). The resulting synthetic strain, however, showed growth defects when using PET substrates as a sole carbon source. Furthermore, the degrading activity of the secreted HiC cutinases was not effective enough to break down PET into monomers (Brandenberg et al. 2022). Similarly, in a second study, P. putida KT2440 was engineered for simultaneously synthesizing and secreting the LCC cutinase (Fig. 1c) (Liu et al. 2022a). PET degradation was partially achieved by the enzymatic activity of the secreted LCC. The strain was constructed by genomically integrating EG and TPA assimilation pathways (see above) and the catA gene (see above), while expressing the LCC cutinase gene from a plasmid. The new microbial strain utilized its own-made PET hydrolyzates for producing muconic acid (MA). However, the rate-limiting secretion of LCC and its poor catalytic activity at the bacterial growth temperature impaired the overall efficiency. The poor performances of these two engineered strains may be related to the modest depolymerization activity of the secreted PET hydrolases. The consequent low availability of carbon sources in the form of PET-derived monomers likely reduces the activity of central carbon metabolism for biomass and energy production. These issues may eventually be overcome by further evolving PET hydrolases together with the bacterial secretion apparatus and/or by utilizing engineered microbial consortia. To this end, the yeast Y. lipolytica carrying the IsPETase gene and the TPA-degrading bacterium P. stutzeri expressing a three-gene operon for polyhydroxybutyrate (PHB) biosynthesis from Ralstonia eutropha (Fig. 3), were efficiently co-cultured on PET powder for PHB production (Table 2) (Liu et al. 2021). Extracellular secretion of IsPETase was directed by the endogenous SPLIP2, and its activity was backed-up by endogenous lipases which could degrade MHET into TPA and EG (Fig. 1a). The artificial microflora maintained high activity in the co-cultivation system; however, the low activity of IsPETase and the low efficiency of the PHB pathway limited the accumulation of PHB (Table 2).
Another self-sustained microbial PET-degrading system was attempted by making use of a four-species consortium. It consisted of two strains of B. subtilis carrying IsPETase or IsMHETase fused to an endogenous SP, the TPA-degrading soil bacteria Rhodococcus jostii, and the EG utilizer P. putida KT2440. After optimization of carbon sources, temperature, and inoculation ratio, the consortia reached stability and achieved a depolymerization rate of amorphous PET film of 23.2% within seven days (Qi et al. 2021).
Although the whole-cell consortium approach seems to offer a more sustainable and integrated system for PET degradation and upcycling, harnessing the contribution of each single microorganism needs careful optimization. This may require complex metabolic engineering designs to improve, beside the efficiency of the individual strain, microorganism co-cultivation. The potential of generating a system based on engineered microbes to bio-upcycle PET is enormous and ecosystem engineering could allow synchronizing and controlling microbial metabolic activities. Nevertheless, the application of these systems strictly relies on the primary activity of PET hydrolases whose activity parameters need to match those for optimal bacterial growth.
The chemo-microbial approaches stand as the most valuable approaches so far developed for achieving complete PET upcycling. Chemical PET depolymerization continues to demonstrate superior performance in comparison to the enzymatic or whole-cell microbial PET degradation processes (Uekert et al. 2023). The biochemical routes for degrading and growing on plastic substrates mostly reside on non-conventional microorganisms; thus, harnessing the metabolic diversity of non-model microbial species would be critical for future developments (Wilkes et al. 2023). However, rational strain engineering necessitates a profound comprehension of the regulatory mechanisms governing substrate breakdown, carbon flow within the central carbon metabolism, and energy utilization of each microbe, either used as target for engineering or simple component of a consortium (Calero and Nikel 2019). Sequencing and characterization of new genomes can facilitate the groundwork for comprehensive systems-level biological analysis in non-model microorganisms. Furthermore, the integration of multi-omics data, like transcriptomics, metabolomics and proteomics will provide invaluable insights in promoting understanding of microbial behavior of still untapped microorganisms and will define a framework of guiding principles to develop high-throughput strain engineering platforms for PET degradation and valorization.
5 Conclusions
This review covers the current state of PET enzymatic degradation technology and some microbial-based valorization systems, including enzyme engineering strategies for generating improved PET-degrading enzyme variants, and several metabolic engineering strategies applied for PET degradation and valorization via microorganisms. We anticipate that machine learning programs will greatly assist the future implementation of high-throughput screening of enzymes and the design of synthetic metabolic pathways for the rational engineering of efficient microbial biocatalysts to perform PET upcycling. The utilization of synthetic microbial biocatalysts or hybrid biochemical methods for PET upcycling holds great promise as a sustainable and large-scale solution for managing PET waste. The applications of innovative synthetic microbiology and metabolic engineering approaches would support the scaling up of microbial biocatalysts from laboratory bioreactors to enable PET upcycling at a commercial level. This advancement would foster a circular material economy and play a significant role in safeguarding the environment from the negative impacts of PET waste.
Abbreviations
- AP:
-
Anchor peptide
- BHET:
-
Bis(2-hydroxyethyl) terephthalate
- CBM:
-
Carbohydrate binding modules
- Cry-PET:
-
Semicrystalline PET
- EG:
-
Ethylene glycol
- HBM:
-
Hydrophobic binding modules
- MHET:
-
Mono-(2-hydroxyethyl) terephthalate
- PC-PET:
-
Post-consumer PET
- PET:
-
Polyethylene terephthalate
- SDSP:
-
Sec-dependent signal peptides
- SP:
-
Signal peptide
- Topt :
-
Optimal temperatures
- Tm :
-
Denaturation midpoint
- TPA:
-
Terephthalate
References
Ackermann YS, Li W-J, Op De Hipt L, Niehoff P-J, Casey W, Polen T, Köbbing S, Ballerstedt H, Wynands B, O’connor K, Blank LM, Wierckx N (2021) Engineering adipic acid metabolism in Pseudomonas putida. Metab Eng 67:29–40
Aer L, Jiang Q, Gul I, Qi Z, Feng J, Tang L (2022) Overexpression and kinetic analysis of Ideonella sakaiensis PETase for polyethylene terephthalate (PET) degradation. Environ Res 212:113472
Anandharaj M, Lin Y-J, Rani RP, Nadendla EK, Ho M-C, Huang C-C, Cheng J-F, Chang J-J, Li W-H (2020) Constructing a yeast to express the largest cellulosome complex on the cell surface. Proc Natl Acad Sci 117(5):2385–2394
Antelava A, Damilos S, Hafeez S, Manos G, Al-Salem SM, Sharma BK, Kohli K, Constantinou A (2019) Plastic solid waste (PSW) in the context of life cycle assessment (LCA) and sustainable management. Environ Manag 64(2):230–244
Bååth JA, Borch K, Jensen K, Brask J, Westh P (2021) Comparative biochemistry of four polyester (PET) hydrolases**. ChemBioChem 22(9):1627–1637
Baek M, Anishchenko I, Humphreys IR, Cong Q, Baker D, Dimaio F (2023) Efficient and accurate prediction of protein structure using RoseTTAFold2. bioRxiv: 2023.2005.2024.54217
Bateman A (2019) UniProt: a worldwide hub of protein knowledge. Nucl Acids Res 47:D506–D515
Bayry J, Aimanianda V, Guijarro JI, Sunde M, Latgé JP (2012) Hydrophobins–unique fungal proteins. PLoS Pathog 8(5):e1002700
Beal J, Rogers M (2020) Levels of autonomy in synthetic biology engineering. Mol Syst Biol 16(12):e10019
Bell EL, Finnigan W, France SP, Green AP, Hayes MA, Hepworth LJ, Lovelock SL, Niikura H, Osuna S, Romero E, Ryan KS, Turner NJ, Flitsch SL (2021) Biocatal Nat Rev Methods Primers 1(1):46
Bell EL, Smithson R, Kilbride S, Foster J, Hardy FJ, Ramachandran S, Tedstone AA, Haigh SJ, Garforth AA, Day PJR, Levy C, Shaver MP, Green AP (2022) Directed evolution of an efficient and thermostable PET depolymerase. Nat Catal 5(8):673–681
Benavides Fernández CD, Guzmán Castillo MP, Quijano Pérez SA, Carvajal Rodríguez LV (2022) Microbial degradation of polyethylene terephthalate: a systematic review. SN Appl Sci 4(10):263
Bhak G, Méndez-Ardoy A, Escobedo A, Salvatella X, Montenegro J (2020) An adhesive peptide from the C-terminal domain of α-synuclein for single-layer adsorption of nanoparticles onto substrates. Bioconjug Chem 31(12):2759–2766
Billig S, Oeser T, Birkemeyer C, Zimmermann W (2010) Hydrolysis of cyclic poly (ethylene terephthalate) trimers by a carboxylesterase from thermobifida fusca KW3. Appl Microbiol Biotechnol 87(5):1753–1764
Blank LM, Narancic T, Mampel J, Tiso T, O’connor K (2020) Biotechnological upcycling of plastic waste and other non-conventional feedstocks in a circular economy. Curr Opin Biotechnol 62:212–219
Blázquez-Sánchez P, Engelberger F, Cifuentes-Anticevic J, Sonnendecker C, Griñén A, Reyes J, Díez B, Guixé V, Richter PK, Zimmermann W, Ramírez-Sarmiento CA (2022) Antarctic polyester hydrolases degrade aliphatic and aromatic polyesters at moderate temperatures. Appl Environ Microbiol 88(1):01842–11821
Boots B, Russell CW, Green DS (2019) Effects of microplastics in soil ecosystems: above and below ground. Environ Sci Technol 53(19):11496–11506
Brandenberg OF, Schubert OT, Kruglyak L (2022) Towards synthetic PETtrophy: engineering pseudomonas putida for concurrent polyethylene terephthalate (PET) monomer metabolism and PET hydrolase expression. Microb Cell Fact 21(1):119
Büscher N, Sayoga GV, Rübsam K, Jakob F, Schwaneberg U, Kara S, Liese A (2019) Biocatalyst immobilization by anchor peptides on an additively manufacturable material. Org Process Res Dev 23(9):1852–1859
Calero P, Nikel PI (2019) Chasing bacterial chassis for metabolic engineering: a perspective review from classical to non-traditional microorganisms. Microb Biotechnol 12(1):98–124
Carniel A, Santos AG, Júnior LSC, De Castro AM, Coelho MMAZ (2023) Biotransformation of ethylene glycol to glycolic acid by yarrowia lipolytica: a route for poly (ethylene terephthalate) (PET) upcycling. Biotechnol J 18(6):2200521
Castilla A, Giordano SR, Irazoqui G (2022) Microbial extremozymes. Academic Press, Cambridge, pp 207–222
Castro AMD, Carniel A, Stahelin D, Chinelatto Junior LS, Honorato HDA, De Menezes SMC (2019) High-fold improvement of assorted post-consumer poly (ethylene terephthalate) (PET) packages hydrolysis using humicola insolens cutinase as a single biocatalyst. Process Biochem 81:85–91
Chain PSG, Denef VJ, Konstantinidis KT, Vergez LM, Agulló L, Reyes VL, Hauser L, Córdova M, Gómez L, González M, Land M, Lao V, Larimer F, Lipuma JJ, Mahenthiralingam E, Malfatti SA, Marx CJ, Parnell JJ, Ramette A, Richardson P, Seeger M, Smith D, Spilker T, Sul WJ, Tsoi TV, Ulrich LE, Zhulin IB, Tiedje JM (2006) Burkholderia xenovorans LB400 harbors a multi-replicon, 9.73-Mbp genome shaped for versatility. Proc Natl Acad Sci 103(42):15280–15287
Chandra P, Enespa SR, Arora PK (2020) Microbial lipases and their industrial applications: a comprehensive review. Microb Cell Fact 19(1):169
Chen C-C, Han X, Li X, Jiang P, Niu D, Ma L, Liu W, Li S, Qu Y, Hu H, Min J, Yang Y, Zhang L, Zeng W, Huang J-W, Dai L, Guo R-T (2021) General features to enhance enzymatic activity of poly(ethylene terephthalate) hydrolysis. Nat Catal 4(5):425–430
Chen Z, Duan R, Xiao Y, Wei Y, Zhang H, Sun X, Wang S, Cheng Y, Wang X, Tong S, Yao Y, Zhu C, Yang H, Wang Y, Wang Z (2022) Biodegradation of highly crystallized poly(ethylene terephthalate) through cell surface codisplay of bacterial PETase and hydrophobin. Nat Commun 13(1):7138
Chew SY, Chee WJY, Than LTL (2019) The glyoxylate cycle and alternative carbon metabolism as metabolic adaptation strategies of candida glabrata: perspectives from Candida albicans and Saccharomyces cerevisiae. J Biomed Sci 26(1):52
Child J, Willetts A (1978) Microbial metabolism of aliphatic glycols bacterial metabolism of ethylene glycol. Biochim Biophys Acta (BBA) Gen Subj 538(2):316–327
Choi KY, Kim D, Sul WJ, Chae J-C, Zylstra GJ, Kim YM, Kim E (2005) Molecular and biochemical analysis of phthalate and terephthalate degradation by Rhodococcus sp. strain DK17. FEMS Microbiol Lett 252(2):207–213
Correddu D, Di Nardo G, Gilardi G (2021) Self-sufficient class VII cytochromes P450: from full-length structure to synthetic biology applications. Trends Biotechnol 39(11):1184–1207
Cui L, Qiu Y, Liang Y, Du C, Dong W, Cheng C, He B (2021a) Excretory expression of ispetase in E. coli by an enhancer of signal peptides and enhanced PET hydrolysis. Int J Biol Macromol 188:568–575
Cui Y, Chen Y, Liu X, Dong S, Tian YE, Qiao Y, Mitra R, Han J, Li C, Han X, Liu W, Chen Q, Wei W, Wang X, Du W, Tang S, Xiang H, Liu H, Liang Y, Houk KN, Wu B (2021b) Computational redesign of a PETase for plastic biodegradation under ambient condition by the GRAPE strategy. ACS Catal 11(3):1340–1350
Da Costa AM, De Oliveira LVR, Vidal L, Nicaud J-M, De Castro AM, Coelho MAZ (2020) Poly(ethylene terephthalate) (PET) degradation by yarrowia lipolytica: investigations on cell growth, enzyme production and monomers consumption. Process Biochem 95:81–90
Danso D, Schmeisser C, Chow J, Zimmermann W, Wei R, Leggewie C, Li X, Hazen T, Streit WR (2018) New insights into the function and global distribution of polyethylene terephthalate (PET)-degrading bacteria and enzymes in marine and terrestrial metagenomes. Appl Environ Microbiol 84(8):e02773-e12717
DeLano WL (2002) Pymol: an open-source molecular graphics tool. CCP4 Newsl Protein Crystallogr 40(1):82–92
Deng B, Yue Y, Yang J, Yang M, Xing Q, Peng H, Wang F, Li M, Ma L, Zhai C (2023) Improving the activity and thermostability of PETase from Ideonella sakaiensis through modulating its post-translational glycan modification. Commun Biol 6(1):39
Diao J, Hu Y, Tian Y, Carr R, Moon TS (2023) Upcycling of poly (ethylene terephthalate) to produce high-value bio-products. Cell Rep 42(1):111908
Ding Z, Xu G, Miao R, Wu N, Zhang W, Yao B, Guan F, Huang H, Tian J (2023) Rational redesign of thermophilic PET hydrolase LCCICCG to enhance hydrolysis of high crystallinity polyethylene terephthalates. J Hazard Mater 453:131386
Dittrich J, Brethauer C, Goncharenko L, Bührmann J, Zeisler-Diehl V, Pariyar S, Jakob F, Kurkina T, Schreiber L, Schwaneberg U, Gohlke H (2022) Rational design yields molecular insights on leaf-binding of anchor peptides. ACS Appl Mater Interfaces 14(25):28412–28426
Eberl A, Heumann S, Brückner T, Araujo R, Cavaco-Paulo A, Kaufmann F, Kroutil W, Guebitz GM (2009) Enzymatic surface hydrolysis of poly(ethylene terephthalate) and bis(benzoyloxyethyl) terephthalate by lipase and cutinase in the presence of surface active molecules. J Biotechnol 143(3):207–212
Eiamthong B, Meesawat P, Wongsatit T, Jitdee J, Sangsri R, Patchsung M, Aphicho K, Suraritdechachai S, Huguenin-Dezot N, Tang S, Suginta W, Paosawatyanyong B, Babu MM, Chin JW, Pakotiprapha D, Bhanthumnavin W, Uttamapinant C (2022) Discovery and genetic code expansion of a polyethylene terephthalate (PET) hydrolase from the human saliva metagenome for the degradation and bio-functionalization of PET. Angew Chem Int Ed Engl 61(37):e202203061
Erickson E, Gado JE, Avilán L, Bratti F, Brizendine RK, Cox PA, Gill R, Graham R, Kim D-J, König G, Michener WE, Poudel S, Ramirez KJ, Shakespeare TJ, Zahn M, Boyd ES, Payne CM, Dubois JL, Pickford AR, Beckham GT, Mcgeehan JE (2022) Sourcing thermotolerant poly(ethylene terephthalate) hydrolase scaffolds from natural diversity. Nat Commun 13(1):7850
Frazão CJR, Wagner N, Rabe K, Walther T (2023) Construction of a synthetic metabolic pathway for biosynthesis of 2,4-dihydroxybutyric acid from ethylene glycol. Nat Commun 14(1):1931
Gaston LW, Stadtman ER (1963) Fermentation of ethylene glycol by Clostridium glycolicum, sp. n. J Bacteriol 85(2):356–362
Gautom T, Dheeman D, Levy C, Butterfield T, Alvarez Gonzalez G, Le Roy P, Caiger L, Fisher K, Johannissen L, Dixon N (2021) Structural basis of terephthalate recognition by solute binding protein TphC. Nat Commun 12(1):6244
Giangeri G, Morlino MS, De Bernardini N, Ji M, Bosaro M, Pirillo V, Antoniali P, Molla G, Raga R, Treu L, Campanaro S (2022) Preliminary investigation of microorganisms potentially involved in microplastics degradation using an integrated metagenomic and biochemical approach. Sci Total Environ 843:157017
Gluth A, Xu Z, Fifield LS, Yang B (2022) Advancing biological processing for valorization of plastic wastes. Renew Sustain Energy Rev 170:112966
Graham R, Erickson E, Brizendine RK, Salvachúa D, Michener WE, Li Y, Tan Z, Beckham GT, Mcgeehan JE, Pickford AR (2022) The role of binding modules in enzymatic poly (ethylene terephthalate) hydrolysis at high-solids loadings. Chem Catal 2(10):2644–2657
Grunwald P (2018) Immobilized biocatalysts. Catalysts 8(9):386
Guo D, Li M, Jiang M, Cong G, Liu Y, Wang C, Li X (2022) Enhanced extracellular production and characterization of sucrose isomerase in Bacillus subtilis with optimized signal peptides. Foods 11(16):2468
Guo W, Duan J, Shi Z, Yu X, Shao Z (2023a) Biodegradation of PET by the membrane-anchored PET esterase from the marine bacterium Rhodococcus pyridinivorans P23. Commun Biol 6(1):1090
Guo W, Shi Z, Yu X, et al (2023b) Biodegradation of PET plastic by a marine strain Rhodococcus pyridinivorans P23 with a membrane anchoring PET esterase in a biofilm model. PREPRINT (Version 1) available at Research Square
Han X, Liu W, Huang J-W, Ma J, Zheng Y, Ko T-P, Xu L, Cheng Y-S, Chen C-C, Guo R-T (2017) Structural insight into catalytic mechanism of PET hydrolase. Nat Commun 8(1):2106
Hartline CJ, Schmitz AC, Han Y, Zhang F (2021) Dynamic control in metabolic engineering: theories, tools, and applications. Metab Eng 63:126–140
He B, Chen W, Mi L, Wu S, Zhu Y, Chen S (2014) Strong secretory signal peptide enhancing small peptide motif and use thereof.
Herrero Acero E, Ribitsch D, Steinkellner G, Gruber K, Greimel K, Eiteljoerg I, Trotscha E, Wei R, Zimmermann W, Zinn M, Cavaco-Paulo A, Freddi G, Schwab H, Guebitz G (2011) Enzymatic surface hydrolysis of PET: effect of structural diversity on kinetic properties of cutinases from thermobifida. Macromolecules 44(12):4632–4640
Horvat M, Winkler M (2020) In vivo reduction of medium- to long-chain fatty acids by carboxylic acid reductase (CAR) enzymes: limitations and solutions. ChemCatChem 12(20):5076–5090
Huang X, Cao L, Qin Z, Li S, Kong W, Liu Y (2018) Tat-Independent secretion of polyethylene terephthalate hydrolase PETase in Bacillus subtilis 168 mediated by its native signal peptide. J Agric Food Chem 66(50):13217–13227
Hughes AM, Darby JF, Dodson EJ, Wilson SJ, Turkenburg JP, Thomas GH, Wilkinson AJ (2022) Peptide transport in Bacillus subtilis–structure and specificity in the extracellular solute binding proteins OppA and DppE. Microbiology. https://doi.org/10.1099/mic.0.001274
Islam S, Apitius L, Jakob F, Schwaneberg U (2019) Targeting microplastic particles in the void of diluted suspensions. Environ Int 123:428–435
Jehanno C, Alty JW, Roosen M, De Meester S, Dove AP, Chen EYX, Leibfarth FA, Sardon H (2022) Critical advances and future opportunities in upcycling commodity polymers. Nature 603(7903):803–814
Jemli S, Ayadi-Zouari D, Hlima HB, Bejar S (2016) Biocatalysts: application and engineering for industrial purposes. Crit Rev Biotechnol 36(2):246–258
Ji M, Giangeri G, Usman M, Liu C, Bosaro M, Sessa F, Canu P, Treu L, Campanaro S (2023a) An integrated metagenomic-pangenomic strategy revealed native microbes and magnetic biochar cooperation in plasticizer degradation. Chem Eng J 468:143589
Ji M, Giangeri G, Yu F, Sessa F, Liu C, Sang W, Canu P, Li F, Treu L, Campanaro S (2023b) An integrated metagenomic model to uncover the cooperation between microbes and magnetic biochar during microplastics degradation in paddy soil. J Hazard Mater 458:131950
Ji M, Xiao L, Usman M, Liu C, Sang W, Treu L, Campanaro S, Luo G, Zhang Y (2023c) Different microplastics in anaerobic paddy soils: altering methane emissions by influencing organic matter composition and microbial metabolic pathways. Chem Eng J 469:144003
Jia Y, Samak NA, Hao X, Chen Z, Wen Q, Xing J (2022) Hydrophobic cell surface display system of PETase as a sustainable biocatalyst for PET degradation. Front Microbiol 13:1005480
Johan UUM, Rahman RNZRA, Kamarudin NHA, Ali MSM (2021) An integrated overview of bacterial carboxylesterase: structure, function and biocatalytic applications. Colloids Surf B 205:111882
Joo S, Cho IJ, Seo H, Son HF, Sagong HY, Shin TJ, Choi SY, Lee SY, Kim KJ (2018) Structural insight into molecular mechanism of poly(ethylene terephthalate) degradation. Nat Commun 9(1):382
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596(7873):583–589
Kang MJ, Kim HT, Lee M-W, Kim K-A, Khang TU, Song HM, Park SJ, Joo JC, Cha HG (2020) A chemo-microbial hybrid process for the production of 2-pyrone-4,6-dicarboxylic acid as a promising bioplastic monomer from PET waste. Green Chem 22(11):3461–3469
Karunatillaka I, Jaroszewski L, Godzik A (2022) Novel putative polyethylene terephthalate (PET) plastic degrading enzymes from the environmental metagenome. Proteins 90(2):504–511
Khairul Anuar NFS, Huyop F, Ur-Rehman G, Abdullah F, Normi YM, Sabullah MK, Abdul Wahab R (2022) An overview into polyethylene terephthalate (PET) hydrolases and efforts in tailoring enzymes for improved plastic degradation. Int J Mol Sci 23(20):12644
Kim HT, Kim JK, Cha HG, Kang MJ, Lee HS, Khang TU, Yun EJ, Lee D-H, Song BK, Park SJ, Joo JC, Kim KH (2019) Biological valorization of poly(ethylene terephthalate) monomers for upcycling waste PET. ACS Sustain Chem Eng 7(24):19396–19406
Kim JW, Park S-B, Tran Q-G, Cho D-H, Choi D-Y, Lee YJ, Kim H-S (2020) Functional expression of polyethylene terephthalate-degrading enzyme (PETase) in green microalgae. Microb Cell Fact 19(1):97
Kim DH, Han DO, In Shim K, Kim JK, Pelton JG, Ryu MH, Joo JC, Han JW, Kim HT, Kim KH (2021) One-pot chemo-bioprocess of PET depolymerization and recycling enabled by a biocompatible catalyst Betaine. ACS Catal 11(7):3996–4008
Kincannon WM, Zahn M, Clare R, Lusty Beech J, Romberg A, Larson J, Bothner B, Beckham GT, Mcgeehan JE, Dubois JL (2022) Biochemical and structural characterization of an aromatic ring–hydroxylating dioxygenase for terephthalic acid catabolism. Proc Natl Acad Sci 119(13):e2121426119
Kleeberg I, Hetz C, Kroppenstedt RM, Müller RJ, Deckwer WD (1998) Biodegradation of aliphatic-aromatic copolyesters by Thermomonospora fusca and other thermophilic compost isolates. Appl Environ Microbiol 64(5):1731–1735
Knott BC, Erickson E, Allen MD, Gado JE, Graham R, Kearns FL, Pardo I, Topuzlu E, Anderson JJ, Austin HP, Dominick G, Johnson CW, Rorrer NA, Szostkiewicz CJ, Copié V, Payne CM, Woodcock HL, Donohoe BS, Beckham GT, Mcgeehan JE (2020) Characterization and engineering of a two-enzyme system for plastics depolymerization. Proc Natl Acad Sci 117(41):25476–25485
Ko H, Kang M, Kim M-J, Yi J, Kang J, Bae J-H, Sohn J-H, Sung BH (2021) A novel protein fusion partner, carbohydrate-binding module family 66, to enhance heterologous protein expression in Escherichia coli. Microb Cell Fact 20(1):232
Koelmans AA, Mohamed Nor NH, Hermsen E, Kooi M, Mintenig SM, De France J (2019) Microplastics in freshwaters and drinking water: critical review and assessment of data quality. Water Res 155:410–422
Korendovych IV (2018) Protein engineering: methods and protocols. Springer, New York, pp 15–23
Kosiorowska KE, Moreno AD, Iglesias R, Leluk K, Mirończuk AM (2022) Production of PETase by engineered yarrowia lipolytica for efficient poly(ethylene terephthalate) biodegradation. Sci Total Environ 846:157358
Kunjapur AM, Tarasova Y, Prather KLJ (2014) Synthesis and accumulation of aromatic aldehydes in an engineered strain of Escherichia coli. J Am Chem Soc 136(33):11644–11654
Lazarevic D, Aoustin E, Buclet N, Brandt N (2010) Plastic waste management in the context of a European recycling society: comparing results and uncertainties in a life cycle perspective. Resour Conserv Recycl 55(2):246–259
Lee S, Lee YR, Kim SJ, Lee J-S, Min K (2023) Recent advances and challenges in the biotechnological upcycling of plastic wastes for constructing a circular bioeconomy. Chem Eng J 454:140470
Li W-J, Jayakody LN, Franden MA, Wehrmann M, Daun T, Hauer B, Blank LM, Beckham GT, Klebensberger J, Wierckx N (2019) Laboratory evolution reveals the metabolic and regulatory basis of ethylene glycol metabolism by Pseudomonas putida KT2440. Environ Microbiol 21(10):3669–3682
Li A, Sheng Y, Cui H, Wang M, Wu L, Song Y, Yang R, Li X, Huang H (2023) Discovery and mechanism-guided engineering of BHET hydrolases for improved PET recycling and upcycling. Nat Commun 14(1):4169
Liang X, Zou H (2023) Biotechnological application of cutinase: a powerful tool in synthetic biology. SynBio 1(1):54–64
Liu P, Zhang T, Zheng Y, Li Q, Su T, Qi Q (2021) Potential one-step strategy for PET degradation and PHB biosynthesis through co-cultivation of two engineered microorganisms. Eng Microbiol 1:100003
Liu P, Zheng Y, Yuan Y, Zhang T, Li Q, Liang Q, Su T, Qi Q (2022a) Valorization of polyethylene terephthalate to muconic acid by engineering Pseudomonas putida. Int J Mol Sci 23(19):10997
Liu Z, Zhang Y, Wu J (2022b) Enhancement of PET biodegradation by anchor peptide-cutinase fusion protein. Enzyme Microb Technol 156:110004
Lu H, Diaz DJ, Czarnecki NJ, Zhu C, Kim W, Shroff R, Acosta DJ, Alexander BR, Cole HO, Zhang Y, Lynd NA, Ellington AD, Alper HS (2022) Machine learning-aided engineering of hydrolases for PET depolymerization. Nature 604(7907):662–667
Ma Y, Zhang N, Vernet G, Kara S (2022) Design of fusion enzymes for biocatalytic applications in aqueous and non-aqueous media. Front Bioeng Biotechnol 10:944226
Macek B, Forchhammer K, Hardouin J, Weber-Ban E, Grangeasse C, Mijakovic I (2019) Protein post-translational modifications in bacteria. Nat Rev Microbiol 17(11):651–664
Macindoe I, Kwan AH, Ren Q, Morris VK, Yang W, Mackay JP, Sunde M (2012) Self-assembly of functional, amphipathic amyloid monolayers by the fungal hydrophobin EAS. Proc Natl Acad Sci 109(14):E804–E811
Markel U, Essani KD, Besirlioglu V, Schiffels J, Streit WR, Schwaneberg U (2020) Advances in ultrahigh-throughput screening for directed enzyme evolution. Chem Soc Rev 49(1):233–262
Martin HG, Radivojevic T, Zucker J, Bouchard K, Sustarich J, Peisert S, Arnold D, Hillson N, Babnigg G, Marti JM, Mungall CJ, Beckham GT, Waldburger L, Carothers J, Sundaram S, Agarwal D, Simmons BA, Backman T, Banerjee D, Tanjore D, Ramakrishnan L, Singh A (2023) Perspectives for self-driving labs in synthetic biology. Curr Opin Biotechnol 79:102881
Mccaul N, Braakman I (2022) Hold the fold: how delayed folding aids protein secretion. EMBO J 41(23):e112787
Meyer-Cifuentes IE, Öztürk B (2021) Mle046 is a marine mesophilic MHETase-like enzyme. Front Microbiol 12:693985
Meys R, Frick F, Westhues S, Sternberg A, Klankermayer J, Bardow A (2020) Towards a circular economy for plastic packaging wastes–the environmental potential of chemical recycling. Resour Conserv Recycl 162:105010
Mohanan N, Montazer Z, Sharma PK, Levin DB (2020) Microbial and enzymatic degradation of synthetic plastics. Front Microbiol 11:580709
Monterrey DT, Ayuso-Fernández I, Oroz-Guinea I, García-Junceda E (2022) Design and biocatalytic applications of genetically fused multifunctional enzymes. Biotechnol Adv 60:108016
Moog D, Schmitt J, Senger J, Zarzycki J, Rexer K-H, Linne U, Erb T, Maier UG (2019) Using a marine microalga as a chassis for polyethylene terephthalate (PET) degradation. Microb Cell Fact 18(1):171
Mückschel B, Simon O, Klebensberger J, Graf N, Rosche B, Altenbuchner J, Pfannstiel J, Huber A, Hauer B (2012) Ethylene glycol metabolism by Pseudomonas putida. Appl Environ Microbiol 78(24):8531–8539
Mudondo J, Lee HS, Jeong Y, Kim TH, Kim S, Sung BH, Park SH, Park K, Cha HG, Yeon YJ, Kim HT (2023) Recent advances in the chemobiological upcycling of polyethylene terephthalate (pet) into value-added chemicals. J Microbiol Biotechnol 33(1):1–14
Müller R-J, Schrader H, Profe J, Dresler K, Deckwer W-D (2005) Enzymatic degradation of poly (ethylene terephthalate): rapid hydrolyse using a hydrolase from T. fusca. Macromol Rapid Commun 26(17):1400–1405
Narancic T, Salvador M, Hughes GM, Beagan N, Abdulmutalib U, Kenny ST, Wu H, Saccomanno M, Um J, O’connor KE, Jiménez JI (2021) Genome analysis of the metabolically versatile Pseudomonas umsongensis GO16: the genetic basis for PET monomer upcycling into polyhydroxyalkanoates. Microb Biotechnol 14(6):2463–2480
Nechwatal A, Blokesch A, Nicolai M, Krieg M, Kolbe A, Wolf M, Gerhardt M (2006) A contribution to the investigation of enzyme-catalysed hydrolysis of poly (ethylene terephthalate) oligomers. Macromol Mater Eng 291(12):1486–1494
Nguyen PQ, Botyanszki Z, Tay PKR, Joshi NS (2014) Programmable biofilm-based materials from engineered curli nanofibres. Nat Commun 5(1):4945
Nikolaivits E, Taxeidis G, Gkountela C, Vouyiouka S, Maslak V, Nikodinovic-Runic J, Topakas E (2022) A polyesterase from the Antarctic bacterium Moraxella sp. degrades highly crystalline synthetic polymers. J Hazard Mater 434:128900
Oda M, Yamagami Y, Inaba S, Oida T, Yamamoto M, Kitajima S, Kawai F (2018) Enzymatic hydrolysis of PET: functional roles of three Ca2+ ions bound to a cutinase-like enzyme, cut190*, and its engineering for improved activity. Appl Microbiol Biotechnol 102(23):10067–10077
OECD (2022) Global Plastics Outlook.
Palm GJ, Reisky L, Böttcher D, Müller H, Michels EAP, Walczak MC, Berndt L, Weiss MS, Bornscheuer UT, Weber G (2019) Structure of the plastic-degrading Ideonella sakaiensis MHETase bound to a substrate. Nat Commun 10(1):1717
Panda S, Zhou JFJ, Feigis M, Harrison E, Ma X, Fung Kin Yuen V, Mahadevan R, Zhou K (2023) Engineering Escherichia coli to produce aromatic chemicals from ethylene glycol. Metab Eng 79:38–48
Pandit AV, Srinivasan S, Mahadevan R (2017) Redesigning metabolism based on orthogonality principles. Nat Commun 8(1):15188
Pandit AV, Harrison E, Mahadevan R (2021) Engineering Escherichia coli for the utilization of ethylene glycol. Microb Cell Fact 20(1):22
Patrauchan MA, Florizone C, Dosanjh M, Mohn WW, Davies J, Eltis LD (2005) Catabolism of benzoate and phthalate in Rhodococcus sp. strain RHA1: redundancies and convergence. J Bacteriol 187(12):4050–4063
Perez-Garcia P, Chow J, Costanzi E, Gurschke M, Dittrich J, Dierkes RF, Molitor R, Applegate V, Feuerriegel G, Tete P, Danso D, Thies S, Schumacher J, Pfleger C, Jaeger K-E, Gohlke H, Smits SHJ, Schmitz RA, Streit WR (2023) An archaeal lid-containing feruloyl esterase degrades polyethylene terephthalate. Commun Chem 6(1):193
Pirillo V, Orlando M, Battaglia C, Pollegioni L, Molla G (2022a) Efficient polyethylene terephthalate degradation at moderate temperature: a protein engineering study of LC-cutinase highlights the key role of residue 243. FEBS J 290:3185–3202
Pirillo V, Orlando M, Tessaro D, Pollegioni L, Molla G (2022b) An efficient protein evolution workflow for the improvement of bacterial PET hydrolyzing enzymes. Int J Mol Sci 23(1):264
Prabakaran S, Lippens G, Steen H, Gunawardena J (2012) Post-translational modification: nature’s escape from genetic imprisonment and the basis for dynamic information encoding. Wires Syst Biol Med 4(6):565–583
Puspitasari N, Tsai S-L, Lee C-K (2021) Fungal hydrophobin RolA Enhanced PETase hydrolysis of polyethylene terephthalate. Appl Biochem Biotechnol 193(5):1284–1295
Qi X, Ma Y, Chang H, Li B, Ding M, Yuan Y (2021) Evaluation of PET degradation using artificial microbial consortia. Front Microbiol. https://doi.org/10.3389/fmicb.2021.778828
Qiu X-Y, Xie S-S, Min L, Wu X-M, Zhu L-Y, Zhu L (2018) Spatial organization of enzymes to enhance synthetic pathways in microbial chassis: a systematic review. Microb Cell Fact 17(1):120
Rahimi A, García JM (2017) Chemical recycling of waste plastics for new materials production. Nat Rev Chem 1(6):0046
Ramazi S, Zahiri J (2021) Posttranslational modifications in proteins: resources, tools and prediction methods. Database (oxford). https://doi.org/10.1093/database/baab012
Rapp JT, Bremer BJ, Romero PA (2024) Self-driving laboratories to autonomously navigate the protein fitness landscape. Nat Chem Eng 1(1):97–107
Redondo-Hasselerharm PE, Gort G, Peeters ETHM, Koelmans AA (2020) Nano- and microplastics affect the composition of freshwater benthic communities in the long term. Sci Adv 6(5):eaay4054
Rennison AP, Westh P, Møller MS (2023) Protein-plastic interactions: the driving forces behind the high affinity of a carbohydrate-binding module for polyethylene terephthalate. Sci Total Environ 870:161948
Ribitsch D, Herrero Acero E, Greimel K, Dellacher A, Zitzenbacher S, Marold A, Rodriguez RD, Steinkellner G, Gruber K, Schwab H, Guebitz GM (2012) A new esterase from thermobifida halotolerans hydrolyses polyethylene terephthalate (PET) and polylactic acid (PLA). Polymers 4(1):617–629
Ribitsch D, Yebra AO, Zitzenbacher S, Wu J, Nowitsch S, Steinkellner G, Greimel K, Doliska A, Oberdorfer G, Gruber CC, Gruber K, Schwab H, Stana-Kleinschek K, Acero EH, Guebitz GM (2013) Fusion of binding domains to thermobifida cellulosilytica cutinase to tune sorption characteristics and enhancing PET hydrolysis. Biomacromol 14(6):1769–1776
Ribitsch D, Acero EH, Przylucka A, Zitzenbacher S, Marold A, Gamerith C, Tscheließnig R, Jungbauer A, Rennhofer H, Lichtenegger H, Amenitsch H, Bonazza K, Kubicek CP, Druzhinina IS, Guebitz GM (2015a) Enhanced cutinase-catalyzed hydrolysis of polyethylene terephthalate by covalent fusion to hydrophobins. Appl Environ Microbiol 81(11):3586–3592
Ribitsch D, Herrero Acero E, Przylucka A, Zitzenbacher S, Marold A, Gamerith C, Tscheließnig R, Jungbauer A, Rennhofer H, Lichtenegger H, Amenitsch H, Bonazza K, Kubicek CP, Druzhinina IS, Guebitz GM (2015b) Enhanced cutinase-catalyzed hydrolysis of polyethylene terephthalate by covalent fusion to hydrophobins. Appl Environ Microbiol 81(11):3586–3592
Richter PK, Blázquez-Sánchez P, Zhao Z, Engelberger F, Wiebeler C, Künze G, Frank R, Krinke D, Frezzotti E, Lihanova Y, Falkenstein P, Matysik J, Zimmermann W, Sträter N, Sonnendecker C (2023) Structure and function of the metagenomic plastic-degrading polyester hydrolase PHL7 bound to its product. Nat Commun 14(1):1905
Ronkvist ÅM, Xie W, Lu W, Gross RA (2009) Cutinase-catalyzed hydrolysis of poly (ethylene terephthalate). Macromolecules 42(14):5128–5138
Sadler JC, Wallace S (2021) Microbial synthesis of vanillin from waste poly (ethylene terephthalate). Green Chem 23(13):4665–4672
Sagong H-Y, Seo H, Kim T, Son HF, Joo S, Lee SH, Kim S, Woo J-S, Hwang SY, Kim K-J (2020) Decomposition of the PET film by MHETase using Exo-PETase function. ACS Catal 10(8):4805–4812
Saleski TE, Chung MT, Carruthers DN, Khasbaatar A, Kurabayashi K, Lin XN (2021) Optimized gene expression from bacterial chromosome by high-throughput integration and screening. Sci Adv 7(7):eabe1767
Sasoh M, Masai E, Ishibashi S, Hara H, Kamimura N, Miyauchi K, Fukuda M (2006) Characterization of the terephthalate degradation genes of Comamonas sp. strain E6. Appl Environ Microbiol 72(3):1825–1832
Scheffen M, Marchal DG, Beneyton T, Schuller SK, Klose M, Diehl C, Lehmann J, Pfister P, Carrillo M, He H, Aslan S, Cortina NS, Claus P, Bollschweiler D, Baret J-C, Schuller JM, Zarzycki J, Bar-Even A, Erb TJ (2021) A new-to-nature carboxylation module to improve natural and synthetic CO2 fixation. Nat Catal 4(2):105–115
Seeley ME, Song B, Passie R, Hale RC (2020) Microplastics affect sedimentary microbial communities and nitrogen cycling. Nat Commun 11(1):2372
Seo H, Kim S, Son HF, Sagong H-Y, Joo S, Kim K-J (2019) Production of extracellular PETase from Ideonella sakaiensis using sec-dependent signal peptides in E. coli. Biochem Biophys Res Comm 508:250–255
Shi L, Liu H, Gao S, Weng Y, Zhu L (2021) Enhanced extracellular production of IsPETase in Escherichia coli via engineering of the pelB signal peptide. J Agric Food Chem 69(7):2245–2252
Shi L, Liu P, Tan Z, Zhao W, Gao J, Gu Q, Ma H, Liu H, Zhu L (2023) Complete depolymerization of PET wastes by an evolved PET hydrolase from directed evolution. Angew Chem Int Ed 62(14):e202218390
Shigematsu T, Yumihara K, Ueda Y, Morimura S, Kida K (2003) Purification and gene cloning of the oxygenase component of the terephthalate 1,2-Dioxygenase system from delftia tsuruhatensis strain T7. FEMS Microbiol Lett 220(2):255–260
Sinha V, Patel MR, Patel JV (2010) Pet waste management by chemical recycling: a review. J Polym Environ 18(1):8–25
Son HF, Cho IJ, Joo S, Seo H, Sagong H-Y, Choi SY, Lee SY, Kim K-J (2019) Rational protein engineering of thermo-stable PETase from Ideonella sakaiensis for highly efficient PET degradation. ACS Catal 9(4):3519–3526
Sonnendecker C, Oeser J, Richter PK, Hille P, Zhao Z, Fischer C, Lippold H, Blázquez-Sánchez P, Engelberger F, Ramírez-Sarmiento CA, Oeser T, Lihanova Y, Frank R, Jahnke H-G, Billig S, Abel B, Sträter N, Matysik J, Zimmermann W (2022) Low carbon footprint recycling of post-consumer PET plastic with a metagenomic polyester hydrolase. Chemsuschem 15(9):e202101062
Su L, Chen K, Bai S, Yu L, Sun Y (2022) Cutinase fused with C-terminal residues of α-synuclein improves polyethylene terephthalate degradation by enhancing the substrate binding. Biochem Eng J 188:108709
Sulaiman S, You D-J, Kanaya E, Koga Y, Kanaya S (2014) Crystal structure and thermodynamic and kinetic stability of metagenome-derived LC-cutinase. Biochem 53(11):1858–1869
Sullivan KP, Werner AZ, Ramirez KJ, Ellis LD, Bussard JR, Black BA, Brandner DG, Bratti F, Buss BL, Dong X, Haugen SJ, Ingraham MA, Konev MO, Michener WE, Miscall J, Pardo I, Woodworth SP, Guss AM, Román-Leshkov Y, Stahl SS, Beckham GT (2022) Mixed plastics waste valorization through tandem chemical oxidation and biological funneling. Science 378(6616):207–211
Świderek K, Velasco-Lozano S, Galmés MÀ, Olazabal I, Sardon H, López-Gallego F, Moliner V (2023) Mechanistic studies of a lipase unveil effect of pH on hydrolysis products of small PET modules. Nat Commun 14(1):3556
Thakur B, Singh J, Singh J, Angmo D, Vig AP (2023) Biodegradation of different types of microplastics: molecular mechanism and degradation efficiency. Sci Total Environ 877:162912
Tournier V, Topham CM, Gilles A, David B, Folgoas C, Moya-Leclair E, Kamionka E, Desrousseaux ML, Texier H, Gavalda S, Cot M, Guémard E, Dalibey M, Nomme J, Cioci G, Barbe S, Chateau M, André I, Duquesne S, Marty A (2020) An engineered PET depolymerase to break down and recycle plastic bottles. Nature 580(7802):216–219
Trifunović D, Schuchmann K, Müller V (2016) Ethylene glycol metabolism in the acetogen Acetobacterium woodii. J Bacteriol 198(7):1058–1065
Trudeau DL, Edlich-Muth C, Zarzycki J, Scheffen M, Goldsmith M, Khersonsky O, Avizemer Z, Fleishman SJ, Cotton CAR, Erb TJ, Tawfik DS, Bar-Even A (2018) Design and in vitro realization of carbon-conserving photorespiration. Proc Natl Acad Sci USA 115(49):E11455–E11464
Uekert T, Singh A, Desveaux JS, Ghosh T, Bhatt A, Yadav G, Afzal S, Walzberg J, Knauer KM, Nicholson SR, Beckham GT, Carpenter AC (2023) Technical, economic, and environmental comparison of closed-loop recycling technologies for common plastics. ACS Sustain Chem Eng 11(3):965–978
Vaaje-Kolstad G, Westereng B, Horn SJ, Liu Z, Zhai H, Sørlie M, Eijsink VGH (2010) An oxidative enzyme boosting the enzymatic conversion of recalcitrant polysaccharides. Science 330(6001):219–222
Van Geem KM (2023) Plastic waste recycling is gaining momentum. Science 381(6658):607–608
Vollmer I, Jenks MJF, Roelands MCP, White RJ, Van Harmelen T, De Wild P, Van Der Laan GP, Meirer F, Keurentjes JTF, Weckhuysen BM (2020) Beyond mechanical recycling: giving new life to plastic waste. Angew Chem Int Ed 59(36):15402–15423
Von Haugwitz G, Han X, Pfaff L, Li Q, Wei H, Gao J, Methling K, Ao Y, Brack Y, Mican J, Feiler CG, Weiss MS, Bednar D, Palm GJ, Lalk M, Lammers M, Damborsky J, Weber G, Liu W, Bornscheuer UT, Wei R (2022) structural insights into (tere)phthalate-ester hydrolysis by a carboxylesterase and its role in promoting PET depolymerization. ACS Catal 12(24):15259–15270
Wagner N, Wen L, Frazão CJR, Walther T (2023a) Next-generation feedstocks methanol and ethylene glycol and their potential in industrial biotechnology. Biotechnol Adv 69:108276
Wagner N, Bade F, Straube E, Rabe K, Frazão CJR, Walther T (2023b) In vivo implementation of a synthetic metabolic pathway for the carbon-conserving conversion of glycolaldehyde to acetyl-CoA. Front Bioeng Biotechnol 11:1125544
Walther T, Topham CM, Irague R, Auriol C, Baylac A, Cordier H, Dressaire C, Lozano-Huguet L, Tarrat N, Martineau N, Stodel M, Malbert Y, Maestracci M, Huet R, André I, Remaud-Siméon M, François JM (2017) Construction of a synthetic metabolic pathway for biosynthesis of the non-natural methionine precursor 2,4-dihydroxybutyric acid. Nat Commun 8(1):15828
Wang N, Guan F, Lv X, Han D, Zhang Y, Wu N, Xia X, Tian J (2020) Enhancing secretion of polyethylene terephthalate hydrolase PETase in Bacillus subtilis WB600 mediated by the SPamy signal peptide. Lett Appl Microbiol 71(3):235–241
Weber J, Petrović D, Strodel B, Smits SHJ, Kolkenbrock S, Leggewie C, Jaeger K-E (2019) Interaction of carbohydrate-binding modules with poly (ethylene terephthalate). Appl Microbiol Biotechnol 103(12):4801–4812
Wei R, Breite D, Song C, Gräsing D, Ploss T, Hille P, Schwerdtfeger R, Matysik J, Schulze A, Zimmermann W (2019) Biocatalytic degradation efficiency of postconsumer polyethylene terephthalate packaging determined by their polymer microstructures. Adv Sci 6(14):1900491
Wei R, Von Haugwitz G, Pfaff L, Mican J, Badenhorst CPS, Liu W, Weber G, Austin HP, Bednar D, Damborsky J, Bornscheuer UT (2022) Mechanism-based design of efficient PET hydrolases. ACS Catal 12(6):3382–3396
Werner AZ, Clare R, Mand TD, Pardo I, Ramirez KJ, Haugen SJ, Bratti F, Dexter GN, Elmore JR, Huenemann JD, Peabody GL, Johnson CW, Rorrer NA, Salvachúa D, Guss AM, Beckham GT (2021) Tandem chemical deconstruction and biological upcycling of poly(ethylene terephthalate) to β-ketoadipic acid by Pseudomonas putida KT2440. Metab Eng 67:250–261
Wilkes RA, Waldbauer J, Caroll A, Nieto-Domínguez M, Parker DJ, Zhang L, Guss AM, Aristilde L (2023) Complex regulation in a COMAMONAS platform for diverse aromatic carbon metabolism. Nat Chem Biol 19(5):651–662
Wu G, Yan Q, Jones JA, Tang YJ, Fong SS, Koffas MAG (2016) Metabolic burden: cornerstones in synthetic biology and metabolic engineering applications. Trends Biotechnol 34(8):652–664
Wu B, Cui Y, Chen Y, Sun J, Zhu T, Pang H, Li C, Geng W (2023) Deep learning-aided redesign of a hydrolase for near 100% PET depolymerization under industrially relevant conditions.
Xu A, Zhou J, Blank LM, Jiang M (2023) Future focuses of enzymatic plastic degradation. Trends Microbiol 31(7):668–671
Xue R, Chen Y, Rong H, Wei R, Cui Z, Zhou J, Dong W, Jiang M (2021) Fusion of chitin-binding domain from Chitinolyticbacter meiyuanensis SYBC-H1 to the leaf-branch compost cutinase for enhanced PET hydrolysis. Front Bioeng Biotechnol 9:762854
Yang KK, Wu Z, Arnold FH (2019) Machine-learning-guided directed evolution for protein engineering. Nat Methods 16(8):687–694
Yang P, Liu W, Chen Y, Gong A-D (2022) Engineering the glyoxylate cycle for chemical bioproduction. Front Bioeng Biotechnol 10:1066651
Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Kimura Y, Oda K (2016) A bacterium that degrades and assimilates poly(ethylene terephthalate). Science 351(6278):1196–1199
Zeng W, Li X, Yang Y, Min J, Huang J-W, Liu W, Niu D, Yang X, Han X, Zhang L, Dai L, Chen C-C, Guo R-T (2022) Substrate-binding mode of a thermophilic PET hydrolase and engineering the enzyme to enhance the hydrolytic efficacy. ACS Catal 12(5):3033–3040
Zhang Y, Wang L, Chen J, Wu J (2013) Enhanced activity toward PET by site-directed mutagenesis of Thermobifida fusca cutinase–CBM fusion protein. Carbohyd Polym 97(1):124–129
Zhang Y, Guan F, Xu G, Liu X, Zhang Y, Sun J, Yao B, Huang H, Wu N, Tian J (2022) A novel thermophilic chitinase directly mined from the marine metagenome using the deep learning tool preoptem. Bioresour Bioprocess 9(1):54
Zhou H, Liao X, Wang T, Du G, Chen J (2010) Enhanced l-phenylalanine biosynthesis by co-expression of pheAfbr and aroFwt. Biores Technol 101(11):4151–4156
Zhu B, Wang D, Wei N (2022a) Enzyme discovery and engineering for sustainable plastic recycling. Trends Biotechnol 40(1):22–37
Zhu B, Ye Q, Seo Y, Wei N (2022b) Enzymatic degradation of polyethylene terephthalate plastics by bacterial curli display PETase. Environ Sci Technol Lett 9(7):650–657
Zurier HS, Goddard JM (2023) A high-throughput expression and screening platform for applications-driven PETase engineering. Biotechnol Bioeng 120(4):1000–1014
Acknowledgements
This study was carried out within the MICS (Made in Italy—Circular and Sustainable) Extended Partnership and received funding from Next-GenerationEU (Italian PNRR—M4 C2, Invest 1.3—D.D. 1551.11-10-2022, PE00000004). This manuscript reflects only the authors' views and opinions, neither the European Union nor the European Commission can be considered responsible for them.
Funding
Open access funding provided by Università degli Studi di Padova within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
Alessandro Satta: conceptualization, formal analysis, writing—original draft, visualization, review and editing. Guido Zampieri: editing, Giovanni Loprete: editing, Stefano Campanaro: review and editing. Laura Treu: review and editing, funding acquisition. Elisabetta Bergantino: review and editing, funding acquisition.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Satta, A., Zampieri, G., Loprete, G. et al. Metabolic and enzymatic engineering strategies for polyethylene terephthalate degradation and valorization. Rev Environ Sci Biotechnol 23, 351–383 (2024). https://doi.org/10.1007/s11157-024-09688-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11157-024-09688-1