Abstract
In this study, a curdlan-producing bacterium was isolated from Cow pea soil and identified as Priestia megaterium based on 16 S rRNA sequencing. To identify the most suitable carbon and nitrogen sources for curdlan production, submerged fermentation studies with different sources was carried out. To enhance the curdlan yield, optimization by one-factor-at-a-time approach was conducted. The optimal fermentation media consisted of 15% (w/v) sucrose, 0.1% (w/v) urea, 0.1% (w/v) KH2PO4, 0.04% (w/v) MgSO4·7H2O, trace elements, initial pH of 7.0 with 10% (v/v) inoculum size and agitation speed of 180 rpm. Kinetics of growth, curdlan yield, sucrose and ammonia depletion were studied for a period of 168 h. Maximum curdlan yield (0.31 g/L) was achieved at 96 h of fermentation. At this point, the fermentation media had an optical density of 9.68, biomass concentration of 4.26 mg/mL, and viable count of 2.4 × 104 CFU/mL. Additionally, the maximum percentage consumption of sucrose and ammonia over 168 h of fermentation were 75 and 62.5%, respectively. Finally, the identity of biopolymer curdlan was validated through characterization techniques such as Fourier Transform Infrared Spectroscopy (FTIR), Scanning Electron Microscopy (SEM) and Thermogravimetric analysis (TGA). Some characteristic features of curdlan such as the β-1,3-linkage was depicted by the absorption band at 890 cm−1 in FTIR, flaky granules with irregularities as seen in SEM, and thermal degradation between 235 and 350 °C by TGA. To the best of our knowledge, this is the first report on curdlan production from Priestia megaterium.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Curdlan is a bacterial exopolysaccharide, a linear homopolysaccharide, with repeating glucose monomers bonded by β-1,3-glycosidic linkages [1]. The general chemical formula of curdlan is (C6H10O5)n [2]. In 1962, Harada and co-workers discovered a biopolymer which could curdle when heated, and later in 1966 they named it as curdlan [3]. Curdlan exhibits insolubility in water but demonstrates solubility in alkaline solutions due to the absence of hydrophilic side chains [4] It lacks taste, odour, or colour [5]. Curdlan has molecular weight between 5.3 × 104 and 2.0 × 106 Da and its degree of polymerization (DP) usually varies between 135 and 455 [6]. When curdlan is dissolved in an aqueous solution and heated, it can form two distinct types of gels depending on the temperature: the thermo-reversible or low-set gel (~ 55 °C), and the thermo-irreversible or high-set gel (~ 85 °C). This gel forming property of curdlan can be attributed to its structure which forms condensed rod-like triple helices under higher temperatures [6]. Curdlan production has been reported in certain bacteria such as Agrobacterium sp., Rhizobium sp., Bacillus sp., Pseudomonas sp., Cellulomonas sp [7,8,9,10,11]. The crdASC operon consists of four essential genes which are necessary for biosynthesis of curdlan. These genes are crdA, crdS, crdC and crdR. Of these, the crdR gene is responsible for the expression of the crdASC operon [7].
Curdlan is biodegradable and non-toxic towards humans and the environment. It is gaining attention in the food industry as well as the pharmaceutical industry [6]. It has been in use in Japan, Taiwan, and Korea since 1989. In 1996, the U.S. FDA (Food and Drug Administration) granted approval for its utilization in food industries. It was introduced in the U.S. market under the brand Pureglucan™ after FDA approval [12]. Because of its exemplary rheological properties, curdlan has been utilized as a thickening and gelling agent in food. It also renders stability to food by acting as a food stabilizer. Due to these properties, the texture of food also improves [13]. In Japan, noodles are made by using curdlan to modify texture [14]. Edible food packaging films have also been manufactured from a combination of xanthan and curdlan gums. This eliminates the use of plastic and other non-biodegradable packaging materials [15]. In the biomedical and pharmaceutical industry, curdlan and its modified forms have found application as carriers for drug-delivery due to their capacity to control and prolong drug release. Curdlan sulfate has successfully shown anti-AIDS activity [12]. Protein drug delivery has also been carried out with curdlan hydroxyethyl derivatives [16]. The aforementioned features of curdlan make it an ideal raw material for many sectors. Therefore, to meet the demand of these applications, multiple sources of curdlan or good availability of curdlan is required.
Hence, in this study, various sources were screened to obtain a curdlan-producing bacteria which was identified as Priesta megaterium. The organism can be easily isolated from soil, and it does not have very stringent growth requirements. It has other biotechnological applications as well which are outlined in the results section. Notably, this study represents the first documentation of curdlan production from Priestia megaterium. The bacterial production of curdlan was optimized by the ‘One-factor-at-a-time’ method to obtain the best process conditions. To elucidate the relationship between cell growth and curdlan production, fermentation kinetics were also studied extensively. Further, the structure, morphology, and thermal properties of curdlan were studied by different characterization techniques.
Materials and Methods
Isolation and Screening of Curdlan-Producing Bacteria
Curdlan-producing bacteria was isolated from the nodules of leguminous plants such as Mimosa pudica, Vigna unguiculata, Vigna unguiculata ssp. Sesquipedalis and root soils of different plants such as Vigna unguiculata, Parthenium, Solanum lycopersicum, Solanum melongena, Jasminum sambac, Cocos nucifera, Musa, Areca catechu, Syzygium sp., Solanum melongena, Artocarpus heterophyllus, and moist and dry soil from a tree park (Table 1).
Curdlan-producing bacteria from nodules were isolated as described elsewhere [17] In brief, nodules were washed with distilled water and surface sterilized with hydrogen peroxide (3%). This was followed by squashing the nodules between two clean, grease-free glass slides. The liquid released from the nodules was plated onto Nutrient agar, and incubated at 30 °C for a period ranging from 24 to 48 h.
Curdlan-producing bacteria from soil samples were obtained from a depth ranging between 10 and 15 cm beneath the ground surface level. 10 g of soil was added to 100 mL of distilled water and kept in a rotary shaker at 30 °C at 100 rpm for 15 min. 1 mL of the soil suspension was added to 9 mL of distilled water and serially diluted up to 10−5. 0.1 mL was taken from each diluted sample and plated on nutrient agar by the spread plate method [18]. On incubation at 30 °C for 24 h, colonies displaying distinct morphological characteristics were selected and plated on aniline blue agar medium. The isolated colonies, screened for curdlan-producing bacteria on Aniline blue medium (consisting of 10 g/L glucose, 5 g/L yeast extract, 0.05 g/L aniline blue, and 15 g/L agar at pH 7.0), were incubated at 30 °C for 5 days to allow the development of blue coloured colonies, and those showing a deep blue colour were selected for fermentation studies [19].
Screening of Curdlan-Producing Bacterial Isolates Using Submerged Fermentation
The aniline blue positive isolates were individually inoculated in the growth medium consisting of (g/L) sucrose 20, yeast extract 5, peptone 5 at pH 7.0. The isolates were incubated at 180 rpm for 18 h at 30 °C on a rotary shaker. After growth, 10% inoculum (v/v) was transferred to the fermentation medium consisting of (g/L) sucrose, 100; (NH4)2HPO4, 2.3; KH2PO4, 1.0; MgSO4·7H2O, 0.4; 1% trace elements solution (g/L): FeSO4·7H2O, 5; MnSO4·H2O, 2; CoCl2·6H2O, 1; ZnCl2, 1 in 0.1 N HCl [10]. The initial pH of the fermentation medium was 7.0. The temperature and agitation speed were kept constant at 30 °C and 180 rpm, respectively. Incubations were performed at 96 h.
Recovery of Biopolymer Curdlan
The fermented media was centrifuged at 10,000 rpm for 15 min at 4 °C and the resulting pellet was used for extraction of curdlan. The pellet, consisting of cell-bound curdlan, was solubilized in 2 N NaOH overnight in a rotary shaker at 30 °C. After solubilization, centrifugation was carried out again at 10,000 rpm for 15 min at 4 °C to separate the cells. 2 N HCl was added to the supernatant to precipitate out the curdlan. The precipitated curdlan was rinsed with distilled water and subsequently dried in hot air oven at 60 °C for 7 to 8 h [10, 20]. Curdlan yield was determined by weighing the dried precipitate.
Identification of Curdlan-Producing Bacterium
The bacterial isolate with the highest curdlan yield amongst the other isolates was identified using 16s rRNA sequencing from National Centre for Microbial Research (NCMR), Pune, India. The EzBioCloud database was used for generating the identification report. Sequence alignment was done using CLUSTAL W program. The phylogenetic tree was constructed using > 1200 bp long aligned sequences by the neighbour joining method using General Time Reversible (GTR) parameter with bootstrap replications of 100 in MEGA 11 software.
Screening of Suitable Carbon and Nitrogen Sources in the Fermentation Medium
Carbon sources namely glucose, sucrose, maltose, fructose, lactose at 10% were selected to study their effect on the yield of curdlan. Similarly, nitrogen sources such as diammonium hydrogen phosphate, urea, peptone, ammonium nitrate and ammonium sulphate at 0.2% were studied [19]. The other media constituents (% w/v) FeSO4·7H2O (5), MnSO4·H2O (2), CoCl2·6H2O (1), ZnCl2 (1) and process parameters such as pH, fermentation temperature, and agitation speed were kept constant at 7.0, 30 °C, and 170 rpm, respectively.
Optimization of Media and Culture Conditions for Curdlan Production by the ‘One-Factor-at-a-Time’ Approach
In the One-factor-at-a-time (OFAT) method of optimization, one factor is varied while the others are kept constant [19, 21]. The advantage of employing the OFAT approach is the ease and convenience by which experiments can be carried out. OFAT technique is the best tool to understand about the effect of media components on product yield [21] The parameters chosen for OFAT optimization were sucrose (5–30% w/v), urea (0.05–2.4% w/v), KH2PO4 (0.05–0.3% w/v), MgSO4·7H2O (0.02–0.16% w/v), pH (6–9), inoculum size (5–25%), agitation speed (140–240 rpm); whereas concentration of trace elements (% w/v): FeSO4·7H2O (5), MnSO4·H2O (2), CoCl2·6H2O (1), ZnCl2 (1) in 0.1 N HCl were kept constant.
Fermentation Kinetics of Curdlan Production
Fermentation kinetics of growth, curdlan yield, sucrose and ammonia utilization was studied at optimized conditions for the production of curdlan by P. megaterium. Fermentation characteristics of the organism were studied over a period of 168 h (7 days). Samples were collected at regular intervals for analysis.
Different kinetic parameters such as the biomass yield coefficient based on substrate utilization (Yxs), product yield coefficient based on substrate utilization (Yps), product yield coefficient based on biomass (Ypx), specific rate of substrate utilization (qs), and specific rate of product formation (qp) were evaluated as per the methods described by Doran [22]. The kinetics equations employed are given below:
Analytical Methods
Estimation of Cell Growth
Cell growth was quantified by measuring optical density at 600 nm using spectrophotometer [23], viable cell count (CFU/mL), and biomass concentration (mg/mL). In brief, viable cell count was determined by serial dilution followed by spread plate method. The plates with maximum of 300 colonies were selected and recorded in CFU/mL [24]. Biomass was determined using the dry weight method [25]. In brief, the biomass pellet was washed with distilled water and dried to constant weight. The difference between the final and initial weight was equal to the biomass weight.
Estimation of Curdlan Yield
Curdlan yield was quantified by solubilization of cell pellet using 2 N NaOH, and further neutralization was carried out by 2 N HCl. The curdlan precipitate was dried in hot air oven to a constant weight as described in the section ‘Recovery of biopolymer curdlan’.
Estimation of Sucrose and Ammonia Concentration
The sucrose concentration in the fermentation media was estimated by hydrolysing it with 3 N HCl and thereafter reducing sugars were determined by modified 3,5-dinitrosalicylic acid (DNSA) method. The detailed methodology is shown in Fig. 1 [10].
The ammonia concentration in the fermentation media was estimated by the Indophenol method where phenol and hypochlorite react with ammonia to form indophenol blue which can be detected by spectrophotometer at 630 nm as shown in Fig. 2 [26].
Statistical Analysis
Data are represented as the mean ± standard deviation (SD) of three replicates. Statistical analyses were carried out using Microsoft Excel 2016.
Characterization
[27]FTIR Analysis
FTIR analysis was conducted to investigate the existence of functional groups in the biopolymer. Shimadzu spectrometer 00254 via ATR probing was used for the analysis.
SEM Analysis
To analyse the morphology of curdlan, scanning electron microscopy was carried out in the Zeiss SEM EV018 instrument at 10 kV.
TGA Analysis
A thermogravimetric analyzer was utilized for studying the thermal degradation of curdlan. The Hitachi thermal analysis system STA7200 was used for this purpose. Nitrogen was used as the purge gas. The temperature range was from 30 to 550 °C with heating rate of 15 °C/min [27].
Results and Discussion
Isolation and Identification of the Curdlan-Producing Bacterium
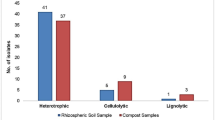
Curdlan production through microbial synthesis is mainly attributed to soil bacteria [2]. Curdlan forms a complex with aniline blue dye and hence is widely used for preliminary screening of curdlan-producing strains. The interaction of the aniline blue dye with the polymer were attributed to its concentration and its degree of polymerization [28]. A total of 52 bacterial strains were isolated on nutrient agar from different environmental sources as shown in Table 1. Of these, 35 isolates which showed blue colour were screened for curdlan production through submerged fermentation. Among the 35 isolates, the strain CPS1 isolated from Cow Pea (Vigna unguiculata) soil showed curdlan precipitation with yield of 0.14 g/L. This is the first report on the isolation of curdlan producing bacteria from cow pea soil. The strain CPS1 was found to be gram positive and spore forming bacilli based on morphology. Further, the strain CPS1 was identified by 16 S rRNA sequencing by amplification of > 1200 bp fragment. The closest match of CPS1 was found to be with Priestia megaterium with 100% homology, a finding verified by phylogenetic analysis as shown in Fig. 3. This confirmed the isolated organism to be Priestia megaterium with bootstrap value of 70. The nucleotide sequence was deposited to GenBank (Accession no. OL960210). The isolate Priestia megaterium was deposited in NCMR, India bearing the accession no. MCC 5044.
P. megaterium is known to have many applications in the biotechnology sector. It was identified to be a natural producer of Vitamin B12. It is also known to produce polyhydroxybutyrate (PHB), in the form of granules inside the cells. It is also involved in the production of recombinant proteins and as a growth-promoting bacteria in plants [29]. However, this study marks the first documentation of P. megaterium producing biopolymer curdlan.
Effect of Carbon and Nitrogen Sources on Curdlan Production
Curdlan production is significantly influenced by the carbon and nitrogen sources being utilized [19] The effect of different carbon sources (sucrose, glucose, maltose, fructose, lactose) on curdlan yield was evaluated. The yields of curdlan when sucrose, glucose, maltose, fructose, and lactose were used were 0.14, 0.13, 0.12, 0.09 and 0 g/L, respectively. Curdlan production was maximum at 0.14 g/L when sucrose was the carbon source. Hence, the order of curdlan yield with respect to carbon sources was sucrose > glucose > maltose > fructose > lactose (Fig. 4a). Since glucose is a monosaccharide, its consumption occurs at a much higher rate as opposed to sucrose. This leads to excess demand of glucose in the media which increases the cost of production [30]. Hence, similar to our study, numerous studies have also used sucrose as the substrate for curdlan production [16, 19, 30, 31].
The effect of different nitrogen sources (diammonium hydrogen phosphate, urea, peptone, ammonium nitrate and ammonium sulphate) on curdlan yield was evaluated. The curdlan yield when diammonium hydrogen phosphate, urea, peptone, ammonium nitrate and ammonium sulphate were used, were 0.15, 0.19, and 0 g/L for all the other three nitrogen sources, respectively. Curdlan yield was maximum when urea was the nitrogen source (0.19 g/L). There was no curdlan production when peptone, ammonium nitrate, and ammonium sulphate were used as nitrogen sources. This was due to very minimal growth of P. megaterium when these nitrogen sources were used in the media thereby affecting curdlan production. Hence, the order of curdlan yield with respect to nitrogen sources was Urea > diammonium hydrogen phosphate > peptone, ammonium nitrate, ammonium sulphate (Fig. 4b). P. megaterium being a ureolytic microorganism hydrolyses urea into ammonia and carbon dioxide for nitrogen source [32]. In comparison with other nitrogen sources, urea was selected as the most ideal nitrogen source in other research studies as well [16, 33].
It was observed that the curdlan yield was maximum at 0.20 g/L with sucrose and urea as the carbon and nitrogen source respectively. Therefore, sucrose and urea were selected as the optimal carbon and nitrogen sources in the fermentation media for P. megaterium.
Media Optimization for Curdlan Production by Priestia megaterium
To improve the production of curdlan, OFAT technique was employed for optimizing the media components such as sucrose, Urea, KH2PO4, MgSO4.7H2O and physical parameters such as pH, Inoculum size, Agitation speed.
Optimization of Sucrose
The carbon source is used for cell growth as well as synthesis of the biopolymer during fermentation [16]. In this study, carbon source sucrose concentration was varied from 5 to 30% (w/v). The optimum sucrose concentration was found to be 15% (w/v) with curdlan yield of 0.15 g/L (Fig. 5a). When sucrose concentration is lower, sugar consumption is higher during the exponential phase, but formation of biopolymer is relatively less. Conversely, at higher sucrose concentrations, more sugar is utilized for maintenance of the cell rather than contributing to biopolymer formation. This could be associated to balancing the osmotic pressure on the cell due to higher concentrations of sucrose [34]. It can probably be inferred that at 15% sucrose, the bacterium is able to balance the utilization of sucrose for cell growth as well as production of curdlan.
Optimization of Urea
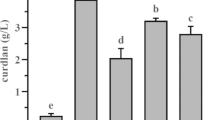
To determine the optimal urea concentration, 15% sucrose concentration was kept constant and urea concentration was varied from 0.05 to 2.4% (w/v). The maximum curdlan production was observed at 0.1% of urea with yield of 0.31 g/L thereby making it the optimum concentration (Fig. 5b). Nitrogen limiting condition being the primary requirement for curdlan production, an increase in curdlan yield is seen at these conditions. This is because at elevated levels of nitrogen, cellular growth is promoted which simultaneously limits the accumulation of curdlan [10]. Under nitrogen-limiting conditions, isoprenoid lipid in the curdlan biosynthetic pathway, readily synthesizes curdlan instead of lipopolysaccharides. Limiting levels of nitrogen curtails bacterial growth while simultaneously favouring production of secondary metabolites such as curdlan [20].
Optimization of KH2PO4
The concentration of KH2PO4 was varied from 0.05 to 0.3% (w/v) with sucrose (15% w/v) and urea (0.1% w/v) at their optimal levels. Curdlan production was maximum at 0.32 g/L at 0.1% of KH2PO4 (Fig. 5c). When curdlan is being produced under conditions of nitrogen scarcity, the phosphate concentration in the media is constant and does not promote cell growth. Instead, this residual phosphate is necessary to produce curdlan. Therefore, it can be inferred that nitrogen limitation and residual phosphate concentration are interdependent for curdlan production [35]. KH2PO4 provides the phosphate and potassium source in the media while simultaneously playing the role of buffering agent [36].
Optimization of MgSO4·7H2O
The concentration of MgSO4·7H2O was varied from 0.02 to 0.16% (w/v) with optimal levels of sucrose, urea, and KH2PO4 concentrations. The maximum curdlan yield was obtained at 0.04% of MgSO4·7H2O (Fig. 5d). The role of magnesium in secondary metabolite formation is not very clear. However, microbial cells have a requirement of Mg2+ ions for fulfilling metabolic needs. It affects the ribosomal activity and stability of the cell, protecting the cell from environmental stress [37]. Sulfur is a vital macronutrient required for structural and metabolic functions [36].
Optimization of pH
To study the effect of pH, the pH of the media was varied from 5.0 to 9.0. The maximum curdlan yield of 0.28 g/L was at pH 7.0 and hence was chosen as the optimal pH of the curdlan fermentation media (Fig. 6a). It was seen that neutral pH of 7.0 supports the growth of the bacteria however, when pH begins to decline it initiates curdlan production in the media after biomass is accumulated [38]. The findings of a study suggested that curdlan biosynthesis was favoured when the initial pH of media was 7.0 and dropped to 5.5 during fermentation [39]. Another study confirmed that an initial neutral pH of 7.0 was required for efficient cell growth followed by curdlan production occurring as the pH gradually decreases [40].
Optimization of Inoculum Size
Different inoculum sizes ranging from 5 to 25% (v/v) were utilized and the curdlan yield was determined. It was observed that the highest yield of 0.32 g/L was obtained at 10% (v/v) inoculum size and then the yield decreased gradually (Fig. 6b). Inoculum size influences the metabolite synthesis and activity [41]. Few other studies have also reported optimal curdlan yield at 10% (v/v) inoculum size [10, 16, 19].
Optimization of Agitation Speed
Agitation is an important factor which affects the production of curdlan as it facilitates aeration and may also lead to increased dissolved oxygen (DO) levels in the media [42]. Agitation speed of 180 rpm yielded 0.35 g/L curdlan which was the maximum curdlan yield (Fig. 6c). It was reported that static condition resulted in lesser biomass and product yield whereas agitation resulted in better curdlan yield. The aeration rate and agitation speed are crucial for maintaining the dissolved oxygen levels in the media. It was observed that the aeration rate had a less significant impact on oxygen transfer compared to the agitation speed. Curdlan production can be enhanced by higher agitation speeds. This is because a good oxygen supply benefits cell growth and curdlan production [42].
Bacillus sp. (Priestia sp. formerly known as Bacillus) using glucose and diammonium hydrogen phosphate as the carbon and nitrogen source, respectively [10].
Fermentation Kinetics
Studying the kinetics of microorganisms during fermentation provides valuable insights into understanding their growth and metabolism. The relationship between curdlan production and cell growth was evaluated by carrying out fermentation kinetic studies which included determination of cell growth parameters (OD600, viable count, biomass concentration); estimation of sucrose and ammonia concentration; determination of substrate and product yield factors [43]. Curdlan production was estimated at intervals of every 4 h and then phased out to every 12 h up to 168 h.
The optical density (OD) of the media gradually increased up to 60 h representing active growth of the microorganism. At the outset, exponential growth does not produce a substantial cell count, but after several generations, the cells accumulate leading to a significant increase in cell count. A decrease in OD is observed at 72 h followed by plateau up to 168 h indicating the stationary phase where growth of cells is balanced by the death of cells (Fig. 7). At this stage, as the substrate concentration diminishes, the growth rate will also correspondingly decrease [44]. In the case of viable count (CFU/mL), there was an increase up to 48 h followed by its decline. After 72 h, the organism stops actively dividing but the production of curdlan increases. The viable count was 26.9 × 104 CFU/mL at the beginning of fermentation but there was a decline seen around 60–72 h i.e., 3.1 × 104 CFU/mL. At the end of 168 h of fermentation, the viable count was as low as 0.21 × 104 CFU/mL, and corresponding biomass concentrations were 0.96 mg/mL, 5.76 mg/mL, and 2.94 mg/mL at 0 h, 72 h, and 168 h, respectively (Fig. 7). As the viable count reduced, curdlan production increased thereby proving that it is indeed a secondary metabolite produced during the stationary phase of microbial growth.
In this study, sucrose and urea were used as the carbon and nitrogen sources respectively. After growth, the organism focuses on using the unutilized sucrose in the media to produce curdlan. Urea is hydrolysed to ammonia and carbon dioxide by enzyme urease produced by the organism [45]. It was seen that as the organism grows and produces curdlan, the carbon and nitrogen in the media starts decreasing over time. A declining trend was observed in sucrose utilization, where the concentration of sucrose decreased from 70 mg/mL to 17.5 mg/mL over the course of 168 h of fermentation. Similarly, a decline was observed in ammonia concentration as well, where initial concentration of 0.8 mg/mL declined to 0.3 mg/mL over time, as depicted in Fig. 8. The percent consumption of sucrose concentration over the course of fermentation was 75% whereas that of ammonia was 62.5%. Based on the results, it is evident that when nitrogen levels reach limiting conditions in the media, cell growth reduces and curdlan production increases. This phenomenon could be attributed to isoprenoid lipid being readily available for synthesis of the exopolysaccharide, as opposed to lipopolysaccharide, under nitrogen-depleting conditions [10]. Hence, it can be concluded that the excess carbon source in the media is utilized when the nitrogen levels drop. This also aligns with a rising pattern of curdlan production which reaches maximum yield of 0.31 g/L at 96 h (Fig. 8). Similar results were also observed in other studies [10, 19, 46].
Over the course of fermentation, the kinetics of cell growth during the production of curdlan were studied. Biomass and product yield factors were determined as per Table 2. At 96 h, the curdlan production was maximum (0.31 g/L), the specific rate of substrate utilization (qs) and specific rate of product formation (qp) were found to be 11.88 and 0.126 h−1 respectively. The average specific rate of product formation (qp) was 0.0735 h−1. The maximum biomass concentration (Xm) was 5.76 mg/mL at 72 h which demonstrated that higher yield of curdlan at 96 h is probably due to a productive strain (high qp).
Characterization of Curdlan Produced by P. megaterium
Commercially available curdlan obtained from Fujifilm Wako (Japan) and curdlan obtained from P. megaterium were used for characterization. The FTIR spectra as shown in Fig. 9 showed some characteristic functional groups of curdlan. The absorption band near 890 cm−1 indicates the presence of characteristic β-1,3-glycosidic bond. The bend at around 3400 cm−1 indicated the O–H group. C–H stretching was shown by the band at 2900 cm−1 and the bend around 1650 cm−1 represented the C=O group. The band at around 1310 cm−1 depicts the C–H bond. This spectrum is consistent with the commercial curdlan sample as well as with the spectra reported by other studies [19, 20, 47].
Morphology of curdlan was determined by Scanning electron microscopy (SEM). Commercial curdlan granules appear invaginated or slightly collapsed (Fig. 10) whereas the curdlan produced by P. megaterium appears flaky with irregularities (Fig. 11). This can be attributed to the methods used for drying. Industries rely mainly on spray-drying whereas at the laboratory scale, lyophilization is commonly employed which exerts high pressure on the particles for drying [5].
The thermogravimetric analysis (TGA) curves are illustrated in Fig. 12. The decomposition profile consisted of two steps. Curdlan underwent dehydration in the first step which occurred around 230 °C. In the second phase, which was between 235 and 350 °C, there was a significant decline in the curve, which was attributed to the overall reduction in weight. This showed the thermal decomposition trend of curdlan. These results were consistent with the commercial curdlan curves and previously reported literature [48].
Conclusion
In this study, a curdlan-producing microorganism identified as Priestia megaterium was isolated from Cow pea soil. With respect to substrate utilization, sucrose exhibited best effect as the carbon source in conjunction with urea as the best nitrogen source. Optimization of curdlan’s fermentative production was carried out by the one-factor-at-a-time method. At optimal conditions, the yield of curdlan increased from 0.15 to 0.35 g/L, which is a 133% improvement. Fermentation kinetics of P. megaterium was also studied wherein it was observed that the specific rate of product formation and specific rate of substrate utilization at maximum curdlan yield (96 h) were 0.0735 and 11.88 h−1, respectively suggesting that curdlan production was due to highly productive strain. From the fermentation time profile and cell growth, it is evident that when the OD of the media remains constant and the viable count declines, production of curdlan begins in the stationary phase denoting that it is a secondary metabolite. This coincides with the depleting sucrose and nitrogen limiting conditions of the media. The kinetics data can be useful for further scaling up of the process. Curdlan extracted from P. megaterium was characterized by FTIR, SEM, and TGA. Further optimization using different strategies such as response surface methodology (RSM) and Artificial neural network (ANN) may help in increasing the yield of curdlan.
Data Availability
No datasets were generated or analysed during the current study.
References
Donot F, Fontana A, Baccou JC, Schorr-Galindo S (2012) Microbial exopolysaccharides: main examples of synthesis, excretion, genetics and extraction. Carbohydr Polym 87:951–962. https://doi.org/10.1016/j.carbpol.2011.08.083
Verma DK, Niamah AK, Patel AR et al (2020) Chemistry and microbial sources of curdlan with potential application and safety regulations as prebiotic in food and health. Food Res Int. https://doi.org/10.1016/j.foodres.2020.109136
Harada T (1977) Production, properties, and application of curdlan. Extracell Microb Polysaccharides 45:265–283
Miyoshi K, Uezu K, Sakurai K, Shinkai S (2004) Proposal of a new hydrogen-bonding form to maintain curdlan triple helix. Chem Biodivers 1:916–924. https://doi.org/10.1002/cbdv.200490073
Mangolim CS, Silva TT, Da Fenelon VC et al (2017) Description of recovery method used for curdlan produced by Agrobacterium sp. IFO 13140 and its relation to the morphology and physicochemical and technological properties of the polysaccharide. PLoS One 12:1–19. https://doi.org/10.1371/journal.pone.0171469
Zhang R, Edgar KJ (2014) Properties, chemistry, and applications of the bioactive polysaccharide curdlan. Biomacromolecules 15:1079–1096. https://doi.org/10.1021/bm500038g
Yu X, Zhang C, Yang L et al (2015) CrdR function in a curdlan-producing Agrobacterium sp. ATCC31749 strain. BMC Microbiol 15:1–10. https://doi.org/10.1186/s12866-015-0356-1
Mamaril JC, Paner ET, Palacpac ES (1988) The production of gel-forming polysaccharides by Rhizobium sp. and curdlan by a mutant cultured in coconut water. Trans Nat Acad Sci Tech 10:339–349
Ben Salah R, Jaouadi B, Bouaziz A et al (2011) Fermentation of date palm juice by curdlan gum production from Rhizobium radiobacter ATCC 6466™: purification, rheological and physico-chemical characterization. LWT Food Sci Technol 44:1026–1034. https://doi.org/10.1016/j.lwt.2010.11.023
Gummadi SN, Kumar K (2005) Production of extracellular water insoluble β-1,3-glucan (Curdlan) from Bacillus sp. SNC07. Biotechnol Bioprocess Eng 10:546–551. https://doi.org/10.1007/BF02932292
Kenyon WJ, Esch SW, Buller CS (2005) The curdlan-type exopolysaccharide produced by Cellulomonas flavigena KU forms part of an extracellular glycocalyx involved in cellulose degradation. Antonie Van Leeuwenhoek 87:143–148. https://doi.org/10.1007/s10482-004-2346-4
West TP (2020) Production of the polysaccharide curdlan by Agrobacterium species on processing coproducts and plant lignocellulosic hydrolysates. Fermentation 6:1–10. https://doi.org/10.3390/fermentation6010016
Martinez CO, Ruiz SP, Fenelon VC et al (2016) Characterization of curdlan produced by Agrobacterium sp. IFO 13140 cells immobilized in a loofa sponge matrix, and application of this biopolymer in the development of functional yogurt. J Sci Food Agric 96:2410–2417. https://doi.org/10.1002/jsfa.7357
Yotsuzuka F (2001) Curdlan. In: Cho SS, Dreher M (eds) Handbook of dietary fiber. Taylor & Francis, Milton Park, pp 734–754
Mohsin A, Zaman WQ, Guo M et al (2020) Xanthan-curdlan nexus for synthesizing edible food packaging films. Int J Biol Macromol 162:43–49. https://doi.org/10.1016/j.ijbiomac.2020.06.008
Shih IL, Yu JY, Hsieh C, Wu JY (2009) Production and characterization of curdlan by Agrobacterium Sp. Biochem Eng J 43:33–40. https://doi.org/10.1016/j.bej.2008.08.006
Marappa N, Dharumadurai D, Nooruddin T (2022) Isolation of Frankia from Casuarina Root Nodule. In: Amaresan N, Patel P, Amin D (eds) Practical handbook on agricultural microbiology. Springer, Berlin
Selvaraj S, Murty VR (2016) Process optimization for tannase production by Bacillus gottheilii M2S2 on inert polyurethane foam support. Biocatal Agric Biotechnol 7:48–55. https://doi.org/10.1016/j.bcab.2016.05.004
Yang M, Zhu Y, Li Y et al (2016) Production and optimization of curdlan produced by Pseudomonas sp. QL212. Int J Biol Macromol 89:25–34. https://doi.org/10.1016/j.ijbiomac.2016.04.027
Prakash S, Rajeswari K, Divya P et al (2018) Optimization and production of curdlan gum using Bacillus cereus PR3 isolated from rhizosphere of leguminous plant. Prep Biochem Biotechnol 48:408–418. https://doi.org/10.1080/10826068.2018.1451886
Singh V, Haque S, Niwas R et al (2017) Strategies for fermentation medium optimization: an in-depth review. Front Microbiol. https://doi.org/10.3389/fmicb.2016.02087
Doran PM (1995) Homogeneous reactions. Bioprocess Engineering principles, 1st edn. Academic press limited, pp 257–295
Zhang Q, Sun J, Wang Z et al (2018) Kinetic analysis of curdlan production by Alcaligenes faecalis with maltose, sucrose, glucose and fructose as carbon sources. Bioresour Technol 259:319–324. https://doi.org/10.1016/j.biortech.2018.03.059
Sanders ER (2012) Aseptic laboratory techniques: plating methods. J Vis Exp. https://doi.org/10.3791/3064
Selvaraj S, Natarajan K, Nowak A, Murty VR (2021) Mathematical modeling and simulation of newly isolated bacillus cereus M1GT for tannase production through semi-solid state fermentation with agriculture residue triphala. S Afr J Chem Eng 35:89–97. https://doi.org/10.1016/j.sajce.2020.10.001
Srienc F, Arnold B, Bailey JE (1984) Characterization of lntracellular accumuIation of poly-/3-H ydroxybutyrate (PHB) in individual cells of AIcaIigenes eutrophus HI6 by flow cytometry. Biotechnol Bioeng 26:982–987
Aquinas N, Bhat MR, Selvaraj S (2022) A review presenting production, characterization, and applications of biopolymer curdlan in food and pharmaceutical sectors. Polym Bull 79:6905–6927. https://doi.org/10.1007/s00289-021-03860-1
Nakanishi I, Kimura K, Suzuki T et al (1976) Demonstration of curdlan-type polysac-charide and some other β-1,3-glucan in microorganisms with aniline blue. J Gen Appl Microbiol 22:1–11. https://doi.org/10.2323/jgam.22.1
Biedendieck R, Knuuti T, Moore SJ, Jahn D (2021) The beauty in the beast-the multiple uses of Priestia megaterium in biotechnology. Appl Microbiol Biotechnol 105:5719–5737. https://doi.org/10.1007/s00253-021-11424-6/Published
Anane RF, Sun H, Zhao L et al (2017) Improved curdlan production with discarded bottom parts of Asparagus spear. Microb Cell Fact 16:1–8. https://doi.org/10.1186/s12934-017-0671-3
Mohsin A, Sun J, Khan IM et al (2019) Sustainable biosynthesis of curdlan from orange waste by using Alcaligenes faecalis: a systematically modeled approach. Carbohydr Polym 205:626–635. https://doi.org/10.1016/j.carbpol.2018.10.047
Mekonnen E, Kebede A, Nigussie A et al (2021) Isolation and characterization of urease-producing soil bacteria. Int J Microbiol. https://doi.org/10.1155/2021/8888641
Jiang L (2013) Effect of nitrogen source on curdlan production by Alcaligenes faecalis ATCC 31749. Int J Biol Macromol 52:218–220. https://doi.org/10.1016/j.ijbiomac.2012.10.010
Mohammad FHA, Badr-eldin SM, El-tayeb M, Abd El-Rahman OA (1995) Polysaccharide production by Aureobasidium pullulans III. The influence of initial sucrose concentration on batch kinetics. Biomass Bioenergy 8:121–129
Kim M-K, Lee I-Y, Lee J-H et al (2000) Residual phosphate concentration under nitrogen-limiting conditions regulates curdlan production in Agrobacterium species. J Indus Microb Biotech. https://doi.org/10.1038/sj.jim.7000053
Elhussiny NI, El-Refai HA, Mohamed SS et al (2023) Aspergillus flavus biomass catalytic lipid modification: optimization of cultivation conditions. Biomass Convers Biorefin. https://doi.org/10.1007/s13399-023-04396-2
Walker GM (1994) The roles of magnesium in biotechnology. Crit Rev Biotechnol 14:311–354. https://doi.org/10.3109/07388559409063643
Lee JH, Lee IY, Kim MK, Park YH (1999) Optimal pH control of batch processes for production of curdlan by Agrobacterium species. J Ind Microbiol Biotechnol 23:143–148. https://doi.org/10.1038/sj.jim.2900714
Zhan XB, Lin CC, Zhang HT (2012) Recent advances in curdlan biosynthesis, biotechnological production, and applications. Appl Microbiol Biotechnol 93:525–531. https://doi.org/10.1007/s00253-011-3740-2
Saudagar PS, Singhal RS (2004) Fermentative production of curdlan. Appl Biochem Biotechnol 118:21–31
Selvaraj S, Vytla RM (2018) Solid state fermentation of Bacillus gottheilii M2S2 in laboratory-scale packed bed reactor for tannase production. Prep Biochem Biotechnol 48:799–807. https://doi.org/10.1080/10826068.2018.1509086
Lee IY, Kim MK, Lee JH et al (1999) Influence of agitation speed on production of curdlan by Agrobacterium species. Bioprocess Eng 20:283–287
Zhu C, Fang B, Wang S (2016) Effects of culture conditions on the kinetic behavior of 1,3-propanediol fermentation by Clostridium butyricum with a kinetic model. Bioresour Technol 212:130–137. https://doi.org/10.1016/j.biortech.2016.04.028
Maier RM, Pepper IL (2015) Bacterial growth. Environmental microbiology, Third Edition. Elsevier Inc., pp 37–56
Hailemariam S, Zhao S, He Y, Wang J (2021) Urea transport and hydrolysis in the rumen: a review. Anim Nutr 7:989–996. https://doi.org/10.1016/j.aninu.2021.07.002
El-Sayed MH, Arafat HH, Elsehemy IA, Basha M (2016) Optimization, purification and physicochemical characterization of curdlan produced by Paenibacillus sp. strain NBR-10. Biosci Biotechnol Res Asia 13:901–909. https://doi.org/10.13005/bbra/2113
Kalyanasundaram GT, Doble M, Gummadi SN (2012) Production and downstream processing of (1→3)-β- D-glucan from mutant strain of Agrobacterium sp. ATCC 31750. AMB Express 2:1–10. https://doi.org/10.1186/2191-0855-2-31
Bai W, Shah F, Wang Q, Liu H (2019) Dissolution, regeneration and characterization of curdlan in the ionic liquid 1-ethyl-3-methylimidazolium acetate. Int J Biol Macromol 130:922–927. https://doi.org/10.1016/j.ijbiomac.2019.01.223
Acknowledgements
The authors wish to thank Department of Biotechnology, Manipal Institute of Technology (MIT), Manipal Academy of Higher Education (MAHE), Manipal for their support in conducting this research. The authors would like to express their gratitude to Karnataka Vision Group on Science and Technology (VGST, GRD-773) for additional instrumentation facilities. The authors also wish to thank Dr. Anil Kumar N V, Professor, Department of Chemistry, Manipal Institute of Technology (MIT), Manipal Academy of Higher Education (MAHE) for his valuable inputs in characterization studies.
Funding
Open access funding provided by Manipal Academy of Higher Education, Manipal. Not applicable.
Author information
Authors and Affiliations
Contributions
NA: conceptualization, methodology, formal analysis, writing—original draft preparation. RMB: conceptualization, formal analysis, supervision, writing—review and editing. SS: conceptualization, formal analysis, supervision, writing—review and editing.
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interests.
Ethical Approval
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Part of this work has been filed for an Indian patent against patent application number 202341048058.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aquinas, N., Bhat, R.M. & Selvaraj, S. Submerged Fermentation and Kinetics of Newly Isolated Priestia megaterium for the Production of Biopolymer Curdlan. J Polym Environ (2024). https://doi.org/10.1007/s10924-024-03224-6
Accepted:
Published:
DOI: https://doi.org/10.1007/s10924-024-03224-6