Abstract
Methyl-TROSY based NMR experiments have over the last two decades become one of the most important means to characterise dynamics and functional mechanisms of large proteins and macromolecular machines in solution. The chemical shift assignment of methyl groups in large proteins is, however, still not trivial and it is typically performed using backbone-dependent experiments in a ‘divide and conquer’ approach, mutations, structure-based assignments or a combination of these. Structure-based assignment of methyl groups is an emerging strategy, which reduces the time and cost required as well as providing a method that is independent of a backbone assignment. One crucial step in available structure-based assignment protocols is linking the two prochiral methyl groups of leucine and valine residues. This has previously been achieved by recording NOESY spectra with short mixing times or by comparing NOESY spectra. Herein, we present a method based on through-bond scalar coupling transfers, a 3D-HMBC-HMQC experiment, to link the intra-residue methyl groups of leucine and valine. It is shown that the HMBC-HMQC method has several advantages over solely using NOESY spectra since a unique intra-residue cross-peak is observed. Moreover, overlap in the methyl-TROSY HMQC spectrum can easily be identified with the HMBC-HMQC experiment, thereby removing possible ambiguities in the assignment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The methyl-bearing residues leucine, isoleucine, and valine are often well dispersed throughout a protein and these residues therefore provide significant coverage of the structure. Using 13CH3 labelled methyl groups as probes within a uniformly deuterated protein has become an indispensable method to characterise large dynamic systems by Nuclear Magnetic Resonance (NMR) spectroscopy. The three-fold rotational axis in combination with sophisticated pulse sequences, which maintain the sensitivity enhancement afforded by the methyl-TROSY effect, have provided a suite of experiments that allow one to probe the structure and dynamics in a wide variety of systems (Tugarinov et al. 2003; Tugarinov and Kay 2003, 2004; Ruschak and Kay 2010; Hansen and Kay 2011; Hansen et al. 2010).
A central challenge faced is the assignment of resonances in the methyl-TROSY HMQC spectrum. A common method used to achieve this is a ‘divide and conquer’ approach where large systems are divided into stable subunits or domains and assigned using standard triple-resonance experiments (Salzmann et al. 2000; Tugarinov et al. 2002) Once the resonances are assigned in each subunit, the methyl resonance assignment is transferred to the full complex. Whilst this approach has proven very successful (Sprangers and Kay 2007; Rosenzweig et al. 2013; Gelis et al. 2007), it can only be applied in cases where the protein can be divided into smaller stable units that are amenable to standard triple-resonance experiments.
Recent structure-based chemical shift assignment methods have recast the problem of methyl resonance assignment as a graph matching problem (Pritišanac et al. 2017; Monneau et al. 2017; Xiao et al. 2015). The aim is to build an experimental graph that represents the connectivity of the methyl resonances in a NOESY spectrum and to overlay this with a methyl–methyl network obtained from a known three-dimensional structure. If a unique overlay between these graphs is found then the assignment can be transferred from the known structure to the NOE graph, providing the desired chemical shift assignment. A significant advantage of these approaches is that, when a representative structure is known, they provide an exact and complete search of the solution space. The structure-based methods provide a significant alternative to previous approaches as they can be performed independently of a backbone assignment and so can be utilised when backbone assignments are impossible or unavailable.
A key part of the structure-based strategies is to reduce the available solution space, which scales approximately with n!, where n is the number of methyl groups. This is achieved by (1) assigning the residue type to each methyl resonance in the methyl-TROSY HMQC spectrum, giving nile!(nval)!(nleu)! solutions, and (2) linking intra-residue valine and leucine methyl resonances into pseudoatoms to further reduce the space to nile!(nval/2)!(nleu/2)! solutions (Pritišanac et al. 2017). Previously, the resonances of the two prochiral methyl groups of valine and leucine were linked using 3D or 4D NOESY spectra recorded with short mixing times. However, this approach has several shortcomings since it can be challenging to distinguish intra-residue NOEs from inter-residue NOEs.
Below we present a 3D-HMBC-HMQC experiment as an efficient method to link intra-residue methyl groups in valine and leucine residues. To demonstrate the applicability of the method, the experiment is applied to two large systems: the 81 kDa Malate synthase G (MSG), and the 360 kDa α7α7 half-proteasome complex of T. acidophilum. Intra-residue correlations are observed for approximately 90% of valine and leucine residues in these two systems.
Materials and methods
Density functional theory calculations
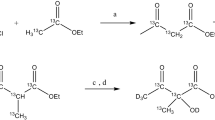
Scalar coupling constants were obtained from Density Functional Theory (DFT) calculations on the Ac-Val-NMe molecule shown in Fig. 1a using the programme Gaussian 09 (Frisch et al. 2016). Initially a structure optimisation was carried out using the B3LYP functional with the 6-31G* basis set (Ditchfield et al. 1971; Hehre et al. 1972). Subsequently, scalar couplings were calculated using a gauge-independent atomic orbital (GIAO) approach, as implemented in Gaussian with the keyword NMR = SpinSpin.
Assessing inter-methyl scalar couplings within a valine side-chain. a Scalar couplings between the two methyl groups of the valine side-chain obtained from DFT calculations. b A one-dimensional 1H NMR (500 MHz) spectrum of {U-[12C, 2H] [13C,1H]2} α-ketoisovaleric acid. The 128 Hz coupling corresponds to the intra-methyl one-bond 1H-13C coupling, whereas the ~ 5 Hz coupling is a three-bond inter-methyl 1H-13C coupling
Protein expression
Isotopically labelled MSG was produced as described previously (Tugarinov and Kay 2003; Pritchard and Hansen 2019; Korzhnev et al. 2004) with a slight modification. Briefly, the MSG gene, with a C-terminal his6 tag, in a kanamycin resistant pET28a vector, was transformed into BL21 (λDE3) E. coli cells for protein expression. A single colony was inoculated in 5 mL of LB media supplemented with kanamycin (50 μg mL−1) at 37 °C. Once the primary culture reached an OD600 between 0.8 and 1.0, it was used to inoculate a 50 ml M9 minimal media culture made with 2H2O and supplemented with 1 g L−1 [1H,15N]-ammonium chloride and 3 g L−1 of [2H,12C]-glucose as the sole nitrogen and carbon sources, respectively. The pre-culture was used to inoculate 1 L of M9 media and grown at 37 °C to OD600 ≈ 0.45. MSG expression was induced for > 16 h with 1 mM IPTG at 21 °C. To achieve the U-[12C, 2H]-LV-[13CH3]2 methyl labelling {U-[12C,2H] [13CH3]2} α-ketoisovaleric acid was added one hour prior to induction, while the U-[12C, 2H]-LV-[13CH3] labelling scheme was achieved by adding {U-[12C,2H] [13CH3]} α-keto-isovalerate.
The cells were lysed by sonication in Lysis buffer (20 mM Tris pH 7.8, 300 mM NaCl, 10 mM 2-mercaptoethanol) supplemented with 10 mg DNase1 (Sigma), 10 mg hen egg lysozyme (Sigma) and 1 complete™ Mini Protease Inhibitor Cocktail tablets (Sigma) per 50 mL lysate. The lysate was pelleted and MSG was purified from the soluble fraction by Ni-NTA affinity chromatography using a HisTrap 5 mL HP column (GE Healthcare), which was pre-equilibrated with Lysis buffer. Protein was eluted from the column using a 10 mM to 250 mM imidazole gradient. The fractions containing MSG were further purified by size exclusion chromatography using a Superdex 200 16/600 gel filtration column (GE Healthcare) (20 mM NaH2PO4, pH 7.1, 5 mM dithiothreitol).
To produce α-subunit complex (α7α7) of T. acidophilum proteasome, the αWT clone, with a N-terminal Histidine tag and a TEV protease site, was transformed into BL21 (λDE3) E. coli cells. The protein expression protocol for this α-subunit complex is same as MSG up to induction. The α-subunit complex culture was induced at OD600 ≈ 0.9 with 1 mM IPTG at 37 °C for 5 h. The U-[12C, 2H]-LV-[13CH3]2 methyl labelling scheme was achieved as described above. The cells were lysed by sonication in lysis buffer (50 mM NaH2PO4 pH 8.0, 0.2 M NaCl, 10 mM imidazole) and purified by Ni-NTA chromatography as before. After purification by Ni-NTA, TEV protease was added and the protein was dialyzed against 2 L of dialysis buffer (50 mM Tris–HCl pH 8.0, 1 mM EDTA, 5 mM 2-mercaptoethanol) overnight at 4 °C. After TEV cleavage the protein was further purified by Ni-NTA chromatography to remove the histidine tag and un-cleaved protein followed by size exclusion chromatography using a Superdex 200 16/600 gel filtration column (GE Healthcare) (50 mM NaH2PO4 pH 7.5, 0.1 M NaCl).
NMR spectroscopy
The NMR experiments on MSG were performed on a ~ 400 µM sample in 20 mM Sodium phosphate buffer pH 7.1, 5 mM DTT, 20 mM MgCl2, 0.05% NaN3 at 37 °C on a Bruker 800 MHz Avance III HD spectrometer equipped with Z-gradient triple-resonance TCI cryoprobe. The 3D-HMBC-HMQC experiment was acquired with 1024, 96, and 64 complex points in the 1H, 13Chmqc, and 13Chmbc dimensions with spectral widths of 11161 Hz, 2632 Hz, and 2632 Hz, respectively. Eight scans were collected per increment with a recycling delay of 1 s leading to a total experiment time of 63 h. The transfer time in the HMBC block was 23.5 ms (n = 3). The 4D-HMQC-NOESY-HMQC (Vuister et al. 1993; Tugarinov et al. 2005) recorded on MSG was acquired with non-uniform sampling (NUS) over the Nyquist grid consisting of 1024, 64, 72, and 96 complex points with spectral widths of 11161 Hz, 3220 Hz, 3220 Hz, and 2121 Hz, respectively. Four scans were collected per increment with a recycle delay of 1 s and the mixing time was 150 ms. The 4D-HMQC-NOESY-HMQC was recorded using a 1.6% NUS sampling schedule generated with a Poisson Gap distribution (Hyberts et al. 2010). The spectra were reconstructed using an iterative soft thresholding (IST) algorithm (Hyberts et al. 2012) and transformed in nmrPipe (Delaglio et al. 1995). The spectra were processed on the UCL Legion and NMRbox servers (Maciejewski et al. 2017).
The experiments recorded on the α7α7 proteasome were performed on 1.2 mM sample (monomer concentration) in 20 mM potassium phosphate pH 6.8, 50 mM NaCl, 1 mM EDTA, 2 mM DTT, 0.03% NaN3 at 50 °C on a Bruker 950 MHz Avance III HD spectrometer equipped with Z-gradient triple-resonance TCI cryoprobe. The 3D-HMBC-HMQC experiment was acquired with 1024, 84, and 48 complex points in the 1H, 13Chmqc, and 13Chmbc dimensions, respectively, with spectral widths of 15244 Hz (1H) and 3333 Hz (13C). 16 scans were collected for each increment with a recycle delay of 1 s for a total experiment time of 80 h.
All spectra were analysed using the CCPN (Vranken et al. 2005) and NMRFAM-Sparky (Lee et al. 2015) software packages. Peak heights used for signal-to-noise calculations were obtained using the inbuilt tools of NMRFAM-Sparky, whereas the noise level was estimated using the inbuilt tool of nmrPipe.
Results and discussion
Density functional theory (DFT) calculations were carried out on the Ac-Val-NMe molecule, Fig. 1a, in order to assess the possibility of obtaining intra-residue through-bond methyl–methyl correlations in large proteins. This molecule mimics a valine side chain within a protein environment. The calculations show the presence of an inter-methyl three-bond 13C-1H scalar coupling, 3J(1Ha,13Cb) = 4.8 Hz, Fig. 1a, which was confirmed experimentally by a 1D 1H NMR spectrum of {U-[12C,2H] [13C,1H]2} α-ketoisovaleric acid, Fig. 1b. The presence of an inter-methyl long-range scalar coupling of ~ 5 Hz, in agreement with previous observations (Reckel et al. 2011; Bax et al. 1994), opens up the possibility for obtaining through-bond methyl–methyl correlations in large proteins for valine and leucine residues.
The 3D-HMBC-HMQC experiment
The proposed HMBC-HMQC pulse sequence shown in Fig. 2 consists of two back-to-back HMQC elements and is designed to obtain intra-residue methyl–methyl correlations. The first HMQC element (HMBC) (Bax and Summers 1986) is optimised for the three-bond, 3JCH, coupling transfer between the proton of one of the prochiral methyl groups, 1Ha, and the carbon of the other methyl group, 13Cb, while suppressing the one-bond coupling transfer. The second HMQC element is optimised for the standard one-bond 1JCH coupling transfer.
Pulse sequence of the 3D-HMBC-HMQC experiment designed to correlate intra-residue methyl groups of leucine and valine side-chains. The carrier positions are 4.7 and 20 ppm for 1H and 13C, respectively. Hard 90° (180°) rf-pulses are indicated by narrow (wide) black bars and are applied at the highest available powers. The delay τ is set to 3.91 ms and is optimised for 1/(2 × 1JCH), where 1JCH is 128 Hz. The constant n is set to an integer depending on the desired evolution time of the long-range scalar coupling 3JCH. 13C decoupling during acquisition is achieved with a 3 kHz GARP4 (Shaka et al. 1985) scheme. Pulses are applied with x phase unless stated otherwise. The phase cycle used is φ1: x, −x, φ3: 2(x), 2(−x), φrec: x, 2(−x), x. φ2 and φ4 are decremented by 90° in accordance with the states-TPPI (Kay et al. 1989) scheme to achieve the required frequency discrimination in F1 and F2, respectively. Gradient pulses of 1 ms are represented by black rectangles and are applied with strengths of g1: 25.1 G cm−1, g2: 5.9 G cm−1, g3: 9.1 G cm−1
During the HMBC element, transverse proton magnetisation is allowed to evolve for a period of 2nτ, where n is an integer and τ = 1/(2 × 1JCH). This allows magnetisation to transfer from 1Ha to 13Cb while refocusing the one-bond proton–carbon coupling. In the product operator formalism (Sørensen et al. 1984) this results in a density operator at point 1 in Fig. 2 proportional to
A \(90_{{ \pm {\text{x}}}}^{\text{o}}\)13C pulse generates and selects the inter-methyl multi-quantum (MQ) coherence, \(\pm2H_{\text{x}}^{\text{a}} C_{{ {\text{y}}}}^{\text{b}},\) which evolves during t1 between 1 and 2. An inversion 180° 1H pulse in the middle of the t1 period refocuses the 1H chemical shift. In a subsequent delay of 2nτ between 2 and 3 the long-range coupling 3JCH refocuses and results in a density element at point 3 proportional to
The last part of the sequence between point 3 and 6 is a standard HMQC element. It is important to note that the second term in Eq. (2), \(2H_{\text{x}}^{\text{a}} C_{\text{z}}^{\text{b}},\) leads to a coherence of the type \(4H_{\text{y}}^{\text{a}} C_{\text{z}}^{\text{a}} C_{\text{z}}^{\text{b}}\) at point 4 and is therefore eliminated by the phase cycle of \(\phi_{3}.\) The first term in Eq. (2), \(H_{\text{y}}^{\text{a}},\) is labelled with the frequency of 13Ca between 4 and 5 and is finally detected at point 6, while decoupling 1H-13C scalar couplings. Fourier transform of the resulting dataset leads to cross-peaks at {ω1(13Cb), ω2(13Ca), ω3(1Ha)}. A key feature of the pulse sequence in Fig. 2 is the absence of 1H 90° pulses, which leads to the preservation of the methyl-TROSY effect (Ollerenshaw et al. 2003), thereby increasing the sensitivity of the experiment when applied to large systems.
The long-range 3JCH coupling that gives rise to the desired inter-methyl coherence in Eq (1), \(2H_{\text{x}}^{\text{a}} C_{\text{z}}^{\text{b}},\) is only approximately 5 Hz. A compromise between relaxation and scalar-coupling transfer should be considered, since a delay of 1/(2 × 3JCH) = 100 ms, which is required for full transfer, is impractical for large proteins due to relaxation. The dominant relaxation pathway is due to transverse relaxation of 1H magnetisation during the 2t = 2(2n + 1)τ coupling-transfer steps. The signal intensity in the final spectrum is approximately proportional to:
where R2 is the apparent proton transverse relaxation rate. Figure 3 shows the calculated intensity versus the constant n for several relaxation rates. The ideal choice for n varies from residue to residue depending on the residue-specific transverse relaxation rate, R2, which in turn depends on the methyl order parameters, S2axis. The HMBC transfer time, 2nτ, can also be optimised using a 2D version of the HMBC-HMQC experiment, where t2 in Fig. 2 is set to 0 s and n is arrayed. Choosing n based on the most rigid side chains seems to work well in our hands, since the more flexible side chains generally provide a higher signal-to-noise to begin with.
Plot of the calculated cross-peak intensity as a function of n (Fig. 2), for three different transverse relaxation rates, R2. The intensity on the y-axis is normalised to the case where R2 is 0 s−1. b Plot of the optimal value of n, nmax versus R2. The three cases from a are shown with coloured circles
Application to the 81 kDa malate synthase G and the 360 kDa α7α7 proteasome
The 3D-HMBC-HMQC approach was initially applied to the 81 kDa MSG, which has 116 leucine and valine residues. The signal-to-noise ratio (S/N) for the 3D-HMBC-HMQC was excellent, with 97% of all possible cross-peaks observed with S/N > 5 (see Fig. S1a), despite the 41 ns effective correlation time (Tugarinov and Kay 2005, 2006). In previous studies 226 leucine and valine methyl-resonances were assigned giving an assignment of 114 residues (L174 and V620 each have one unassigned methyl group), leaving only two residues unassigned. Out of these 226 assigned methyl resonances 9 resonances are marked as ambiguous in the original assignment (Tugarinov and Kay 2003).
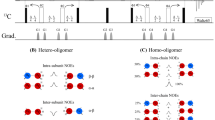
In the 3D-HMBC-HMQC spectra only two cross peaks should in principle be observed for each valine and leucine residue at (ω1, ω2, ω3) = (13Cγ1/δ1, 13Cγ2/δ2, 1Hγ2/δ2) and at (ω1, ω2, ω3) = (13Cγ2/δ2, 13Cγ1/δ1, 1Hγ1/δ1). This allows the two methyl groups to be correlated using the two 13C chemical shifts, Fig. 4a. As also exemplified in Fig. 4a, additional weak diagonal peaks are sometimes observed at (13Cγ1/δ1, 13Cγ1/δ1, 1Hγ1/δ1) and at (13Cγ2/δ2, 13Cγ2/δ2, 1Hγ2/δ2) if the 1JCH coupling is not completely refocused during the HMBC element. In cases where the methyl-TROSY HMQC spectrum is well-resolved, the 3D-HMBC-HMQC and the 2D-HMQC spectrum contain enough information to link all intra-residue methyl groups of valine and leucine residues. For the 81 kDa MSG, intra-residue methyl–methyl correlations could be confidently assigned for ~ 106 residues using solely the 3D-HMBC-HMQC and the 2D-HMQC experiments.
Two 2D planes of the 3D-HMBC-HMQC experiment on MSG showing the assignments of the two methyl groups of L64. b A 2D plane of the 4D-HMQC-NOESY-HMQC and the 3D-HMBC-HMQC spectra for L64 13Cγ1,1Hγ1. c 2D planes of the 4D-HMQC-NOESY-HMQC and the 3D-HMBC-HMQC spectra for linking the two methyl groups of L435. Cyan stars represent diagonal peaks
If the methyl-TROSY HMQC spectrum is crowded it can be challenging to link all the intra-residue methyl cross peaks confidently using only the two 13C shifts. For crowded methyl-TROSY HMQC spectra we therefore propose to use the 3D-HMBC-HMQC in combination with a 4D-HMQC-NOESY-HMQC (Vuister et al. 1993; Tugarinov et al. 2005) spectrum recorded with a long mixing time (up to 350 ms depending on size) and utilising a sample where both the proR and proS methyl groups are labelled with 13C and 1H in an otherwise deuterated background (U-[12C,2H]-LV-[13C,1H]2), Fig. 4b, c. The long mixing time guarantees that the intra-residue NOE is observed, which can then be easily identified based on the 3D-HMBC-HMQC, Fig. 4b, c. Using this methodology it is possible to identify intra-residue methyl–methyl cross peaks for 228 methyl resonances in MSG, which resulted in the successful linking of 98.3% of the leucine and valine methyl coherences. Additionally, the 3D-HMBC-HMQC confirms all nine tentative assignments made previously (Tugarinov and Kay 2003) by a backbone dependent assignment and also permits the assignment of the two missing methyl groups, one for L174 (13C = 25.70 ppm and 1H = 0.95 ppm) and one for V620 (13C = 21.45 ppm and 1H = 0.82 ppm).
Indirect methods based on methyl–methyl NOESY spectra, which are independent of the backbone, have traditionally been used to link intra-residue methyl groups of valine and leucine residues in large proteins (Sprangers and Kay 2007). The most commonly used method is to record HMQC-NOESY-HMQC spectra with a short mixing time, in the range of ca. 25–50 ms depending on the size of protein, utilising a U-[12C,2H]-LV-[13C,1H]2 labelled sample. The aim is to choose a mixing time such that the intra-residue NOE (~ 2.5 Å) can be distinguished from all other inter-residue NOEs. In practice selecting an appropriate mixing time can be challenging without prior knowledge, as exemplified in Fig. 5a for L64 of MSG. This is particularly the case for proteins with varying side-chain dynamics, where the correlation time for the intra-residue methyl–methyl vector can vary greatly throughout the protein and so the observed intra-residue NOEs vary substantially from residue to residue. The scenario is often that for a short NOESY mixing time some valine and leucine residues show no cross peaks, some show a single cross peak, and some may show many cross peaks.
HMBC-HMQC and HMQC-NOESY-HMQC spectra of MSG. a A 2D plane extracted from a 4D-HMQC-NOESY-HMQC spectrum focussing on L85. The spectrum was recorded on a doubly U-[12C,2H]-LV-[13C,1H]2 labelled sample with a mixing time of 150 ms. Multiple cross-peaks with similar intensity are observed, which makes an identification of the intra-residue correlation impossible. b An additional 4D-HMQC-NOESY-HMQC spectrum recorded on a singly labelled sample identifies the intra-residue methyl–methyl correlation. The arrow shows the intra-residual methyl–methyl correlation for L85, identified by absence of the peak in the singly methyl labelled sample. c Overlap in the 2D methyl-TROSY HMQC spectrum is easily resolved using the 3D-HMBC-HMQC spectrum. Two cross peaks corresponding V490 and V556 are observed. Cyan stars represent diagonal peaks
Related to the method above, is a comparison of a HMQC-NOESY-HMQC spectrum recorded on a U-[12C,2H]-LV-[13C,1H]2 sample with a HMQC-NOESY-HMQC spectrum of a sample with only one of the methyl groups labelled, U-[12C,2H]-LV-[13C,1H], Fig. 5b. In the spectrum obtained on the U-[12C,2H]-LV-[13C,1H]-sample, one cross peak should be absent compared to the spectra acquired on the U-[12C,2H]-LV-[13C,1H]2 sample. A key caveat of this approach is that the two labelling schemes lead to different relaxation properties due to the presence/absence of protons on the adjacent methyl, which affects the overall sensitivity. The intra-residue methyl–methyl correlation can be identified confidently using the 3D-HMBC-HMQC spectrum in combination with the HMQC-NOESY-HMQC spectrum, both recorded on a doubly U-[12C,2H]-LV-[13C,1H]2 labelled sample.
Since the 3D-HMBC-HMQC experiment only utilises scalar coupling transfers it provides additional information compared to the NOESY experiments. First and foremost, only a single cross-peak is observed per methyl group in addition to a potentially easily-identified diagonal peak. This means that if multiple cross-peaks are observed in the 3D-HMBC-HMQC for an apparent single peak in the reference 2D-methyl-HMQC spectrum, then it is a clear indication of overlap in the 2D spectrum. Hence overlapping peaks can be readily identified, as two cross-peaks will be observed, Fig. 5c.
To demonstrate the utility of the 3D-HMBC-HMQC experiment on larger protein complexes, a 3D-HMBC-HMQC spectrum was recorded on the 360 kDa α7α7 “half-proteasome” from T. acidophilum, with an effective rotational correlation time of ~ 120 ns at 50 °C (Tugarinov et al. 2007). Still, good signal-to-noise ratios were generally obtained, Fig. S1b, with 76% of the expected cross-peaks having S/N > 5 and 83% of the valine and leucine residues having at least one HMBC-HMQC cross-peak with S/N > 5. In the α-domain of the proteasome there are 19 leucine and 21 valine residues. In previous studies, 74 leucine and valine methyl-resonances were assigned out of the possible 80 methyl-resonances (Sprangers and Kay 2007). Despite its high molecular weight and slow tumbling, cross peaks with S/N > 3 were observed for 80% of the possible methyl resonances in the 3D-HMBC-HMQC experiment (64 out of 80). Overall, 87.5% of the methyl groups in valine and leucine (35 residues out of 40) side chains were correctly paired. Figure 6a–d demonstrate that despite the size of the α7α7 proteasome the 3D-HMBC-HMQC experiment provides good signal-to-noise across a range of order parameters (Tugarinov et al. 2007).
3D-HMBC-HMQC spectra of the 360 kDa α7α7 proteasome to obtain intra-residue methyl–methyl correlations. a–d are 2D planes of the 3D-HMBC-HMQC spectra for L38, L201, V190, and V184 respectively, which have different order parameters and therefore different internal dynamics. Cyan stars represent diagonal peaks
Conclusions
In conclusion, we presented a 3D-HMBC-HMQC scalar coupling based approach to link the two prochiral methyl groups in leucine and valine side-chains in large proteins. The method was demonstrated on the 81 kDa Malate synthase G (MSG) and the 360 kDa α7α7 proteasome complex. Only intra-residue correlations are observed since the method utilises scalar-coupling based transfers, which also means that overlap in the 2D-methyl-TROSY HMQC spectrum can be identified by the presence of multiple cross-peaks in the 3D-HMBC-HMQC spectrum. In challenging cases, the presented method can be combined with HMQC-NOESY-HMQC spectra to remove ambiguities in the assignment of intra-residue methyl resonances. Moreover, when methyl resonances are assigned using a mutational approach (Sprangers and Kay 2007; Rosenzweig et al. 2013), the 3D-HMBC-HMQC could also provide valuable insight to confirm the assignments. Finally, the presented approach only relies on a single sample to pair intra-residue methyl resonances and, in favourable cases, it therefore reduces the number of samples required for obtaining the required input for structure-based assignment of methyl NMR resonances.
References
Bax A, Summers MF (1986) Proton and carbon-13 assignments from sensitivity-enhanced detection of heteronuclear multiple-bond connectivity by 2D multiple quantum NMR. J Am Chem Soc 108:2093–2094
Bax A, Delaglio F, Grzesiek S, Vuister GW (1994) Resonance assignment of methionine methyl groups and χ3 angular information from long-range proton–carbon and carbon–carbon J correlation in a calmodulin peptide complex. J Biomol NMR 4:787–797
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) Nmrpipe—a multidimensional spectral processing system based on unix pipes. J Biomol NMR 6:277–293
Ditchfield R, Hehre WJ, Pople JA (1971) Self-consistent molecular-orbital methods. IX. An extended Gaussian gaussian-type basis for molecular-orbital studies of organic molecules. J Chem Phys 54:724–728
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery JA Jr, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2016) Gaussian 09, Revision A.02. Gaussian, Inc., Wallingford
Gelis I, Bonvin AMJJ, Keramisanou D, Koukaki M, Gouridis G, Karamanou S, Economou A, Kalodimos CG (2007) Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell 131:756–769
Hansen DF, Kay LE (2011) Determining valine side-chain rotamer conformations in proteins from methyl 13C chemical shifts: application to the 360 kDa half-proteasome. J Am Chem Soc 133:8272–8281
Hansen DF, Neudecker P, Kay LE (2010) Determination of isoleucine side-chain conformations in ground and excited states of proteins from chemical shifts. J Am Chem Soc 132:7589–7591
Hehre WJ, Ditchfield R, Pople JA (1972) Self-consistent molecular orbital methods. XII. Further extensions of Gaussian-type basis sets for use in molecular orbital studies of organic molecules. J Chem Phys 56:2257–2261
Hyberts SG, Takeuchi K, Wagner G (2010) Poisson-gap sampling and forward maximum entropy reconstruction for enhancing the resolution and sensitivity of protein NMR data. J Am Chem Soc 132:2145–2147
Hyberts SG, Milbradt AG, Wagner AB, Arthanari H, Wagner G (2012) Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson Gap scheduling. J Biomol NMR 52:315–327
Kay LE, Marion D, Bax A (1989) Practical aspects of 3D heteronuclear NMR of proteins. J Magn Reson 84:72–84
Korzhnev DM, Kloiber K, Kanelis V, Tugarinov V, Kay LE (2004) Probing slow dynamics in high molecular weight proteins by methyl-TROSY NMR spectroscopy: application to a 723-residue enzyme. J Am Chem Soc 126:3964–3973
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327
Maciejewski MW, Schuyler AD, Gryk MR, Moraru II, Romero PR, Ulrich EL, Eghbalnia HR, Livny M, Delaglio F, Hoch JC (2017) NMRbox: a resource for biomolecular NMR computation. Biophys J 112:1529–1534
Monneau YR, Rossi P, Bhaumik A, Huang C, Jiang Y, Saleh T, Xie T, Xing Q, Kalodimos CG (2017) Automatic methyl assignment in large proteins by the MAGIC algorithm. J Biomol NMR 69:215–227
Ollerenshaw JE, Tugarinov V, Kay LE (2003) Methyl TROSY: explanation and experimental verification. Magn Reson Chem 41:843–852
Pritchard RB, Hansen DF (2019) Characterising side chains in large proteins by protonless 13C-detected NMR spectroscopy. Nat Commun 10:1747
Pritišanac I, Degiacomi MT, Alderson TR, Carneiro MG, Eiso AB, Siegal G, Baldwin AJ (2017) Automatic assignment of methyl-NMR spectra of supramolecular machines using graph theory. J Am Chem Soc 139:9523–9533
Reckel S, Gottstein D, Stehle J, Löhr F, Verhoefen M-K, Takeda M, Silvers R, Kainosho M, Glaubitz C, Wachtveitl J, Bernhard F, Schwalbe H, Güntert P, Dötsch V (2011) Solution NMR structure of proteorhodopsin. Angew Chem Int Ed 50:11942–11946
Rosenzweig R, Moradi S, Zarrine-Afsar A, Glover JR, Kay LE (2013) Unraveling the mechanism of protein disaggregation through a ClpB–DnaK interaction. Science 339:1080–1083
Ruschak AM, Kay LE (2010) Methyl groups as probes of supra-molecular structure, dynamics and function. J Biomol NMR 46:75–87
Salzmann M, Pervushin K, Wider G, Senn H, Wüthrich K (2000) NMR assignment and secondary structure determination of an octameric 110 kDa protein using TROSY in triple resonance experiments. J Am Chem Soc 122:7543–7548
Shaka A, Barker P, Freeman R (1985) Computer-optimized decoupling scheme for wideband applications and low-level operation. J Magn Reson 64:547–552
Sørensen OW, Eich GW, Levitt MH, Bodenhausen G, Ernst RR (1984) Product operator formalism for the description of NMR pulse experiments. Prog Nucl Magn Reson Spectrosc 16:163–192
Sprangers R, Kay LE (2007) Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature 445:618–622
Tugarinov V, Kay LE (2003) Ile, Leu, and Val methyl assignments of the 723-residue malate synthase G using a new labeling strategy and novel NMR methods. J Am Chem Soc 125:13868–13878
Tugarinov V, Kay LE (2004) An isotope labeling strategy for methyl TROSY spectroscopy. J Biomol NMR 28:165–172
Tugarinov V, Kay LE (2005) Quantitative 13C and 2H NMR relaxation studies of the 723-residue enzyme malate synthase G reveal a dynamic binding interface. Biochemistry 44:15970–15977
Tugarinov V, Kay LE (2006) Relaxation rates of degenerate 1H transitions in methyl groups of proteins as reporters of side-chain dynamics. J Am Chem Soc 128:7299–7308
Tugarinov V, Muhandiram R, Ayed A, Kay LE (2002) Four-dimensional NMR spectroscopy of a 723-residue protein: chemical shift assignments and secondary structure of malate synthase G. J Am Chem Soc 124:10025–10035
Tugarinov V, Hwang PM, Ollerenshaw JE, Kay LE (2003) Cross-correlated relaxation enhanced 1H–13C NMR spectroscopy of methyl groups in very high molecular weight proteins and protein complexes. J Am Chem Soc 125:10420–10428
Tugarinov V, Kay LE, Ibraghimov I, Orekhov VY (2005) High-resolution four-dimensional 1H–13C NOE spectroscopy using methyl-TROSY, sparse data acquisition, and multidimensional decomposition. J Am Chem Soc 127:2767–2775
Tugarinov V, Sprangers R, Kay LE (2007) Probing side-chain dynamics in the proteasome by relaxation violated coherence transfer NMR spectroscopy. J Am Chem Soc 129:1743–1750
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Vuister GW, Clore GM, Gronenborn AM, Powers R, Garrett DS, Tschudin R, Bax A (1993) Increased resolution and improved spectral quality in four-dimensional 13C/13C-separated HMQC-NOESY-HMQC spectra using pulsed field gradients. J Magn Reson Ser B 101:210–213
Xiao Y, Warner LR, Latham MP, Ahn NG, Pardi A (2015) Structure-based assignment of Ile, Leu, and Val methyl groups in the active and inactive forms of the mitogen-activated protein kinase extracellular signal-regulated kinase 2. Biochemistry 54:4307–4319
Acknowledgements
Prof. Lewis E. Kay (University of Toronto) is acknowledged for providing the αWT proteasome clone (α-subunit complex α7α7 from T. acidophilum), Dr Angelo M. Figueiredo is acknowledged for help with recording NMR spectra. LS and HWM acknowledge the Wellcome Trust and UCB pharma for PhD studentships, respectively. The BBSRC (BB/R000255/1), Wellcome Trust (ref 101569/z/13/z), and the EPSRC are acknowledged for supporting the ISMB NMR facility at University College London. Access to ultra-high field NMR spectrometers was supported by the Francis Crick Institute through provision of access to the MRC Biomedical NMR Centre. The Francis Crick Institute receives its core funding from Cancer Research UK (FC001029), the UK Medical Research Council (FC001029), and the Wellcome Trust (FC001029). This research is supported by the Leverhulme Trust (RPG-2016-268).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Siemons, L., Mackenzie, H.W., Shukla, V.K. et al. Intra-residue methyl–methyl correlations for valine and leucine residues in large proteins from a 3D-HMBC-HMQC experiment. J Biomol NMR 73, 749–757 (2019). https://doi.org/10.1007/s10858-019-00287-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-019-00287-9