Abstract
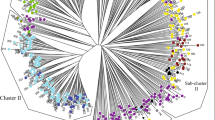
Egyptian clover (Trifolium alexandrinum) has received limited attention regarding its genetic diversity, rendering it susceptible to climate change. To develop resilient Berseem cultivars, exploring its genetic resources is vital for enhancing the diversity within its germplasm, thereby optimizing breeding programs. The examination of 24 Berseem accessions. Using three different molecular marker types; Inter-Simple Sequence Repeat (ISSR) (10 primers), Start Codon Targeted polymorphism (SCoT) (22 primers), and Sequence-Related Amplified Polymorphism (SRAP) (17 primer combinations) revealed a high genetic variability. ISSR markers had the highest polymorphism rate (82.2%) compared to SCoT (69.6%) and SRAP (65.2%). Unweighted Pair Group Method with Arithmetic Mean clustering analysis divided the studied accessions into two major groups, supported by STRUCTURE analysis, which also split samples into two subpopulations (∆K = 2). Additionally, this classification was verified by both principal component analysis and a heat map. The study underscores the importance of Berseem’s genetic diversity in crop development and breeding, with ISSR markers offering valuable insights due to their high polymorphism. Despite the diverse geographic origins of the accessions, the lack of geographic clustering implies significant genetic mixing, possibly because of high outcrossing facilitated by insect pollination or self-incompatibility. The combined use of multiple markers deepens our understanding of Berseem’s genetics by providing more detailed and comprehensive coverage of the clover genome, allowing for increased resolution, validation of results, and a holistic perspective on the genomic landscape, furthermore, identifying and recommending specific Berseem accessions (NGB-43724, NGB-19680, Gemeiza1, and Giza6) as prospective candidates for future breeding programs in Egypt.



Similar content being viewed by others
Data availability
The data and material are part of a Ph.D. thesis. and are available upon request.
References
Abdel-Fatah BE, Bakheit BR (2019) Genetic diversity between two Egyptian clover varieties and QTL analysis for some agro-morphological traits. Mol Biol Rep 46(1):897–908. https://doi.org/10.1007/s11033-018-4546-4
Abuzaid AS, Jahin HS, Asaad AA, Fadl ME, AbdelRahman MA, Scopa A (2021) Accumulation of potentially toxic metals in Egyptian alluvial soils, berseem clover (Trifolium alexandrinum L.), and groundwater after long-term wastewater irrigation. Agriculture 11(8):713
Aljrf A, Ksibe B, Alnaddaf L, Saleh M (2021) The protein fingerprinting and genetic variations among different genotypes of Aegilops belonging to four sections. J Genet Environ Resour Conserv 9(1):90–97
Amiryousefi A, Hyvönen J, Poczai P (2018) iMEC: online marker efficiency calculator. Appl Plant Sci 6(6):e01159. https://doi.org/10.1002/aps3.1159
Bakheit BR (2013) Egyptian clover (Trifolium alexandrinum L.) breeding in Egypt: a review. Asian J Crop Sci 5(4):325–337. https://doi.org/10.3923/ajcs.2013.325.337
Borcard D, Gillet F, Legendre P (2011) Numerical ecology with R. Springer, Cham
Cabo S, Ferreira L, Carvalho A, Martins-Lopes P, Martín A, Lima-Brito JE (2014) Potential of Start Codon Targeted (SCoT) markers for DNA fingerprinting of newly synthesized tritordeums and their respective parents. J Appl Genet 55:307–312. https://doi.org/10.1007/s13353-014-0211-3
Chapman MA, Hvala J, Strever J, Burke JM (2010) Population genetic analysis of safflower (Carthamus tinctorius; Asteraceae) reveals a Near Eastern origin and five centers of diversity. Am J Bot 97(5):831–840. https://doi.org/10.3732/ajb.0900137
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
El-Badan GE, Amin AW, Ashour FM, El Sadek LM (2022) Characterization of the genetic diversity of some species of genus vicia using ISSR and ITS molecular techniques. Egypt J Bot 62(2):475–492
El-Esawi MA, Elashtokhy MM, Shamseldin SA, El-Ballat EM, Zayed EM, Heikal YM (2022) Analysis of genetic diversity and phylogenetic relationships of wheat (Triticum aestivum L.) genotypes using phenological Molecular and DNA Barcoding Markers. Genes 14(1):34. https://doi.org/10.3390/genes14010034
Etminan A, Pour-Aboughadareh A, Noori A, Ahmadi-Rad A, Shooshtari L, Mahdavian Z, Yousefiazar-Khanian M (2018) Genetic relationships and diversity among wild Salvia accessions revealed by ISSR and SCoT markers. Biotechnol Biotechnol Equip 32(3):610–617. https://doi.org/10.1080/13102818.2018.1447397
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
FAO (2014) Egyptian Clover (Trifolium alexandrinum): King of Forage Crops. Food and agriculture organization of the United Nations. https://www.fao.org/publications/card/es/c/a9f7cc85-714a-47b4-a90e-21e2e753c7fd/
Faville MJ, Griffiths AG, Baten A, Cao M, Ashby R, Ghamkhar K, Hong W, Larking A, Williamson M, Webber Z (2020) Genomic assessment of white clover and perennial ryegrass genetic resources. J New Zealand Grassl. https://doi.org/10.33584/jnzg.2020.82.437
George J, Dobrowolski MP, van de Zijll Jong E, Cogan NO, Smith KF, Forster JW (2006) Assessment of genetic diversity in cultivars of white clover (Trifolium repens L.) detected by SSR polymorphisms. Genome 49(8):919–930. https://doi.org/10.1139/g06-079
Golkar P, Bakhtiari MA (2020) Evaluation of genetic diversity in the world collection of Eruca sativa L. using oil content, fatty acids and molecular markers. Ind Crops Prod 148:112280. https://doi.org/10.1016/j.indcrop.2020.112280
Griffiths AG, Moraga R, Tausen M, Gupta V, Bilton TP, Campbell MA, Ashby R, Nagy I, Khan A, Larking A (2019) Breaking free: the genomics of allopolyploidy-facilitated niche expansion in white clover. Plant Cell 31(7):1466–1487. https://doi.org/10.1105/tpc.18.00606
Haerinasab M, Ali-Farsangi F, Bordbar F, Eslami Farouji A (2020) Genetic diversity and infraspecific relationships of Trifolium fragiferum L. in Iran. Iran J Sci Technol Trans : Sci 44:345–354. https://doi.org/10.1007/s40995-020-00834-2
Hamrick J, Godt M, Murawski D, Loveless M (1991) Correlations between species traits and allozyme diversity: implications for conservation biology. Genet Conserv Rare Plants 3:30
Jaiswal RK, Jha S, Patel DCA (2023) Growth and yield analysis of berseem (Trifolium alexandrinum L.) genotypes under different cutting management practices in Chhattisgarh plains
Katoch R (2022) Nutritional quality of important forages. In Techniques in forage quality analysis (pp. 173-185). Singapore: Springer Nature Singapore.
Khal LH, AL-Anbari AK, Al-Hadeethi MA, Abdulrazaq RT (2020) Phylogenetic diversity of Trifolium L. Species in Iraq. J Plant Prod 11(11):1041–1044
Khan MA, von Witzke-Ehbrecht S, Maass BL, Becker HC (2009) Relationships among different geographical groups, agro-morphology, fatty acid composition and RAPD marker diversity in safflower (Carthamus tinctorius). Genet Resour Crop Evol 56:19–30. https://doi.org/10.1007/s10722-008-9338-6
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685. https://doi.org/10.1038/227680a0
Lê S, Josse J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25:1–18
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21(9):2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Ma S, Han C, Zhou J, Hu R, Jiang X, Wu F, Tian K, Nie G, Zhang X (2020) Fingerprint identification of white clover cultivars based on SSR molecular markers. Mol Biol Rep 47:8513–8521. https://doi.org/10.1007/s11033-020-05893-7
Mardi M, Naghavi M, Pirseyedi S, Kazemi AM, Rashidi MS, Ahkami A, Omidbakhsh M, Alavi N, Salehi SP, Katsiotis A (2011) Comparative assessment of SSAP, AFLP and SSR markers for evaluation of genetic diversity of durum wheat (Triticum turgidum L. var. durum). https://api.semanticscholar.org/CorpusID:55837808
Marwal A, Gaur RK (2020) Molecular markers: tool for genetic analysis. In animal biotechnology. Elsevier, Amsterdam, pp 353–372
Metsalu T, Vilo J (2015) ClustVis: a web tool for visualizing clustering of multivariate data using principal component analysis and heatmap. Nucleic Acids Res 43(W1):W566–W570. https://doi.org/10.1093/nar/gkv468
Nay MM, Grieder C, Frey LA, Amdahl H, Radovic J, Jaluvka L, Palmé A, Skøt L, Ruttink T, Kölliker R (2023) Multi-location trials and population-based genotyping reveal high diversity and adaptation to breeding environments in a large collection of red clover. Front Plant Sci 14:1128823. https://doi.org/10.3389/fpls.2023.1128823
Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20(2):289–290. https://doi.org/10.1093/bioinformatics/btg412
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
Rahimi-Ashtiani S, Sahab S, Panter S, Mason J, Spangenberg G (2015) Clovers (Trifolium spp.). Agrobact Protoc 1:223–235. https://doi.org/10.1007/978-1-4939-1695-5_18
Rai MK (2023) Start codon targeted (SCoT) polymorphism marker in plant genome analysis: current status and prospects. Planta 257(2):34. https://doi.org/10.1007/s00425-023-04067-6
Rayan WA, Osman SA (2019) Phylogenetic relationships of some Egyptian soybean cultivars (Glycine max L.) using SCoT marker and protein pattern. Bull Natl Res Centre 43(1):1–10. https://doi.org/10.1186/s42269-019-0197-4
Sammour R (1999) SDS-PAGE analysis of the seed proteins of some Trifolium taxa. Plant Var Seeds 12(1):11–21
Sokal R, Michener C (1958) University of Kansas. A statistical method for evaluating systematic relationships. University of Kansas science bulletin, Lawrence
Wu F, Ma S, Zhou J, Han C, Hu R, Yang X, Nie G, Zhang X (2021a) Genetic diversity and population structure analysis in a large collection of white clover (Trifolium repens L.) germplasm worldwide. PeerJ 9:e11325. https://doi.org/10.7717/peerj.11325
Wu F, Zhang S, Gao Q, Liu F, Wang J, Wang X (2021b) Genetic diversity and population structure analysis in a large collection of Vicia amoena in China with newly developed SSR markers. BMC Plant Biol 21(1):1–12. https://doi.org/10.1186/s12870-021-03330-w
Yazıcılar B, Jannati G, Bezirganoglu I (2021) Genetic variations in Turkey cultivar and ecotype Medicago sativa species: cytological, total protein profile, and molecular characterization. J Genet Eng Biotechnol 19(1):1–11. https://doi.org/10.1186/s43141-021-00159-6
Yi L, Dong Z, Lei Y, Zhao J, Xiong Y, Yang J, Xiong Y, Gou W, Ma X (2021) Genetic diversity and molecular characterization of worldwide prairie grass (Bromus catharticus Vahl) accessions using SRAP markers. Agronomy 11(10):2054. https://doi.org/10.3390/agronomy11102054
Yousefi S, Saeidi H, Assadi M (2018) Genetic diversity analysis of red clover (Trifolium pratense L.) in Iran using sequence related amplified polymorphism (SRAP) markers. J Agric Sci Technol 20(2):373–386
Youssef SS, Ibrahim SA, Abdel Azim AM, Khashaba EH, El-Sharkawy AM (2019) Genetic variation among five egyptian clover cultivars using random amplified polymorphic DNA (RAPD) and sequence-related amplified polymorphism (SRAP) molecular markers. Egypt Acad J Biol Sci Physiol Mol Biol 11(2):37–45
Zayed EM (2013) Applications of biotechnology on Egyptian clover (berseem) (Trifolium alexandrinum L.). Int J Agric Sci Res 3(1):99–120
Acknowledgements
We wish to express our gratitude for the partial funding through Project Code (CFP4-PR-142) within the framework of the FAO BSF Project—Fourth Cycle. All aspects of the research, including plant collection, fieldwork, and laboratory analysis, adhered to the ethical principles set forth by the Ministry of Agriculture and Land Reclamation and any other relevant authorities in Egypt. This study was conducted according to the ethical guidelines outlined by these agencies to ensure the responsible and ethical conduct of plant research in the country. This work was performed at AGERI, 9-Elgamaa Street, Giza, Egypt.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
Conceptualization: All authors contributed to the initial idea and research design of the study. Data collection: H.M. Mohamed was responsible for gathering data and conducting experiments. Data analysis: H. M. Mohamed, S. S. El-Assal, and D. A. El-Khishin conducted the statistical analysis, interpreted the data, or developed the models. Writing: H.M. Mohamed and A.Y. Gamal El-Din drafted the manuscript and contributed to the writing process, including drafting sections or editing the text., Review and editing: all authors are responsible for reviewing and revising the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflicts of interest.
Ethical approval and consent to participate
Not applicable. This study did not involve human subjects or animal research.
Consent for publication
We affirm that all authors have reviewed and approved the content of this manuscript and agree to its submission for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mohamed, H.M., El-Assal, S.ED.S.M., Gamal El-Din, AK.Y. et al. Analysis of genetic diversity and population structure in some Egyptian Berseem (Trifolium alexandrinum) accessions based on ISSR, SCoT and SRAP markers. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01940-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01940-5