Abstract
The increase in the global human population and the accompanying challenges in meeting nutritional needs amidst climate change are a worldwide concern. Widespread protein and micronutrient deficiencies contribute to a significant number of individuals experiencing malnutrition, leading to severe health repercussions. This issue can be addressed through genomics-assisted breeding, particularly in enhancing the nutritional profile of vital staple crops like chickpea (Cicer arietinum L.). Chickpea, beyond being a rich source of protein, provides a diverse nutritional spectrum encompassing carbohydrates, fats, and minerals. To explore and improve the genetic basis of nutritional traits in chickpea, a study was conducted using 93 kabuli-type single plant derived lines and five cultivars in 2018, 2022, and 2023. Genotyping by sequencing revealed a total of 165K single nucleotide polymorphisms (SNPs) within this kabuli chickpea mini-core collection. After filtering for a minor allele frequency greater than 5%, 113,512 SNPs were utilized, distributed across eight chromosomes of the chickpea genome. Marker-trait associations were analyzed using genome wide association study, leading to the identification of 27 significantly associated SNPs from across all eight chromosomes linked to three seed nutritional concentrations and 100-seed weight. To unravel the molecular mechanisms governing seed protein, fiber, fat concentrations, and 100-seed weight, 31 candidate genes were determined within a 30 kb window size. This comprehensive approach holds promise for advancing crop breeding strategies to combat malnutrition and improve global food security.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chickpea (Cicer arietinum L., 2n = 16) is one of the most consumed food legumes globally, primarily used for human consumption but also as an animal feed. Chickpea was harvested from 15 million hectares with a production of 15.9 million tons in the world (FAO 2023). It offers a rich source of protein, starch and dietary fiber (Jukanti et al. 2012; Mugabe et al. 2023). Cultivated chickpea is categorized into two groups, desi and kabuli, based on various characteristics such as flower color, seed shape, and seed size (van der Maesen 1972). Desi types have small green, brown, and black seeds and pink flower colors, and display good stress resistance, while kabuli types are favored for their larger seed size, higher yield, and good seed quality parameters (Anbessa et al. 2006; Eker et al. 2022). Kabuli types are also known to have higher protein digestibility for human nutrition (Sanchez-Vioque et al. 1999; Wang et al. 2010).
The rapid increase in the global human population and the associated challenges in meeting nutritional demands amidst climate change have raised concerns worldwide (FAO 2020; FAO 2023). Protein and micronutrient deficiencies have resulted in a significant number of individuals suffering from malnutrition, leading to severe health consequences (WHO 2021; Beermann 2022). Protein deficiency is known to cause muscle deterioration, weakened immune system, growth retardation, and developmental issues (Rytter et al. 2014; Wu 2016). A balanced diet adhering to the Recommended Dietary Allowance (RDA) suggests a daily protein intake of 0.8 g per kilogram of body weight (Wu 2016). This means that females over 14 years of age should aim for 46 g of daily protein, while males over 19 years should consume 56 g (Meyers et al. 2006). According to the USDA (2019), a 100 g serving of chickpeas provides around 20.2 g of protein. Chickpeas, recognized as an economical and highly nutritious legume, have gained popularity as a dietary staple, particularly in developing regions and among individuals following a vegetarian diet (Iqbal et al. 2006; Vandemark et al. 2018).
Chickpea offers a diverse nutritional profile other than protein, including carbohydrates, fat, and minerals. The composition of chickpea seeds varies based on environmental factors, agronomic practices, variety, and type (desi vs. kabuli). Chickpea has higher fat content than other food legumes and some cereals but a lower fat content than oilseed legumes like soybeans and groundnuts. In desi chickpeas, fat content ranges from 3.10 to 4.93%, while kabuli varies from 4.60 to 5.67% (Singh 1985; Jukanti et al. 2012); higher fat kabuli are preferred for the making of the popular dish hummus. The main carbohydrate in chickpea is starch, comprising around 47.4–66.9% of the carbohydrate fraction (Singh 1985); soluble sugars, crude fiber, and dietary fiber contribute to the remaining carbohydrates. Chickpea is rich in dietary fiber, with insoluble and soluble fiber levels of approximately 10–18/100 g and 4–8/100 g, respectively (Tosh and Yada 2010). The presence of dietary fiber offers numerous health benefits, including improved digestion, a reduced risk of certain chronic diseases, and better weight management (Liu et al. 1999; Birketvedt et al. 2005; Petruzziello et al. 2006). Understanding the genetic basis of nutritional traits in chickpea will help in the selection and breeding of varieties with improved protein, fiber, and fat concentrations, thus contributing to the reduction of malnutrition and enhancing global food and nutritional security.
Recent advancements in chickpea genomics have facilitated the production of numerous genetic markers, linkage maps, and genome sequences (Jain et al. 2013; Leonforte et al. 2013; Stephens et al. 2014; Gaur et al. 2020). These developments have aided the use of Genome-wide association studies (GWAS) in chickpea. Genome-wide association study is a powerful tool for mapping complex traits that allows for the screening of diverse crop accessions with high-density markers, enabling the identification of genes associated with phenotypic traits. Identification of genes and the development of molecular markers in or near these genes can aid breeding programs worldwide in the improvement of various important traits in the crop. To date, these efforts have primarily focused on yield, drought resistance, and resistance to diseases such as Ascochyta blight and Fusarium wilt (Anbessa et al. 2009; Cobos et al. 2009). Some studies have explored the genetic architecture underlying chickpea's nutritional traits, but only of a few traits (iron and zinc concentrations and protein content; Jadhav et al. 2015; Upadhyaya et al. 2016; Sab et al. 2020; Mugabe et al. 2023). Therefore, the goals of this research are to study a wider range of nutritional traits to enable enhancement of kabuli chickpea's nutritional value.
The aim of this study is to determine the genetic factors affecting protein, fat, fiber concentrations, and 100-seed weight through GWAS. The findings from this research will not only contribute to the development of chickpea varieties with enhanced nutritional profiles but will also identify genotypes with high 100-seed weight and nutritional quantity. These identified genotypes can serve as breeding lines for the development of improved cultivars. By addressing malnutrition and promoting global food and nutritional security, this research aims to meet the nutritional demands of a growing population in a sustainable manner. Additionally, it strives to make a substantial contribution to the field, offering insights that support the creation of more nutritious chickpea varieties and sustainable agricultural practices.
Materials and methods
Plant material
In this study, 88 kabuli-type single plant derived lines were selected from the USDA Chickpea Core Collection (Kumar et al. 2014; Simon and Hannan 1995; GRIN-Global (ars-grin.gov). Plots of the selected 88 lines were grown with five check cultivars (‘Dwelley’, ‘Frontier’, ‘Sierra’, ‘Spanish White’, and ‘UC5’) in an irrigated field study at the Central Ferry farm, Washington (46°39′5.1″ N; 117°45′45.4″ W, elevation 198 m above sea level), in 2018, 2022, and 2023. The experimental design adopted was a randomized complete block design (RCBD), featuring single plots with four replications. Thirty seeds were planted in double rows with 30 cm center spacing, keeping 152 cm between rows and plots, in each of the 152 cm long rows. At maturity, the plots were hand harvested, followed by a standardized drying process to achieve uniform moisture content. Threshing was carried out using a Vogel thresher and the final seed cleaning phase was performed by use of a seed blower.
Phenotyping and descriptive statistics
Seed protein concentration was calculated for three years and fiber and fat analyses were calculated for seed for the first two years. Whole seed analyses were performed by NIR (Bruker Matrix-1). The spectrometer was calibrated by grinding 120 samples from 2020 plots. Total N concentration was determined using the LECO C/N analyzer. Nitrogen concentrations were converted to protein concentration using a 6.25 conversion factor (Jones 1931) (Table S1). The OPUS calibration software was used to calibrate protein. Fat and fiber calibrations were performed using the Bruker NIR according to the manufacturer’s instructions. A total of 200 g of harvested seeds per sample was used for each accession for NIR for 2018 and 2022. Three NIR scans were performed for each sample, and protein, fiber, and fat concentrations were estimated using an average of the three scans. The 2023 seed yields were low with insufficient seed for NIR analysis, so 2023 plot samples were ground and analyzed for total N using the LECO C/N analyzer and 6.25 N to protein conversion factor (Jones 1931).
Hundred-seed weight (100-SW) was measured for each accession. The four replications were measured individually by weighing 100 randomly selected seeds that had been dried to an average of 15% moisture after harvest, and the weight was recorded in grams and averaged over replications (Table S1). All analyses were expressed on a dry weight basis. For descriptive statistics, the range, mean, and standard error were calculated, and ANOVA and Pearson correlation between traits were conducted using SPSS 26.
Genotyping and haplotype analysis
DNA was extracted from young leaves of the accessions under controlled greenhouse conditions, employing the DNeasy Plant 96 kit (Qiagen Corp., Valencia, CA, USA). Single-nucleotide polymorphisms (SNPs) were discovered following single-enzyme (ApeKI) genotyping-by-sequencing (Elshire et al. 2011), conducted by a commercial company (LGC Biosearch Technologies, Berlin, Germany). The FreeBayes software (Garrison and Marth 2012) was utilized to call the identification of genetic variants, using the reference genome of kabuli chickpea, ‘CDC Frontier’ (Varshney et al. 2013). This process implemented with BamTools (Barnett et al. 2011) and the FreeBayes variant caller (Garrison and Marth 2012). The accessions, USDA Chickpea Kabuli Mini-Core Collection used were previously genotyped by Mugabe et al. (2023). The SNPs were filtered with minor allele frequency (MAF) > 5% SNP data. A final total of 113,645 markers across the eight chromosomes of the chickpea genome were obtained for genetic analysis.
Marker trait association analysis
A genome-wide marker-trait analyses were conducted to identify SNP markers associated with 100-SW and seed protein, fiber, and oil concentrations. Marker-trait associations (MTAs) were performed with the BLINK (Bayesian-information and Linkage-disequilibrium Iteratively Nested Keyway) model using the R/GAPIT 3.0 package (Wang and Zhang 2021; Huang et al. 2019). In the BLINK model, a fixed effect model is used to correct false negatives and false positives using Bayesian information. It also uses linkage disequilibrium information to eliminate the need for markers to be evenly distributed throughout the genome (Huang et al. 2019). Bonferroni correction was used to evaluate the significance of the SNP marker-trait association. To establish a significance of association between SNPs and the phenotypic traits, a threshold value of P ≤ 0.05 was applied.
Analysis of Linkage Disequilibrium
Linkage Disequilibrium (LD) analysis was performed using TASSEL v5.0. Significantly associated SNPs within an LD window were BLASTed against the chickpea genome (https://www.pulsedb.org), and potential candidate genes were determined for in the Arabidopsis homolog (https://www.gramene.org). LD decay was calculated in R using the following formula (Remington et al. 2001).
the expected value of r2 under drift-recombination equilibrium is denoted as E(r2) and is calculated as 1/(1 + C), where N represents the effective population size, c is the recombination fraction between sites, and C is determined by the equation 4Nc.
Results
Phenotypic traits and correlations
Protein, fiber, fat concentrations and 100-SW in 2018, 2022, and 2023 years of USDA Chickpea Core Collection and check cultivars is presented in Table S1. Seed protein concentrations were determined for the years 2018, 2022, and 2023 for 88 chickpea accessions and five check cultivars, while seed fiber and fat concentrations were determined only in the years 2018 and 2022. The seed protein concentrations varied between 16.3−21.4%, 17.4−23% and 19.9−25.2% for the years 2018, 2022, and 2023, respectively (Table 1). The fiber concentrations ranged between 13.3–20.8% in 2018 and between 14.6–18.2% in 2022. Fat concentrations ranged between 3.3–5.9% and 3.0–5.5% in 2018 and 2022, respectively. The 100-SW obtained in 2022 and 2023 showed a wide range in both years, recorded between 15−52.7 g, and 16.4−56.2 g, respectively (Table 1). The averages and standard errors with years and across years can also be seen in Table 1. According to analysis of variance, statistically significant (P < 0.01) differences were found between the accessions for each trait. Significant differences were found between environments at P < 0.01 for protein and fat traits, and significant differences at P < 0.05 for fiber and 100-SW traits. Genotype × Environment interaction was found to be significant in all traits except fat (Table 2).
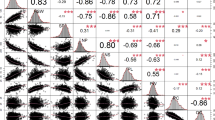
Pearson correlations were calculated between seed nutritional concentrations and 100-SW traits in the kabuli chickpea mini-core collection. There was a significant but not high positive relationship (r =.352**) between protein concentration and 100-SW. A high and significant negative relationship (r = − .747**) was observed between fat and fiber concentrations, and a positive but much lower significant relationship (r = .327**) was found between fat concentration and 100-SW (Table 3).
Principal component analysis
A Principal Components analysis (PCA) was run to determine population substructure in the panel and remove the effect from the GWAS. The PCA of the 88 USDA Chickpea Core Collection illustrated that the lines were divided into four clusters. One was composed exclusively of accessions from Iran; the second cluster contained only accessions from the Middle East; the third cluster consisted of two accessions from the Americas and several more from the Middle East; and the last and largest cluster was comprised of accessions from nearly every country in the study. Further structure within this fourth subpopulation was lacking, and the grouping of these lines into one cluster is likely the result of germplasm exchange between nations (Fig. S1).
Genome-wide association analysis
Utilizing genotyping by sequencing (GBS), a total of 165K SNPs were identified within the kabuli chickpea mini-core collection. Following the filtration of minor allele frequency to > 5%, this study utilized 113,512 SNPs distributed across eight chromosomes of the chickpea genome. The analysis of marker-trait associations employed the BLINK model, which was also compared with two alternative models: Mixed Linear Model (MLM) (Yu et al. 2006), and the Fixed and random model Circulating Probability Unification (FarmCPU) within the GAPIT package Version 3 (Huang et al. 2019). Notably, the BLINK model demonstrated a good fit of test statistics on Q-Q plots. Consequently, considering this favorable performance, BLINK was selected as the optimal model for association analysis in this study. Genome-wide association analysis identified 27 SNPs significantly associated with three seed nutritional concentrations and 100-SW across all eight chromosomes (Fig. 1; Table 4).
Joint Manhattan plot for the GWAS analysis of three seed nutritional traits (protein, fiber, and fat in 3, 2, and 2 years, respectively, and averaged over years) and 100 seed weight (100-SW) in 2 years and averaged over years using the BLINK model. Statistical significance threshold is shown with the green horizontal line; multiple associations to the same SNP are shown with vertical lines
Three marker-trait associations (MTAs) were discovered for seed protein concentration on chromosomes 1, 5, and 7, explaining a phenotypic variation range of 10.4–29.3% in 2018. Two MTAs were identified for protein concentration on chromosomes 4 and 7 based on the three-year average (multiyear). The MTA on chromosome 7 was common to both (Fig. 2).
Six MTAs were identified for fiber concentration for 2018, and four for the average over multiple years. A total of 10 MTAs associated to fiber concentration were distributed to all chromosomes except 7th chromosome explaining a phenotypic variation range of 2.9–38.6% (Fig. 3; Table 4). None of the MTAs were found in common between 2018 and the average over years.
A total of seven MTAs associated with fat concentration were identified on chromosomes 1, 2 and 4. Two MTAs located on chromosomes 1 and 2 were common to all three environments (2018, 2022, and multiyear). The MTAs explained 9.3–54.8% of the phenotypic variation within the three environments (Fig. 4; Table 4).
A total of five MTAs associated with 100-SW were found: three MTAs on chromosomes 1, 2, and 4 in 2018, one MTA on chromosome 1 in 2023, and one MTA on chromosome 2 in the multiyear analysis. The five MTAs identified for 100-SW explained between 16.1–45.4% of phenotypic variation (Table 4; Fig. 5). These MTAs were not found in common between years or the average over years.
Discussion
Chickpeas have gained attention as a versatile and nutritious food source. Rich in protein, fiber, vitamins, and minerals, chickpeas offer a range of health benefits. Chickpeas are an excellent source of plant-based protein, essential for muscle growth and overall health (Jukanti et al. 2012). Chickpeas are low in saturated fat and high in unsaturated fats and high in dietary fiber and contribute to heart and digestive health and can help prevent conditions such as colon diseases from constipation to cancer (Gill et al. 2021; Gupta et al. 2017; Mugabe et al. 2023). Chickpeas contain essential vitamins and minerals, including folate, iron, phosphorus, and manganese, contributing to overall well-being (Derbyshire and Delange 2021). Moreover, they are affordable and have a long shelf life, making them a viable option for improving food security. As a cost-effective source of nutrition, chickpeas can play a crucial role in providing accessible and nutritious food to vulnerable populations. Governments, non-governmental organizations, breeders, and international agencies can play a pivotal role in promoting chickpea cultivation and consumption. This includes investing in breeding programs and agricultural practices that enhance chickpea production, educating communities about the nutritional benefits of chickpeas, and integrating chickpeas into food aid programs.
This study conducted on a kabuli chickpea mini-core collection revealed substantial variation in seed nutritional concentrations and 100-seed weight (100-SW) across multiple years. Notably, the seed protein concentration showed a wide range from 16.3 to 25.2% over the years 2018, 2022, and 2023. This variability, also identified in other studies (Cobos et al. 2009; Farida Traore et al. 2022), underscores the potential for selective breeding to enhance protein concentration in chickpea accessions. We observed a negative correlation between fat and fiber concentrations, a positive relationship of protein and fat concentrations, and a positive correlation between protein concentration and 100-SW. This all indicates that the seed has finite storage capacity for nutrients, and an increase in one may lead to a decrease in others, unless the seed size is increased. In addition, the source-sink relationship of photosynthates and upstream metabolites that lead to the creation of these seed nutrients is often competitive, thus resulting in an intricate interplay of these traits in chickpeas (Ereifej et al. 2001; Özer et al. 2010). This will affect the breeding of these traits, as an increase in one may lead to a decrease in another.
The GBS used in this study identified a sufficient number of SNPs to enable GWAS analysis, and the BLINK model demonstrated a good fit of test statistics on Q-Q plots in identifying marker-trait associations (MTAs). The identification of marker-trait associations (MTAs) for seed protein, fiber and fat concentration, and 100-SW in kabuli chickpeas provides useful insights into the genetic regulation of these important nutritional traits. The identification of 27 significantly associated SNPs linked to seed nutritional concentrations and 100-SW across all eight chromosomes reaffirms the polygenic nature of these traits in chickpeas (Upadhyaya et al. 2016; Karaca et al. 2019; Srungarapu et al. 2022). In other studies, in chickpea, Samineni et al. (2022) reported 46 MTAs for protein concentration, Upadhyaya et al. (2016) found seven MTAs, and Srungarapu et al. (2022) identified four. The MTA identified on chromosome 4 by Srungarapu et al. (2022) was consistent in both years they studied and appeared to be close to the MTA on chromosome 4 (4,583,239 bp) identified in our study, within the linkage disequilibrium (LD) decay distance reported in chickpea (Srungarapu et al. 2022). The variation in numbers of associated SNPs and the genomic regions identified across studies may reflect the diverse germplasm used, different environmental conditions and the genotyping methods employed.
Looking at other traits in the current study, the 10 MTAs discovered for fiber concentration do not seem to be in common with the only other GWAS of this trait published to date, who reported two MTAs (Mugabe et al. 2023). The current study identified seven MTAs for fat concentration, two of which were very consistent across both years and the average over years and explained a considerable proportion of the phenotypic variation (9.3–54.8%). These two MTAs thus show a substantial genetic impact on fat concentration that is stable across environments. Multiple genomic regions were also found to be associated with seed fat concentration by Mugabe et al. (2023) but not in the same genomic locations as those reported here, and not of such large effect. Finally, multiple MTAs were identified for 100-SW in the current study, as well as other GWAS (Srungarapu et al. 2022; Thudi et al. 2023) and QTL studies (Kujur et al. 2014; Bajaj et al. 2015; Das et al. 2015; Verma et al. 2015; Roorkiwal et al. 2016; Wang et al. 2019). Co-localization of associated genomic regions between studies is rare, emphasizing the quantitative nature of the trait, but fairly high phenotypic effect and stability across environments within a study suggest that these QTL are all potentially useful to increase the trait in a breeding study, possibly by marker assisted selection to pyramid them into one genetic background.
To understand potential mechanisms of action of the MTAs identified in this study, candidate genes within a window of 30 kb were sought for the 22 MTA for seed protein, fiber, fat concentrations and 100-SW. Detailed information on the 31 candidate genes thus identified is presented in Table 5. The SNP (SCA1_V1.0_KABULI_455050) on chromosome 1 associated with protein concentration is 1586 bp away from a potential candidate gene whose homolog, AT1G22940 (TH1), has biological functions in the thiamine biosynthetic process, metabolic process, and phosphorylation (Strobbe et al. 2021). In a study reported by Rohi et al. (2013) determining B vitamins and protein in wheat flour, a strong positive correlation (r = 0.56) was found between thiamine and protein concentration in whole wheat flour, although this was not found in a smaller study of chickpea seed components (Roorkiwal et al. 2016). The same SNP on chromosome 1 for protein concentration is linked to two other genes (Table 5), but how they may be involved in seed protein concentration is unclear. The SNP associated with protein on chromosome 5 (SCA5_V1.0_KABULI_39352536) is in the potential candidate gene (0 bp) whose homolog, AT5G56480 (END2), functions in lipid-transfer and binding and is part of the seed storage 2S albumin superfamily, which has involvement in protein localization. In a GWAS analysis of pea (Pisum sativum), a homolog of the same gene END2 was found to be significantly associated with seed fat concentration (unpublished data). Thus, this gene may influence protein concentration directly or indirectly by influencing a potentially competing seed component, lipid, and fat concentration. The SNP (SCA7_V1.0_KABULI_2145652) on chromosome 7 for protein concentration is 5 kb away from a potential candidate gene whose homolog, AT1G59990 (HEAVY SEED3-HS3), is responsible for seed size in Arabidopsis and tends to be highly expressed in developing seeds (Kanai et al. 2013). This gene may consequently help to explain some of the significant correlation between protein concentration and 100-SW in the current study (Table 3) and suggests the potential for developing genotypes with high protein concentration and larger seed size simultaneously (Panthee et al. 2005; Kulwal and Mhase 2017; Samineni et al. 2022). Two additional genes are associated with the SNP on chromosome 7 (Table 5) and the homolog of one, AT4G23850 (LACS4) is known to affect lipid and fatty acid metabolism, suggesting another mechanism affecting protein levels indirectly. Finally, the SNP (SCA4_V1.0_KABULI_4583239) on chromosome 4 for protein concentration is in the potential candidate gene (0 bp) whose homolog, AT1G61290 (SYP124), regulates protein transport and pollen tube growth, which may not indicate involvement in seed protein concentration. This SNP is also linked (2264) to a gene whose homolog AT2G05990 again involves lipid and fatty acid metabolism. The potential interplay of these two seed components should be more closely studied.
Altogether, 10 MTAs related to fiber concentration and accounted for a fairly high phenotypic variation range of 2.9% to 38.6%. Dietary fibers are mostly indigestible complex starches and carbohydrates, often components of plant cell walls (cellulose, hemicellulose, and pectin), and polysaccharides. The fiber-associated SNP (SCA1_V1.0_KABULI_41348657) on chromosome 1 with a particularly significant p value (2.84E-15) is linked (4799 bp) to a gene whose homolog (AT3G14410) is involved in nucleotide, carbohydrate, and sugar transport, and has been related to glycosylation and polysaccharide biosynthesis (Reyes and Orellana 2008). The fiber SNP (SCA4_V1.0_KABULI_1475346) located on chromosome 4 is within 1615 bp of a regulation of starch biosynthetic gene model (AT2G41680-NTRC). Overexpression of NTRC led to an increase in the accumulation of starch in leaves exposed to light (Toivola et al. 2013). For other fiber-related SNPs, candidate genes were identified within a range of 0 bp–15 kb (Table 5) that were involved with various functions, including mRNA catabolic processes, lipid storage, chromosome condensation, auxin-mediated signaling pathways, regulation of gene expression, sphingolipid metabolic processes, and response to abscisic acid.
The three MTAs identified for fat concentration were particularly significant, and those that were identified in multiple years (on chromosomes 1 and 2) explained between one third and one half of the phenotypic variance (Table 4). The SNP on chromosome 1 (SCA1_V1.0_KABULI_23894664) was 18 kb distant from a gene whose homolog AT4G32770 (VTE1), regulates fatty acid metabolic process and vitamin E biosynthesis in Arabidopsis (Porfirova et al. 2002). The SNP on chromosome 4 (SCA4_V1.0_KABULI_23894664) is 13,168 bp away from a potential candidate gene whose homolog, AT3G12120 (FAD2), is known to be involved in a lipid metabolic process, unsaturated fatty acid biosynthesis, fatty acid metabolic process, and omega-6 fatty acid biosynthesis (Lakhssassi et al. 2017). In a study on the genome-wide identification of genes encoding FAD (fatty acid desaturase) proteins in chickpea, Saini and Kumar (2019) identified a total of 18 FAD genes in both desi and kabuli chickpea genomes, including the same FAD2 found in the current study (Ca_14188). This gene is crucial for lipid and fatty acid-related processes, and an excellent candidate for marker-assisted selection.
The five SNPs associated with 100-SW were linked to six genes primarily involved with embryo development and stress responses. Of particular interest, SNP (SCA1_V1.0_KABULI_1915412) on chromosome 1 for 100-SW is 4931 bp away from the gene whose homolog, AT1G61590, is a protein kinase family protein, having biological function on protein phosphorylation, defense response, and regulation of lignin biosynthetic process. AT1G61590 is a DELLA gene, and these genes were found to regulate seed size in Arabidopsis (Gomez et al. 2023). The SNP SCA4_V1.0_KABULI_32380371 on chromosome 4 is linked to a gene whose homolog, AT3G50870 (MNP) is a GATA type zinc finger transcription factor family protein involved in embryo development and seed dormancy; SNP SCA2_V1.0_KABULI_31697291 on chromosome 2 is linked to a potential candidate gene whose homolog, AT4G13940 (HOG1), is also associated with embryo development and seed dormancy. Godge et al. (2008) reported that HOG1 had a significant influence on plant and seed yield parameters in petunia and identified HOG1 as a key gene with potential in regulating seed and plant development and cytokinin signaling, proposing a promising strategy for improving yield in various crop species through a combination of genetic manipulation and conventional breeding.
Conclusions
Chickpea, a valuable and nutritious food source with diverse health benefits, may be further improved with the genetic information presented in this study. Four phenotypically promising chickpea accessions (CSP-52, CSP-59, CSP-73, and CSP-74) have been identified. These selected accessions exhibit a protein concentration of 22% or higher, a fiber concentration of 15% or higher, and a 100-seed weight of 45 g or more. The GWAS revealed 27 SNPs across all eight chromosomes significantly associated with protein, fiber, fat concentrations and 100-seed weight, with varying phenotypic effects, across different chromosomes and environments. These were linked to 31 candidate genes that may help explain molecular mechanisms underlying these important seed traits. These genes and linked SNPs may also offer valuable tools once validated for breeders in optimizing crop nutritional profiles through marker-assisted selection.
References
Anbessa Y, Warkentin T, Vandenberg A, Bandara M (2006) Heritability and predicted gain from selection in components of crop duration in divergent chickpea cross populations. Euphytica 152:1–8
Anbessa Y, Taran B, Warkentin TD, Tullu A, Vandenberg A (2009) Genetic analyses and conservation of QTL for ascochyta blight resistance in chickpea (Cicer arietinum L.). Theor Appl Genet 119:757–765
Bajaj D, Upadhyaya HD, Khan Y et al (2015) A combinatorial approach of comprehensive QTL-based comparative genome mapping and transcript profiling identified a seed weight-regulating candidate gene in chickpea. Sci Rep 5(1):9264
Barnett DW, Garrison EK, Quinlan AR, Strömberg MP, Marth GT (2011) BamTools: a C++ API and toolkit for analyzing and managing BAM files. Bioinform 27(12):1691–1692
Beermann C (2022) Influence of micro-and macronutrients on the clonal phase of the adaptive immune response. In: Beerman C (ed) Food and the Immune System: Molecular Mechanisms and Nutritional Relevance in Health and Disease, 1st edn. Springer, Cham, pp 127–150
Birketvedt GS, Shimshi M, Erling T, Florholmen J (2005) Experiences with three different fiber supplements in weight reduction. Med Sci Monit 11(1):15–8
Cobos MJ, Winter P, Kharrat M, Cubero JI, Gil J, Millan T, Rubio J (2009) Genetic analysis of agronomic traits in a wide cross of chickpea. Field Crops Res 111(1–2):130–136
Das S, Upadhyaya HD, Bajaj D et al (2015) Deploying QTL-seq for rapid delineation of a potential candidate gene underlying major trait-associated QTL in chickpea. DNA Res 22(3):193–203
Derbyshire E, Delange J (2021) The nutritional value of whole pulses and pulse fractions. In: Tiwari BK, Gowen A, McKenna B (eds) Pulse foods, 2nd edn. Elsevier, Academic Press, pp 9–29
Eker T, Sari D, Sari H, Tosun HS, Toker C (2022) A kabuli chickpea ideotype Sci Rep 12(1):1611
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PloS one 6(5):e19379
Ereifej KI, Al-Karaki GN, Hammouri MK (2001) Seed chemical composition of improved chickpea cultivars grown under semiarid Mediterranean conditions. Int J Food Prop 4(2):239–246
FAO, Ifad, UNICEF, WFP, WHO. (2020). The state of food security and nutrition in the world 2020 transforming food systems for affordable healthy diets. IEEE Journal of Selected Topics in Applied Earth Observations and Remote Sensing
FAO (2023). Online available at: http://faostat.fao.org (Accessed 06 June 2023), Rome
Farida Traore F, El-Baouchi A, En-Nahli Y (2022) Exploring the genetic variability and potential correlations between nutritional quality and agro-physiological traits in kabuli chickpea germplasm collection (Cicer arietinum L.). Front Plant Sci 13:905320
Garrison E, Marth G (2012) Haplotype-based variant detection from short-read sequencing. arXiv preprint arXiv:1207.3907
Gaur R, Verma S, Pradhan S, Ambreen H, Bhatia S (2020) A high-density SNP-based linkage map using genotyping-by-sequencing and its utilization for improved genome assembly of chickpea (Cicer arietinum L.). Funct Integr Genomics 20:763–773
Gill SK, Rossi M, Bajka B, Whelan K (2021) Dietary fibre in gastrointestinal health and disease. Nat Rev Gastroenterol Hepatol 18(2):101–116
Godge MR, Kumar D, Kumar PP (2008) Arabidopsis HOG1 gene and its petunia homolog PETCBP act as key regulators of yield parameters. Plant Cell Rep 27:1497–1507
Gomez MD, Cored I, Barro-Trastoy D, Sanchez-Matilla J, Tornero P, Perez-Amador MA (2023) DELLA proteins positively regulate seed size in Arabidopsis. Development 150(15):201853
GRIN-Global. https://ars-grin.gov/gringlobal/search
Gupta RK, Gupta K, Sharma A, Das M, Ansari IA, Dwivedi PD (2017) Health risks and benefits of chickpea (Cicer arietinum) consumption. J Agric Food Chem 65(1):6–22
Huang M, Liu X, Zhou Y, Summers RM, Zhang Z (2019) BLINK: a package for the next level of genome-wide association studies with both individuals and markers in the millions. Gigascience 8(2):154
Iqbal A, Khalil IA, Ateeq N, Khan MS (2006) Nutritional quality of important food legumes. Food Chem 97(2):331–335
Jadhav AA, Rayate SJ, Mhase LB et al (2015) Marker-trait association study for protein content in chickpea (Cicer arietinum L.). J Genet 94:279–286
Jain M, Misra G, Patel RK et al (2013) A draft genome sequence of the pulse crop chickpea (Cicer arietinum L.). The Plant J 74(5):715–729
Jones, DB. (1931). Factors for converting percentages of nitrogen in foods and feeds into percentages of proteins 183. US Department of Agriculture
Jukanti AK, Gaur PM, Gowda CLL, Chibbar RN (2012) Nutritional quality and health benefits of chickpea (Cicer arietinum L.): a review. Br J Nutr 108(S1):11–26
Kanai M, Hayashi M, Kondo M, Nishimura M (2013) The plastidic DEAD-box RNA helicase 22, HS3, is essential for plastid functions both in seed development and in seedling growth. Plant Cell Physiol 54(9):1431–1440
Karaca N, Ates D, Nemli S et al (2019) Genome-wide Association studies of protein, lutein, vitamin C, and fructose concentration in wild and cultivated chickpea seeds. Crop Sci 59(6):2652–2666
Kujur A, Bajaj D, Saxena MS (2014) An efficient and cost-effective approach for genic microsatellite marker-based large-scale trait association mapping: identification of candidate genes for seed weight in chickpea. Mol Breed 34:241–265
Kulwal PL, Mhase LB (2017) Protein content exhibits a significant positive correlation with seed weight in chickpea germplasm collection. Plant Genet Resour 15(3):283–285
Kumar Y, Kwon SJ, Coyne CJ, Hu J, Grusak MA, Kisha TJ, McGee RJ, Sarker A (2014) Target region amplification polymorphism (TRAP) for assessing genetic diversity and marker-trait associations in chickpea (Cicer arietinum L.) germplasm. Genet Resour Crop Evol 61:65–977
Lakhssassi N, Zhou Z, Liu S, Colantonio V, AbuGhazaleh A, Meksem K (2017) Characterization of the FAD2 gene family in soybean reveals the limitations of gel-based TILLING in genes with high copy number. Front Plant Sci 8:324
Leonforte A, Sudheesh S, Cogan NO et al (2013) SNP marker discovery, linkage map construction and identification of QTLs for enhanced salinity tolerance in field pea (Pisum sativum L.). BMC Plant Biol 13(1):1–14
Liu S, Stampfer MJ, Hu FB et al (1999) Whole-grain consumption and risk of coronary heart disease: results from the Nurses’ Health Study. Am J Clin Nutr 70(3):412–419
Meyers LD, Hellwig JP, Otten JJ (2006) Dietary reference intakes: the essential guide to nutrient requirements. National Academies Press
Mugabe D, Frieszell CM, Warburton ML, Coyne CJ, Sari H, Uhdre R, Wallace L, Ma Y, Zheng P, McGee RJ, Ganjyal GM (2023) Kabuli chickpea seed quality diversity and preliminary genome-wide association study identifies markers and potential candidate genes. Agrosyst Geosci Environ 6(4):e20437
Özer S, Karaköy T, Toklu F, Baloch FS, Kilian B, Özkan H (2010) Nutritional and physicochemical variation in Turkish kabuli chickpea (Cicer arietinum L.) landraces. Euphytica 175:237–249
Panthee DR, Pantalone VR, West DR, Saxton AM, Sams CE (2005) Quantitative trait loci for seed protein and oil concentration, and seed size in soybean. Crop Sci 45(5):2015–2022
Petruzziello L, Iacopini F, Bulajic M, Shah S, Costamagna G (2006) Uncomplicated diverticular disease of the colon. Aliment Pharmacol Ther 23(10):1379–1391
Porfirova S, Bergmüller E, Tropf S, Lemke R, Dörmann P (2002) Isolation of an Arabidopsis mutant lacking vitamin E and identification of a cyclase essential for all tocopherol biosynthesis. PNAS 99(19):12495–12500
Remington DL, Thornsberry JM, Matsuoka Y et al (2001) Structure of linkage disequilibrium and phenotypic associations in the maize genome. PNAS 98(20):11479–11484
Reyes F, Orellana A (2008) Golgi transporters: opening the gate to cell wall polysaccharide biosynthesis. Curr Opin Plant Biol 11(3):244–251
Rohi M, Pasha I, Butt MS, Nawaz H (2013) Variation in the levels of B-vitamins and protein content in wheat flours. Pak J Nutr 12(5):441
Roorkiwal M, Rathore A, Das RR et al (2016) Genome-enabled prediction models for yield related traits in chickpea. Front Plant Sci 7:1666
Rytter MJH, Kolte L, Briend A, Friis H, Christensen VB (2014) The immune system in children with malnutrition—a systematic review. PloS One 9(8):e105017
Sab S, Lokesha R, Mannur DM et al (2020) Genome-wide SNP discovery and mapping QTLs for seed iron and zinc concentrations in chickpea (Cicer arietinum L.). Front Nutr 7:559120
Saini R, Kumar S (2019) Genome-wide identification, characterization and in-silico profiling of genes encoding FAD (fatty acid desaturase) proteins in chickpea (Cicer arietinum L.). Plant Gene 18:100180
Samineni S, Mahendrakar MD, Hotti A, Chand U, Rathore A, Gaur PM (2022) Impact of heat and drought stresses on grain nutrient content in chickpea: genome-wide marker-trait associations for protein. Fe and Zn Environ Exp Bot 194:104688
Sánchez-Vioque R, Clemente A, Vioque J, Bautista J, Millán FJFC (1999) Protein isolates from chickpea (Cicer arietinum L.): chemical composition, functional properties and protein characterization. Food Chem 64(2):237–243
Simon CJ, Hannan RM (1995) Development and use of core subsets of cool-season food legume germplasm collections. HortSci 30(4):907C–907
Singh U (1985) Nutritional quality of chickpea (Cicer arietinum L.): current status and future research needs. Plant Foods Hum Nutr 35:339–351
Srungarapu R, Mahendrakar MD, Mohammad LA et al (2022) Genome-wide association analysis reveals trait-linked markers for grain nutrient and agronomic traits in diverse set of chickpea germplasm. Cells 11(15):2457
Stephens A, Lombardi M, Cogan NO, Forster JW, Hobson K, Materne M, Kaur S (2014) Genetic marker discovery, intraspecific linkage map construction and quantitative trait locus analysis of Ascochyta blight resistance in chickpea (Cicer arietinum L.). Mol Breed 33:297–313
Strobbe S, Verstraete J, Stove C, Van Der Straeten D (2021) Metabolic engineering provides insight into the regulation of thiamin biosynthesis in plants. Plant Physiol 186(4):1832–1847
Thudi M, Samineni S, Li W, Boer MP, Roorkiwal M, Yang Z, Ladejobi F, Zheng C, Chitikineni A, Nayak S, He Z (2023) Whole genome resequencing and phenotyping of MAGIC population for high resolution mapping of drought tolerance in chickpea. The Plant Genome 30:e20333
Toivola J, Nikkanen L, Dahlström KM, Salminen TA, Lepistö A, Vignols HF, Rintamäki E (2013) Overexpression of chloroplast NADPH-dependent thioredoxin reductase in Arabidopsis enhances leaf growth and elucidates in vivo function of reductase and thioredoxin domains. Front Plant Sci 4:389
Tosh SM, Yada S (2010) Dietary fibres in pulse seeds and fractions: characterization, functional attributes, and applications. Food Res Int 43(2):450–460
Upadhyaya HD, Bajaj D, Das S et al (2016) Genetic dissection of seed-iron and zinc concentrations in chickpea. Sci Rep 6(1):1–12
USDA (2019) https://fdc.nal.usda.gov/fdc-app.html#/food-details/173756/nutrients
van der Maesen LJG (1972) Cicer L., a monograph of the genus, with special reference to the chickpea (Cicer arietinum L.), its ecology and distribution. Mendelingen Landbouwhogeschool. Wageningen, Wageningen, The Netherlands 1–341
Vandemark GJ, Grusak MA, McGee RJ (2018) Mineral concentrations of chickpea and lentil cultivars and breeding lines grown in the US Pacific Northwest. Crop J 6(3):253–262
Varshney RK, Song C, Saxena RK, Azam S, Yu S, Sharpe AG, Cannon S, Baek J, Rosen BD, Tar'an B, Draft Millan T (2013) Draft genome sequence of chickpea (Cicer arietinum) provides a resource for trait improvement. Nat Biotech 31(3):240–246
Verma S, Gupta S, Bandhiwal N, Kumar T, Bharadwaj C, Bhatia S (2015) High-density linkage map construction and mapping of seed trait QTLs in chickpea (Cicer arietinum L.) using genotyping-by-sequencing (GBS). Sci Rep 5(1):17512
Wang J, Zhang Z (2021) GAPIT version 3: boosting power and accuracy for genomic association and prediction. GPB 19(4):629–640
Wang X, Gao W, Zhang J, Zhang H, Li J, He X, Ma H (2010) Subunit, amino acid composition and in vitro digestibility of protein isolates from Chinese kabuli and desi chickpea (Cicer arietinum L.) cultivars. Food Res Int 43(2):567–572
Wang R, Gangola MP, Irvine C, Gaur PM, Båga M, Chibbar RN (2019) Co-localization of genomic regions associated with seed morphology and composition in a desi chickpea (Cicer arietinum L.) population varying in seed protein concentration. TAG 132:1263–1281
WHO World Health Organization (2021) Levels and trends in child malnutrition UNICEF
Wu G (2016) Dietary protein intake and human health. Food Funct 7(3):1251–1265
Yu J, Pressoir G, Briggs WH et al (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38(2):203–208
Acknowledgements
This study was funded by USDA CRIS project 2090-21000-037-00D. We would like to thank the Scientific and Technological Research Council of Turkiye (TUBITAK) for funding Dr. Hatice Sari. Data analysis assistance of Dr. Ping Zheng gratefully acknowledged. Field work assistance of Kurt Tetrick gratefully acknowledged
Author information
Authors and Affiliations
Contributions
HS: Formal analysis; methodology; writing—original draft; writing—review and editing. RU: Data curation; validation; writing—review and editing. LW: Investigation; writing—review and editing. CJC: Conceptualization; project administration; resources; supervision; writing—review and editing. BB: Validation; writing—review and editing. ZZ: Data curation; methodology; validation; writing—review and editing. MR: Validation; writing—review and editing. AK: Validation; writing—review and editing. MLW: Supervision; conceptualization; methodology; writing—review and editing. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sari, H., Uhdre, R., Wallace, L. et al. Genome-wide association study in Chickpea (Cicer arietinum L.) for yield and nutritional components. Euphytica 220, 84 (2024). https://doi.org/10.1007/s10681-024-03338-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-024-03338-x