Abstract
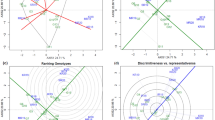
Wheat (Triticum aestivum L.) plays a significant role in the global agricultural economy. Breeding programs need to identify environments where genotype discrimination allows effective selection, as this is crucial for developing more productive wheat genotypes through genetic breeding. Factor analytic models can be applied to the analysis of the multi-environment trial. This approach is based on multivariate models that utilize factor analysis (FA) structures and consider environments (E), genotypes (G), and G × E effects. This study evaluated eight wheat genotypes in fifteen environments in southern Brazil to understand the G × E interactions of wheat genotypes. The primary objectives were identifying stable and responsive genotypes across a set of environments (mega-environments) and determining the main environmental covariates, such as temperature, solar radiation, precipitation, relative humidity, wind speed, and altitude, that can explain G × E interactions. The Bayesian factor analytic model was applied to the grain yield as the response variable. The model with four factors, FA(4), provided the best fit, allowing the creation of four mega-environments. Using latent regressions, we identified genotypes that are suitable for each mega-environment, as they demonstrate responsiveness and wide adaptability to those specific groups of environments. The correlations between the environmental loadings and the environmental covariates revealed that precipitation, maximum temperature, and wind speed negatively affected yield in these environments. On the other hand, the altitude covariate positively affected the grain yield in these environments. The occurrence of four mega-environments in this study suggests that the same information about genotypes can be obtained from fewer environments, indicating the possibility of reducing costs in breeding programs.


Similar content being viewed by others
References
Araghi A, Maghrebi M, Olesen JE (2022) Effect of wind speed variation on rainfed wheat production evaluated by the CERES-Wheat model. Int J Biometeorol 66:225–233. https://doi.org/10.1007/s00484-021-02209-7
Burgueño J, Crossa J, Cornelius PL, Yang R (2008) Using factor analytic models for joining environments and genotypes without crossover genotype × environment interaction. Crop Sci 48:1291–1305. https://doi.org/10.2135/cropsci2007.11.0632
CONAB (2017) A cultura do trigo. https://www.conab.gov.br/uploads/arquivos/17_04_25_11_40_00_a_cultura_do_trigo_versao_digital_final.pdf. Accessed 5 May 2022
CONAB (2023) Acompanhamento da Safra Brasileira de Grãos - safra 2023/24. https://www.conab.gov.br/info-agro/safras/graos/boletim-da-safra-de-graos. Accessed 5 May 2022
Cooper M, Hammer GL (1996) Plant adaptation and crop improvement. IRRI
Cruz CD, Regazzi AJ, Carneiro PCS (2012) Modelos biométricos aplicados ao melhoramento. UFV, Viçosa
Curtis BC, Rajaram S, Gómez Macpherson H (2002) Bread wheat: improvement and production. Food and Agriculture Organization of the United Nations (FAO)
de Los Campos G, Gianola D (2007) Factor analysis models for structuring covariance matrices of additive genetic effects: a Bayesian implementation. Genet Sel Evol 39:481. https://doi.org/10.1186/1297-9686-39-5-481
FAO (2021) Food and Agriculture Organization of the United Nations. https://www.fao.org/home/en. Accessed 5 May 2022
Geweke J (1992) Evaluating the accuracy of sampling-based approaches to the calculations of posterior moments. Bayesian Stat 4:641–649
Hox J, McNeish D (2020) Small samples in multilevel modeling. Small sample size Solutions 215–225
Kohli MM, McMahon MA (1988) A perspective of research needs for nonirrigated tropical conditions. In: Wheat production constraints in tropical environments. Chiang Mai, Thailand. Accessed19–23 Jan 1987
Niu L, Feng S, Ding W, Li G (2016) Influence of speed and rainfall on large-scale wheat lodging from 2007 to 2014 in China. PLoS ONE 11:e0157677. https://doi.org/10.1371/journal.pone.0157677
Nuvunga JJ, da Silva CP, de Oliveira LA et al (2019) Bayesian factor analytic model: an approach in multiple environment trials. PLoS ONE 14:e0220290. https://doi.org/10.1371/journal.pone.0220290
Oliveira ICM, Guilhen JHS, de Oliveira Ribeiro PC et al (2020) Genotype-by-environment interaction and yield stability analysis of biomass sorghum hybrids using factor analytic models and environmental covariates. F Crop Res 257:107929. https://doi.org/10.1016/j.fcr.2020.107929
Ortiz R, Sayre KD, Govaerts B et al (2008) Climate change: Can wheat beat the heat? Agric Ecosyst Environ 126:46–58. https://doi.org/10.1016/j.agee.2008.01.019
Plummer M, Best N, Cowles K, Vines K (2006) CODA: convergence diagnosis and output analysis for MCMC. R News 6:7–11
RCBPTT (2019) 13a Reunião da Comissão Brasileira de Pesquisa de Trigo e Triticale
Resende MDV (2007) Matemática e estatística na análise de experimentos e no melhoramento genético. Embrapa Florestas
Rodrigues O (2002) Efeito de redutor de crescimento cycocel e de altas doses de adubação nitrogenada em trigo. Embrapa Trigo
Sae-Lim P, Komen H, Kause A, Mulder HA (2014) Identifying environmental variables explaining genotype-by-environment interaction for body weight of rainbow trout (Onchorynchus mykiss): reaction norm and factor analytic models. Genet Sel Evol 46:16. https://doi.org/10.1186/1297-9686-46-16
Smith AB, Cullis BR (2018) Plant breeding selection tools built on factor analytic mixed models for multi-environment trial data. Euphytica 214:143. https://doi.org/10.1007/s10681-018-2220-5
Smith AB, Ganesalingam A, Kuchel H, Cullis BR (2015) Factor analytic mixed models for the provision of grower information from national crop variety testing programs. Theor Appl Genet 128:55–72. https://doi.org/10.1007/s00122-014-2412-x
Spiegelhalter DJ, Best NG, Carlin BP, van der Linde A (2002) Bayesian measures of model complexity and fit. J R Stat Soc Ser B Stat Methodol 64:583–639. https://doi.org/10.1111/1467-9868.00353
Thomson AM, Brown RA, Ghan SJ et al (2002) Elevation dependence of winter wheat production in eastern Washington State with climate change: a methodological study. Clim Change 54:141–164
Tomar V, Singh D, Dhillon GS et al (2021) Increased predictive accuracy of multi-environment genomic prediction model for yield and related traits in spring wheat (Triticum aestivum L.). Front Plant Sci 12:720123. https://doi.org/10.3389/fpls.2021.720123
Torres LG, Rodrigues MC, Lima NL et al (2018) Multi-trait multi-environment Bayesian model reveals G × E interaction for nitrogen use efficiency components in tropical maize. PLoS ONE 13:e0199492. https://doi.org/10.1371/journal.pone.0199492
van de Schoot R, Depaoli S, King R et al (2021) Bayesian statistics and modelling. Nat Rev Methods Prim 1:1. https://doi.org/10.1038/s43586-020-00001-2
Acknowledgements
The authors are grateful for the financial support granted by Fundação de Amparo à Pesquisa do Estado de Minas Gerais—FAPEMIG, Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—CAPES, and Conselho Nacional de Desenvolvimento Científico e Tecnológico—CNPq.
Funding
The Fundação de Amparo à Pesquisa do Estado de Minas Gerais—FAPEMIG, the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—CAPES and the Conselho Nacional de Desenvolvimento Científico e Tecnológico—CNPq.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Consent for publication
The authors have no financial or proprietary interests in any material discussed in this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Azevedo, C.F., Barreto, C.A.V., Nascimento, M. et al. Genotype-by-environment interaction of wheat using Bayesian factor analytic models and environmental covariates. Euphytica 219, 95 (2023). https://doi.org/10.1007/s10681-023-03223-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-023-03223-z