Abstract
Leaf rust is a common and costly wheat disease. The introduction of resistance genes derived from wheat wild relatives is one way to improve disease control. However, due to the rapid development of virulence in the casual pathogen Puccinia triticina, the long-term effectiveness of some leaf rust resistance (Lr) genes is questionable. To assess this, we examined 130 accessions of 13 wheat wild relative species from the Aegilops genus. The presence of 9 Lr genes was determined by PCR using primers complementary to Lr gene markers. The studied individuals were exposed to artificial inoculation with P. triticina under greenhouse and field conditions, respectively. Based on infection rate, the degree of resistance for each individual was estimated. The relationship between the presence of Lr genes and the degree of manifested resistance was assessed using canonical correlation analysis (CCA) and analysis of variance (ANOVA). Our results indicate the effective participation of Lr9 in seedling resistance while Lr58 is involved in adult plant resistance.
Similar content being viewed by others
Introduction
Wheat rust diseases pose a serious threat to wheat production worldwide. Although not as devastating as stem rust or stripe rust, the global and frequent occurrence of leaf rust makes it responsible for considerable annual wheat losses (Huerta-Espino et al., 2011). This fungal disease is caused by Puccinia triticina Eriks, an obligate biotrophic parasite with a lifecycle that consists of two phases: i) an asexual cycle (major phase) completed on wheat, and ii) a sexual cycle (minor phase) completed on an alternate host, Thalictrum sp., that gives the pathogen the advantage of acquiring genetic variability through recombination (Kolmer et al., 2013).
Various strategies have been developed to counter the adverse effects of leaf rust. However, agricultural methods, such as crop rotation, deep tillage and adjusting the sowing/harvesting time (Krupinsky et al., 2002), as well as use of chemical applications are not successful in the long run. Although fungicide use has enabled farmers to intensify production, overall crop losses have not declined for decades (Oerke, 2006). Thus, the development of genetic resistance is now seen as the most efficient, economical and environment- friendly approach.
There are two types of host genetic resistance against P. triticina. The first is seedling resistance, which is expressed at all growth stages and is typically controlled by single genes that have a major effect. This type of resistance is generally qualitative and triggers a hypersensitive response that results in necrosis and prevents the pathogen from spreading. The second is adult plant resistance (APR), which occurs at a post-seedling stage and is commonly quantitative. The interplay of both types of resistance genes creates the overall host resistance to the pathogen.
To date, more than 80 leaf rust resistance (Lr) genes providing either seedling or adult plant resistance have been reported. However, the presence of a known Lr gene does not always guarantee resistance, with several studies documenting loss of efficiency over time (Kolmer, 2019; McCallum et al., 2021; Park et al., 2002). While single-gene resistance provides mostly short-term effectiveness, long term resistance is influenced by the specific combination of multiple resistance genes in the host (Nelson et al., 2018). For this reason, resistance breeding is an ongoing process that systematically tests wild relatives, landraces and other germplasm to identify new sources of genetic resistance to a successful pathogen. Aegilops, a genus of annual grass plants, is genetically related to bread wheat and carries Lr genes. It consists of 23 species with C, M, N, S, U and D genomes. The D genome, derived from Ae. tauschii, and the B genome, derived from a species closely related to Ae. speltoides, are part of the bread wheat (Triticum aestivum) hexaploid genome AABBDD. A notable number of Lr genes have been identified in various species of the Aegilops genus (Prasad et al., 2020). Since the first introgression of Lr9 from Ae. umbellulata into the wheat genome (Sears, 1956), many more resistance genes have been introduced, including those from Ae. tauschii (Lr21, Lr22a, Lr39) and Ae. speltoides (Lr28, Lr35, Lr36, Lr37, Lr51). Despite the considerable number of Lr genes that have already been introduced into wheat from Aegilops species, further screening studies are required to identify new gene(s) or combinations of genes in distinct genotypes.
To address this, 130 accessions of the Aegilops genus were screened for the presence of seedling and adult plant resistance Lr genes. After artificial exposure to P. triticina under greenhouse and field conditions, the seedlings and adult plants, respectively, were symptomatically evaluated. Comparing the outcomes enabled us to evaluate the efficiency of the tested genes and identify genotypes suitable for breeding. Moreover, we identified phenotypically resistant genotypes without the screened Lr genes whose genetically conditioned resistance is worth further investigation.
Material and methods
Plant material and phenotyping
The ex-situ gene bank Triticeae collection, maintained by the Czech Gene Bank, comprises 1763 accessions, of which the Aegilops genus accounts for 1230 accessions, encompassing 21 out of all 23 species. The material selected for this contribution was one set received from the Kazakh Gene Bank Almalybak/Almaty on exchange basis. It comes partly from extensive collecting missions by Kazakh partners and partly from other sources. Selected 130 accessions, belonging to 13 species, (Table 1, S1) were regenerated and phenotyped, and data recorded to the national information system GRIN Czech (GRIN Czech Release 1.10.3 (vurv.cz)). A more detailed characterization of the origin and location of material collection is available in Table S2.
Phenotyping under infection
The National Program on Conservation and Utilization of Microbial Genetic Resources and Invertebrates of Agricultural Importance provided the pathotypes of the pathogen. The spores were collected from leaves from the previous seasons from different localities to simulate the range of races that appear naturally. A collection of 6 pathotypes with varying virulence levels were utilized for resistance tests, as shown in Table 2. Seedlings of the NILs (Nearly Isogenic Lines) were inoculated in the greenhouse by rubbing leaves with an urediniospore water suspension, then they were sprayed with water and placed in glass cylinders for 24—48 h. Thereafter plants were kept in the greenhouse at 18—22 °C. Infection types were scored after 10—14 days based on the scale by Stakman et al. (1962).
Due to pure germination, from overall 130 genotypes screened with molecular markers, 122 genotypes were evaluated in the field and only 77 were possible to evaluate in greenhouse conditions. To provide clarity throughout the text, the SMM (Set tested with Molecular Markers) designation will now refer to 130 genotypes that were screened using molecular markers. The SF (Set tested on Field) designation will represent the 122 genotypes that were evaluated under field conditions, and the SG (Set tested in Greenhouse) designation will refer to the 77 genotypes that were evaluated in a greenhouse environment. It's important to note that the smaller set always contains only representatives from the larger one. Rust spreaders (cv. Michigan Amber) regularly distributed in the trial were inoculated with a mixture of the same P. triticina urediniospores as used in greenhouse experiment to test SF cultivars for resistance (disease severity). Artificial infection was performed by injecting water suspension of urediniospores into the stems of the susceptible variety spreader at the beginning of stem elongation phase. Visual estimation of disease severity took place in several terms according to the development of the disease using a disease severity scale of 1–9, where 9 stands for no symptoms observed and 1 represents heaviest sporulation leading to defoliation.
DNA extraction
DNA of 130 accessions (Table S1) was purified from leaf tissue by following optimized CTAB procedure (Murray & Thompson, 1980). Homogenized plant tissue was incubated in 2% CTAB solution within 30 min at 65 °C and subsequently cooled for 5 min on ice. After incubation with chloroform-isoamylalcohol (24:1) solution and series of centrifugation, DNA was precipitated using 99% ethanol and afterwards purified by 75% ethanol. Dried nucleic acid was incubated with RNase A (20 mg/ml) for 10 min at 37 °C and diluted in TE solution. The integrity of isolated DNA was assessed on 0.8% gel supplemented with ethidium bromide and its concentration measured spectrophotometrically. To set DNA work concentration on 100 ng/μl, DNA was further resuspended in nuclease-free water.
Lr genes detection
Polymerase chain reactions (PCR) were performed in 15 μl reaction volume with specific primers (Table 3) and DNA solution (100 ng/μl). The amplification profile and composition of reaction mixture are listed in Table S3. All amplification products were visualized using electrophoresis on 2% agarose gel stained with ethidium bromide. GeneRuler 100 bp DNA ladder was used to estimate amplicon size.
SSR analysis
The microsatellite markers were selected primarily for the diversity evaluation of wheat by our and French lab (INRA) in collaboration as a selection from hundreds SSR markers. Before this study, we tested these markers on several species of Aegilops and their spontaneous hybrids with wheat (Holubec et al., 2019). It was clear, that only a part of them could be used and applied on genotypes in this study. The final set consisted of 38 markers. Microsatellite analyses were performed according to Röder et al. (1998). PCR products were separated using a capillary electrophoresis in ABI PRISM 3130 sequencer (Life Technology, Foster City, CA, USA). A multiplexed configuration of four reactions was used in one analysis. As to interne size standard LIZ500 (Applied Biosystems) was used. Electrophoretograms were processed by GeneMapper software.
Data analysis
The gels were scored for the presence or absence of reproducible products with corresponding lengths. The molecular data from all genotypes screened with nine markers were transformed into a binary data matrix as discrete variables (1 = presence, 0 = absence). The results of symptomatic evaluation on field were transformed into a value between 0 and 1 depending on score in 1–9 scale. Data generated by symptomatic evaluation in greenhouse were also transformed into a value between 0 and 1 depending on number of resistant cases against 5 possible races. Canonical correlation analysis was used in R environment to identify and measure associations among genotypic and phenotypic data. One-way ANOVA was used to determine the significant difference in symptomatic response between genotypes with presence and absence of particular Lr gene.
A matrix of distances between all the samples was calculated from microsatellite data in the DARwin software (http://darwin.cirad.fr/darwin; Perrier & Jacquemoud-Collet, 2006). For clustering, a weighted Neighbour-joining method (WNJ) was used as its cophenetic coefficient r showed the highest value (0.947). The support for the phenogram branches was obtained using 1,000 bootstrap resampling. Analysis of population structure was performed with STRUCTURE 2.3.4 (Pritchard et al., 2000). The number of populations (K) was selected by testing K = 1 to K = 10 using ten independent iterations of 50 000 MCMC Reps after 20,000 burnin period length. The optimum K-value was identified according to methodology of Evanno et al. (2005).
Results
Leaf rust resistance genes and their phenotypic manifestation
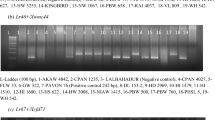
SMM set of genotypes was screened for the presence of 9 Lr genes (1, 9, 19, 21, 24, 28, 35, 37, 58) (Fig. 1a-c). Tested Lr genes were selected with respect to their role in APR/ASR resistance and based on their origin from wild relatives. All of genes were detected at least once with various frequencies (Table 4, Table S4), however in 32% of accessions none of tested gene was present. Marker for Lr24 was present just in one accession representing the only representative of Aegilops speltoides. The frequency of the remaining eight markers ranged from 6.9% (Lr35) to 31.5% (Lr37). Three genotypes were tested positive for five resistance genes, although most genotypes were tested positive for just one resistance gene at once. Based on these results, SF genotypes were divided into three groups. First group was represented by genotypes with at least two detected resistance genes. Remaining two groups consisted of genotypes with one and no detected Lr gene, respectively. All of 122 SF accessions were evaluated in field tests with score 1–9 (Table S4) and their variances were analyzed by one-way ANOVA. Groups showed statistically significant differences (F = 11.973; p ≤ 0.001).
Occurrence of Lr gene markers in each of 122 genotypes from SF set. Symptomatic evaluation in field tests was scored from 1 – susceptible to 9 – resistant, in greenhouse from 0 – susceptible to 1 – resistant. Genotypes are sorted by pattern of marker presence (a), symptomatic evaluation in field conditions (b), clustering according to DARwin software (c)
The relationship between genotypic and phenotypic data from SG genotypes was analyzed by CCA generating two canonical dimensions, both with statistical significance according to Wilks’s lambdas. For the first dimension, the canonical correlation was equal to 0.557 (p ≤ 0.01), for the second equal to 0.496 (p ≤ 0.01). With respect to standardized canonical coefficients of genotypic variables, the first canonical dimension was most strongly influenced by Lr24 (-0.47) and Lr58 (-0.81), the second by Lr1 (-0.45), Lr9 (-0.58), Lr21 (0.45), Lr28 (0.55) and Lr58 (0.53). Concerning phenotypic data, first dimension was influenced mainly by field evaluation (-0.97) and second by greenhouse (-1.12) variable.
For a more detailed interpretation of CCA results, one-way ANOVA was used. Statistically significant differences in symptomatic evaluation were observed in case of Lr9 (F = 7.702; p ≤ 0.01) and Lr24 (F = 5.865; p ≤ 0.05) in greenhouse conditions and Lr58 (F = 17.9; p ≤ 0.01) in field conditions.
All species counting 10 and more genotypes were further evaluated by one-way ANOVA to compare the average infestation rate in the presence and absence of a particular Lr gene (Table 5). A statistically significant difference for both field and seedling resistance was observed only for Lr37 in Ae. cylindrica. The rate of seedling infestation was significantly different for Ae. tauschii and Lr21, while Lr28 gene significantly affected the infestation degree of Ae. triuncialis on the field.
Microsatellites
To determine whether genetic variability corresponds to the occurrence of Lr genes, and subsequently to the degree of pathogen infestation, SMM genotypes were evaluated using 38 microsatellite markers. A total of 339 alleles were detected from the 38 amplified loci. The number of alleles per locus ranged from 2 to 22, with an average of 8.9 alleles per locus in the range of analyzed genotypes.
Cluster analysis based on microsatellite data showed three main clusters (A, B, C) supported by bootstrap resampling. Individual clusters were further divided into subclusters, all (except A.1) representing individual species (Fig. 2).
Similarly, the results obtained within DARwin software generally aligned with population structure analyzed by STRUCTURE and thus support the idea of three main populations within investigated Aegilops representatives (Fig. 3).
Discussion
The main aim of this study was to verify the extent to which the degree of resistance to the Puccinia triticina is influenced by the presence of Lr genes in Aegilops genus members. No Lr gene was present in one-third of the genotypes, yet some of these accessions showed resistance in both field and greenhouse conditions. These genotypes probably have other Lr genes contributing to their overall resistance and/or their resistance is based on various components of quantitative resistance (Azzimonti et al., 2013).
In our study, the marker for Lr1 was detected in all genotypes of Ae. juvenalis, Ae. neglecta and Ae. speltoides, in 89% of Ae. crassa genotypes, 73% of Ae. tauschii genotypes and 30% of Ae. geniculata genotypes. Lr1, available in many wheat cultivars (McIntosh et al., 1995) and Ae. tauschii accessions, is located on the D genome. D genome is further part of Ae. juvenalis and Ae. crassa, however we also detected Lr1 in accessions with absence of D genome, both Ae. neglecta and Ae. geniculate having the UM genome. This unlikely occurrence of Lr1 could have resulted from spontaneous hybridization of these species and wheat (Holubec et al., 2019), followed by misidentification of the resulting hybrids as representatives of Ae. geniculata and Ae. neglecta. Previous studies conducted on Ae. tauschii have shown resistance in 3.5% (Ling et al., 2004) to 11% (Assefa & Fehrmann, 2000) of tested genotypes accessions, slightly lower than ours 18%. Ling et al., however, counted only the accessions that displayed infection type 0, while we considered as resistant also types 1 and 2.
The marker associated with Lr9 was detected with various frequencies in Ae. biuncialis, Ae. columnaris, Ae. geniculate, Ae. juvenalis, Ae. neglecta, Ae. peregrina and Ae. triuncialis. All these species contain the U genome of diploid progenitor Aegilops umbellulate in which Lr9 was identified for the first time (Sears, 1961). Interestingly, Lr9 was also detected in Ae. markgrafii containing C genome only. Similar results to ours were reported for Ae. columnaris in a study of Aliakbari Sadeghabad et al. (2020) where the presence of Lr9 corresponded with observed seedling resistance.
Although the band corresponding to Lr19 marker was previously found in accessions of Ae. tauschii (Aliakbari Sadeghabad et al., 2020), this study observed it in genotypes of Ae. biuncialis, Ae. comosa, Ae. crassa, Ae. cylindrica, Ae. juvenalis and Ae. peregrina without conferring any observable resistance.
Lr21, first time identified in an Iranian Ae. tauschii accession (Huang et al., 2009), is claimed to provide complete resistance at seedling stage and partial resistance at adult stage under field conditions (Naz et al., 2021). However, there are studies identifying Lr21 as gene carrying APR (Draz et al., 2015; Kumar et al., 2022). Here, Lr21 was present not only in Ae. tauschi genotypes, but also in Ae. crassa. These two species had an equal representation of Lr1, Lr21 and Lr35 genes, but Ae. crassa possess two more Lr genes, namely Lr19 and Lr28. Still, Ae. crassa performed worse in both field and greenhouse tests compared to Ae. tauschi. The comparison of Ae. tauschii accessions revealed significantly lower resistance at seedling stage in presence of Lr21. Although this is in contrast with work of Naz et al., 2021, one of the causes might be high virulence of Czech isolates against Lr21 with average of 94% between 2002–2019 (Hanzalová et al., 2021).
The marker for Lr24, originating from Thinopyrum ponticum, was present in only one representative of the species Ae. speltoides. Although its presence in this species is surprising it has been previously recorded (Aliakbari Sadeghabad et al., 2020). The observed high resistance of this genotype in greenhouse and field conditions could also be mediated by the presence of Lr1, Lr28 and Lr37, whose individual presence did not show increased resistance.
All accessions of Ae. crassa, Ae. juvenalis, Ae. speltoides and one of Ae. triuncialis have been shown to be the residence of the Lr28 gene. The presence of this gene was also observed outside the genus Aegilops, in Triticum timopheevii – a tetraploid member of the Triticeae tribe (Gultyaeva et al., 2014). Out of 12 genotypes possessing this gene, only two showed the maximal resistance in the field (9) and just one was resistant against all examined isolates, despite the satisfactory results of Lr28 in Czech wheat cultivars (Hanzalová & Zelba, 2022). Moreover, the presence of Lr28 in Ae. triuncialis accessions was associated with significantly higher infestation level on the field. Two reasons may explain this result. The first, there was a single genotype of Ae. triuncialis carrying the Lr28 rated 4 on the symptomatic scale. The field resistances of the remaining genotypes were mostly the highest possible shifting the results against the presence of Lr28. Secondly, Lr28 is considered a juvenile resistance gene and therefore may not have such an effect on adult plant resistance.
Interestingly, the presence of the gene Lr35 was not observed in Ae. speltoides, the species from which it was derived. Similarly, overall distribution of this gene was low with its presence in 29% of Ae. crassa accessions, 14% of Ae. tauschii accessions and 4% of Ae. cylindrica accessions, all D-geneome species.
Lr37, the most abundant Lr gene in this study, is also the most represented gene conferring adult plant resistance in Czech wheat cultivars (Hanzalová & Zelba, 2022; Holubec et al., 2014). Although Lr37 did not show a population-wide effect, within Ae. cylindrica was indicated to be a possible factor positively affecting resistance not only of adult plants, but also of seedlings.
Lr58 originates in Ae. triuncialis with accessions carrying effective resistance against leaf rust for many years (Arora et al., 2021). Although it is not yet clear wether Lr58 is more of juvenile resistance or adult plant resistance gene, its performance appeared to be more pronounced in the field. The benefiting genotypes were of Ae. geniculata, Ae. peregrina and Ae. triuncialis. However, the presence of this gene did not translate into increased resistance of species Ae. juvenalis.
Based on the results of ANOVA, one Lr gene was sufficient to observe a significant difference in the rate of infestation in the field (Fig. 1b). However, each additional Lr gene did not result in a statistically significant increase in resistance. This trend is mostly prevalent in Ae. triuncialis accession, where Lr58 alone is sufficient to maintain high levels of resistance (Fig. 1a).
The results verified by canonical correlation analysis and subsequent ANOVA showed that the genotypes carrying Lr9 performed significantly better under greenhouse conditions. This is in agreement with reports declaring efficiency of Lr9 in Europe (Aktar-Uz-Zaman et al., 2017; Volkova et al., 2022) and against Czech isolates used also in this study (Hanzalová et al., 2021). Similarly, if the genotype carried Lr58, it was more likely to be resistant in the field. This was true in all genotypes of Ae. biuncialis and Ae. peregrina and 75% of Ae. triuncialis genotypes with scoring of 9. In Ae. geniculate the presence of Lr58 provided only mean resistance of 4.5.
Genotypes belonging to 13 different species of the genus Aegilops were analyzed to assess genetic variability. Cluster analysis revealed the existence of three groups in the examined population. The largest observed variability was in the group A (mainly from the sect. Aegilops), which clustered into three smaller groups. Within the group B there are two closely related D-genome species, splitting in two subgroups, each representing a different species. The last group C showed the least variability and contained exclusively genotypes of the species Ae. cylindrica (Fig. 2). The results from STRUCTURE confirmed the division of genotypes into three main groups, but the division into subgroups was beyond the distinctiveness of the program (Fig. 3). Abbasov et al. (2019) used microsatellite markers to evaluate the genetic diversity of Aegilops genotypes from Azerbaijan and Georgia evaluating Ae. crassa and Ae. vavilovi as the most diverse species. On the other hand, Ae. cylindrica possessed the lowest diversity, which is congruent not only with our results, but also with findings of previous studies (Goryunova et al., 2004; Pester et al., 2003). Contrary to this, Moradkhani et al. (2015) based on microsatellites and Pour-Aboughadareh et al. (2017) based on SCoT markers, determined a relatively higher diversity in Ae. cylindrica compared with other species. These discrepancies might be caused by the different sample size and Ae. cylindrica germplasms from different countries. However, considering the fact Ae. cylindrica is evolutionary younger species (Gandhi et al., 2005) then their diploid progenitors Ae. markgrafii and Ae. tauschii, our results could be of more accuracy.
In addition, the application of DARwin-based clustering on Lr gene occurrence and symptomatic evaluation revealed that group A consists mainly of resistant genotypes compared to representatives in group B. This is even more pronounced when comparing the group A with group C accessions (Fig. 1c).
In conclusion, we have shown that leaf rust resistance in Aegilops is not strictly based on separate resistance genes, but rather the combination of specific genes is needed to ensure higher resistance levels. Our results will not only contribute to knowledge of the genetically based resistance of the Aegilops genus, but also indicate how the genetic composition translates into a phenotype measured by symptomatic evaluation, given that our results suggest an effective contribution of Lr9 on seedling resistance and Lr58 on resistance of adult plants Moreover, identified resistant genotypes may serve as suitable source of resistance in further breeding. The fact that some of accessions showed increased resistance without any Lr gene present made these accessions worth for further investigation with potential of finding new, yet undiscovered resistance contributors. Although we screened with only nine molecular markers, these were the once with the highest probability of detection due to their origin in wild relatives. However, there are more markers that can reflect on overall resistance and thus a broader set of molecular markers could be used in future studies.
Data Availability
All relevant data are available either in manuscript or in the supplementary file1.
References
Abbasov, M., Brueggeman, R., Raupp, W., et al. (2019). Genetic diversity of Aegilops L. species from Azerbaijan and Georgia using SSR markers. Genetic Resources and Crop Evolution, 66, 1–11. https://doi.org/10.1007/s10722-018-0725-3
Aktar-Uz-Zaman, M., Tuhina-Khatun, M., Hanafi, M. M., & Sahebi, M. (2017). Genetic analysis of rust resistance genes in global wheat cultivars: An overview. Biotechnology and Biotechnological Equipment, 31(3), 431–445. https://doi.org/10.1080/13102818.2017.1304180
Aliakbari Sadeghabad, A., Dadkhodaie, A., & Heidari, B. (2020). Phenotypic and genetic diversity of leaf rust resistance in wheat wild relatives. Journal of Phytopathology, 168, 428–438. https://doi.org/10.1111/jph.12907
Arora, S., Kaur, S., Dhillon, G. S., Singh, R., Kaur, J., Sharma, A., & Chhujena, P. (2021). Introgression and genetic mapping of leaf rust and stripe rust resistance in Aegilops triuncialis. Journal of Genetics, 100, 6. https://doi.org/10.1007/s12041-020-01253-3
Assefa, S., & Fehrmann, H. (2000). Resistance to wheat leaf rust in Aegilops tauschii Coss. and inheritance of resistance in hexaploid wheat. Genetic Resources and Crop Evolution, 47(2), 135–140.
Azzimonti, G., Lannou, C., Sache, I., & Goyeau, H. (2013). Components of quantitative resistance to leaf rust in wheat cultivars: Diversity, variability and specificity. Plant Pathology, 62, 970–981. https://doi.org/10.1111/ppa.12029
Cherukuri, D. P., Gupta, S. K., Charpe, A., Koul, S., Prabhu, K. V., Singh, R. B., & Haq, Q. (2005). Molecular mapping of Aegilops speltoides derived leaf rust resistance gene Lr28 in wheat. Euphytica, 143, 19–26. https://doi.org/10.1007/s10681-005-1680-6
Draz, I. S., Abou-Elseoud, M. S., Kamara, A. E., Alaa-Eldein, O. A., & El-Bebany, A. F. (2015). Screening of wheat genotypes for leaf rust resistance along with grain yield. Annals of Agricultural Science, 60(1), 29–39. https://doi.org/10.1016/j.aoas.2015.01.001
Evanno, G., Regnaut, S., & Goudet, J. (2005). Detecting the number of clusters of individuals using the software structure: A simulation study. Molecular Ecology, 14, 2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Gandhi, H. T., Vales, M. I., Watson, C. J., Mallory-Smith, C. A., Mori, N., Rehman, M., Zemetra, R. S., & Riera-Lizarazu, O. (2005). Chloroplast and nuclear microsatellite analysis of Aegilops cylindrica. TAG. Theoretical and Applied Genetics., 111(3), 561–572.
Goryunova, S. V., Kochieva, E. Z., Chikida, N. N., & Pukhalskyi, V. A. (2004). Phylogenetic relationships and intraspecific variation of D-genome Aegilops L. as revealed by RAPD analysis. Russian Journal of Genetics (Translation of Genetika (Moscow, Russian Federation)), 40, 515–523.
Gultyaeva, E. I., Orina, A. S., Gannibal, P. B., Mitrofanova, O. P., Odintsova, I. G., & Laikova, L. I. (2014). The effectiveness of molecular markers for the identification of Lr28, Lr35, and Lr47 genes in common wheat. Russian Journal of Genetics (Translation of Genetika (Moscow, Russian Federation)), 50, 131–139. https://doi.org/10.1134/S1022795414020069
Gupta, S., Charpe, A., Koul, S., Haq, Q., & Prabhu, K. (2006). Development and validation of SCAR markers co-segregating with an Agropyron elongatum derived leaf rust resistance gene Lr24 in wheat. Euphytica, 150, 233–240. https://doi.org/10.1007/s10681-006-9113-8
Hanzalová, A., Dumalasová, V., & Zelba, O. (2021). Virulence in the Puccinia triticina population in the Czech Republic and resistance genes in registered cultivars 1966–2019. Euphytica, 217, 4. https://doi.org/10.1007/s10681-020-02733-4
Hanzalová, A., & Zelba, O. (2022). Leaf rust (Puccinia triticina Eriks) resistance genes in wheat cultivars registered in the Czech Republic. J Plant Dis Protect. https://doi.org/10.1007/s41348-022-00625-4
Helguera, M., Khan, I. A., Kolmer, J. A., Lijavetzky, D., Zhong-qi, L., & Dubcovsky, J. (2003). PCR assays for the Lr37-Yr17-Sr38 cluster of rust resistance genes and their use to develop isogenic hard red spring wheat lines. Crop Science, 43, 1839–1847.
Holubec, V., Hanzalová, A., Dumalasová, V., & Bartoš, P. (2014). Aegilops conservation and collection evaluation in the Czech Republic. Journal of Systematics and Evolution, 52, 783–789. https://doi.org/10.1111/jse.12117
Holubec, V., Leišová-Svobodová, L., & Matějovič, M. (2019). Spontaneous hybridisation within Aegilops collection and biobanking of crop wild relatives (CWR). Genetic Resources and Crop Evolution, 66, 311–319. https://doi.org/10.1007/s10722-018-0723-5
Huang, L., Brooks, S., Li, W., Fellers, J., Nelson, J. C., & Gill, B. (2009). Evolution of new disease specificity at a simple resistance locus in a crop–weed complex: Reconstitution of the Lr21 gene in wheat. Genetics, 182(2), 595–602. https://doi.org/10.1534/genetics.108.099614
Huang, L., & Gill, B. S. (2001). An RGA-like marker detects all known Lr21 leaf rust resistance gene family members in Aegilops tauschii and wheat. TAG. Theoretical and Applied Genetics., 103(6–7), 1007–1013.
Huerta-Espino, J., Singh, R., Germán, S., McAllum, B. D., Park, R. F., Chen, W. Q., Bhardwaj, S. C., & Goyeau, H. (2011). Global status of wheat leaf rust caused by Puccinia triticina. Euphytica, 179, 143–160. https://doi.org/10.1007/s10681-011-0361-x
Kolmer, J. A. (2019). Virulence of Puccinia triticina, the wheat leaf rust fungus, in the United States in 2017. Plant Disease, 103(8), 2113–2120. https://doi.org/10.1094/PDIS-09-18-1638-SR
Kolmer, J. A., Hanzalová, A., Goyeau, H., Bayles, R., & Morgounov, A. (2013). Genetic differentiation of the wheat leaf rust fungus Puccinia triticina in Europe. Plant Pathology, 62(1), 21–31. https://doi.org/10.1111/j.1365-3059.2012.02626.x
Krupinsky, J. M., Bailey, K. L., McMullen, M. P., Gossen, B. D., & Turkington, T. K. (2002). Managing plant disease risk in diversified cropping systems. Journal of Agronomy, 94, 198–209. https://doi.org/10.2134/agronj2002.1980
Kumar, K., Jan, I., Saripalli, G., Sharma, P., Mir, R., Balyan, H., & Gupta, P. (2022). An update on resistance genes and their use in the development of leaf rust resistant cultivars in wheat. Frontiers in Genetics, 13, 816057. https://doi.org/10.3389/fgene.2022.816057
Kuraparthy, V., Sood, S., Guedira, G., & Gill, B. (2011). Development of a PCR assay and marker-assisted transfer of leaf rust resistance gene Lr58 into adapted winter wheats. Euphytica, 180, 227–234. https://doi.org/10.1007/s10681-011-0383-4
Ling, H. Q., Qiu, J. W., Singh, R. P., & Keller, B. (2004). Identification and genetic characterization of an Aegilops tauschii ortholog of the wheat leaf rust disease resistance gene Lr1. Theoretical and Applied Genetics, 109, 1133–1138. https://doi.org/10.1007/s00122-004-1734-5
Mago, R., Zhang, P., Bariana, H., Verlin, D., Bansal, U., Ellis, J., & Dundas, I. (2009). Development of wheat lines carrying stem rust resistance gene Sr39 with reduced Aegilops speltoides chromatin and simple PCR markers for marker-assisted selection. Theoretical and Applied Genetics, 119, 1441–1450. https://doi.org/10.1007/s00122-009-1146-7
McCallum, B. D., Reimer, E., McNabb, W., Foster, A., Rosa, S., & Xue, A. (2021). Physiologic specialization of Puccinia triticina, the causal agent of wheat leaf rust, in Canada in 2015–2019. Canadian Journal of Plant Pathology, 43, 333–346. https://doi.org/10.1080/07060661.2021.1888156
McIntosh, R. A., Wellings, C. R., & Park, R. F. (1995). Wheat rusts: An atlas of resistance genes. CSIRO Publications.
Moradkhani, H., Mehrabi, A. A., Etminan, A., & Pour-Aboughadareh, A. (2015). Molecular diversity and phylogeny of Triticum-Aegilops species possessing D genome revealed by SSR and ISSR markers. Plant Breeding and Seed Science, 71(1), 81–95.
Murray, M. G., & Thompson, W. F. (1980). Rapid isolation of high molecular weight plant. Nucleic Acids Research, 8, 4321–4325.
Naz, A. A., Bungartz, A., Serfling, A., Kamruzzaman, M., Schneider, M., Wulff, B. B. H., Pillen, K., Ballvora, A., Oerke, E. C., Ordon, F., & Léon, J. (2021). Lr21 diversity unveils footprints of wheat evolution and its new role in broad-spectrum leaf rust resistance. Plant, Cell and Environment, 44(10), 3445–3458. https://doi.org/10.1111/pce.14144
Nelson, R., Wiesner-Hanks, T., Wisser, R., & Balint-Kurti, P. (2018). Navigating complexity to breed disease-resistant crops. Nature Reviews Genetics, 19(1), 21–33. https://doi.org/10.1038/nrg.2017.82
Oerke, E. (2006). Crop losses to pests. Journal of Agricultural Science, 144(1), 31–43. https://doi.org/10.1017/S0021859605005708
Park, R. F., Bariana, H. S., Wellings, C. R., & Wallwork, H. (2002). Detection and occurrence of a new pathotype of Puccinia triticina with virulence for Lr24 in Australia. Australian Journal of Agricultural Research, 53, 1069–1076.
Perrier, X., & Jacquemoud-Collet, J. P. (2006). DARwin Software. http://darwin.cirad.fr/darwin.
Pester, T. A., Ward, S. M., Fenwick, A. L., Westra, P., & Nissen, S. J. (2003). Genetic diversity of jointed goatgrass (Aegilops cylindrica) determined with RAPD and AFLP markers. Weed Science, 51, 287–293.
Pour-Aboughadareh, A., Ahmadi, J., Mehrabi, A. A., Etminan, A., & Moghaddam, M. (2017). Insight into the genetic variability analysis and relationships among some Aegilops and Triticum species, as genome progenitors of bread wheat, using SCoT markers. Plant Biosyst, 152(4), 694–703.
Prasad, P., Savadi, S., Bhardwaj, S. C., & Gupta, P. K. (2020). The progress of leaf rust research in wheat. Fungal Biology, 124(6), 537–550. https://doi.org/10.1016/j.funbio.2020.02.013
Prins, R., Groenewald, J., Marais, G., Sanpe, J., & Koebner, R. (2001). AFLP and STS tagging of Lr19, a gene conferring resistance to leaf rust in wheat. Theoretical and Applied Genetics, 103, 618–624. https://doi.org/10.1017/S002185960500570810.1007/PL00002918
Pritchard, J. K., Stephens, M., & Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics, 155(2), 945–959. https://doi.org/10.1093/genetics/155.2.945
Qiu, J. W., Schürch, A. C., Yahiaoui, N., Dong, L. L., Fan, H. J., Zhang, Z. J., Keller, B., & Ling, H. Q. (2007). Physical mapping and identification of a candidate for the leaf rust resistance gene Lr1 of wheat. TAG. Theoretical and Applied Genetics., 115(2), 159–168. https://doi.org/10.1007/s00122-007-0551-z
Röder, M. S., Korzun, V., Wendehake, K., Plaschke, J., Tixier, M. H., Leroy, P., & Ganal, M. W. (1998). A microsatellite map of wheat. Genetics, 149(4), 2007–2023. https://doi.org/10.1093/genetics/149.4.2007
Schachermayr, G., Siedler, H., Gale, M. D., Winzeler, H., Winzeler, M., & Keller, B. (1994). Identification and localization of molecular markers linked to the Lr9 leaf rust resistence gene of wheat. TAG. Theoretical and Applied Genetics., 88(1), 110–115. https://doi.org/10.1007/BF00222402
Sears, E. R. (1956). The transfer of leaf rust resistance from Aegilops umbellulata into wheat. Brookhaven Symposia in Biology: 1–22.
Sears, E. R. (1961). Identification of the wheat chromosome carrying leaf rust resistance from Aegilops umbellulata. Wheat Information Service, 12, 12–13.
Stakman, E. C., Stewart, D. M., & Loegering, W. O. (1962). Identification of physiologic races of Puccinia graminis var. tritici. US department of agriculture Agricultural Research Service E-617, Washington DC.
Volkova, G., Kudinova, O., Vaganova, O., & Agapova, V. (2022). Effectiveness of leaf rust resistance genes in the adult and juvenile stages in southern russia in 2011–2020. Plants, 11, 793. https://doi.org/10.3390/plants11060793
Funding
Open access publishing supported by the National Technical Library in Prague. This work received funding from the Ministry of Agriculture of the Czech Republic under the project number MZE-RO0423.
Author information
Authors and Affiliations
Contributions
VH and ME provided the materials. AH performed the greenhouse and VH the field experiments. LK and LS performed the molecular methods. LK and MJ analyzed the data. LK wrote and LS, AH and VH reviewed and edited the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Research involving human participants and/or animals
The authors confirm there was no research on any humans or animals.
Consent to participate
All authors have reviewed the manuscript and consented the submission to the European Journal of Plant Pathology.
Competing interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Koláriková, L., Svobodová-Leišová, L., Hanzalová, A. et al. Leaf rust resistance genes in Aegilops genus: occurrence and efficiency. Eur J Plant Pathol 167, 335–348 (2023). https://doi.org/10.1007/s10658-023-02712-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-023-02712-0