Abstract
A problem to implement conservation strategies is that in many cases recognized taxa are in fact complexes of several cryptic species. Failure to properly delineate species may lead to misplaced priorities or to inadequate conservation measures. One such species complex is the yellow-spotted ringlet Erebia manto, which comprises several phenotypically distinct lineages, whose degree of genomic isolation has so far not been assessed. Some of these lineages are geographically restricted and thus possibly represent distinct units with conservation priorities. Using several thousand nuclear genomic markers, we evaluated to which degree the bubastis lineage from the Alps and the vogesiaca lineage from the Vosges, are genetically isolated from the widespread manto lineage. Our results suggest that both lineages are genetically as strongly differentiated from manto as other taxonomically well separated sibling species in this genus from each other, supporting a delineation of bubastis and vogesiaca as independent species. Given the restricted and isolated range of vogesiaca as well as the disjunct distribution of bubastis, our findings have significant implication for future conservation efforts on these formerly cryptic species and highlight the need to investigate the genomic identity within species complexes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Insect diversity and biomass are declining at accelerated rates in Europe and elsewhere (Hallmann et al. 2017, Seibold et al. 2019, van Klink et al. 2020), and insect conservation has become a key topic in conservation biology in the last decades (Dunn 2005; Forister et al. 2019). Poorly known life histories or geographic distributions combined with often challenging identification and unresolved taxonomy may hamper conservation efforts for invertebrates more than vertebrates. An incorrect taxonomy that amalgamates species complexes into single taxa may for example lead to erroneous prioritization in conservation or to inadequate conservation measures (Ceballos et al. 2017). This situation can occur when the overall taxon is classified as being of low concern for conservation due to the broad occurrence of the entire species complex, or if conservation measures are based on ecological assumptions from the entire species complex and do not account for ecologically more specialized cryptic species (Bickford et al. 2007). It is therefore important to examine such species complexes and to delineate units for conservation efforts.
Across Europe, almost 500 species of butterflies have been recognized (Wiemers et al. 2018), many of which are declining in their population sizes and/or distribution ranges (Warren et al. 2021). Numerous taxa comprise different subspecies or lineages (Settele et al. 2008), which may moreover hybridize to varying degrees upon secondary contact (Descimon and Mallet 2009). The genus Erebia is known for its cryptic diversity (Sonderegger 2005; Descimon and Mallet 2009) and also includes “geographical replacement species”, i.e. species or lineages that replace each other by their vicariant sibling in parts of their range (Vodă et al. 2015). The taxonomic status of such species or lineages often remains ambiguous because they are possibly weakly isolated and some level of hybridization is still possible.
For Erebia, the degree of differentiation upon secondary contact between closely related species or lineages varies (Sonderegger 2005). For example, subspecies of Erebia euryale form narrow contact zones with phenotypic intermediates (Cupedo 2014; Cupedo and Doorenweerd 2022). This contrasts with the E. tyndarus-group, where geographic lineages represent reproductively isolated species with nearly interrupted gene flow. In particular, E. cassioides and E. tyndarus are parapatric in the Alps, and a third species, the very restricted E. nivalis, is found in range sympatry with the other two. Recent genetic studies showed that F1 hybrids between E. cassioides and E. tyndarus very rarely occur at their narrow zone of secondary contact, but no F2 hybrids have been reported among these three taxa, which exhibit limited levels of admixture, suggesting near complete reproductive isolation, consistent with distinct species (Gratton et al. 2016; Lucek et al. 2020; Augustijnen et al. 2022). Similar cases of strict parapatry without morphological intergradation also occur in other Erebia, leading to fragmented distributions and raising conservation concerns (Sonderegger 2005). This situation is observed in E. sudetica and E. melampus, two taxa that have non-overlapping distributions in Europe. In Switzerland, the former is limited to a few, small areas where E. melampus is absent.
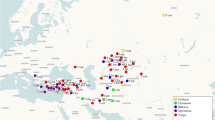
The yellow-spotted ringlet (Erebia manto complex) is a locally common butterfly with an insular-like distribution across Europe (Fig. 1), occurring along disconnected mountain ranges from northern Spain to the Carpathians, including the Alps and the Vosges (Schmitt et al. 2014; Kudrna et al. 2015). This disjunct distribution together with geographic variation in morphology has triggered the description of many allopatric E. manto subspecies in the past (Cupedo 1997; Cupedo and Doorenweerd 2020). The taxonomic status of these subspecies, is, however, debated (Sonderegger 2005; Schmitt et al. 2014; Cupedo and Doorenweerd 2020). Morphological and allozyme studies have indicated that the current distribution of the E. manto complex is likely shaped by post-glacial recolonization events from distinct glacial refugia, suggesting that part of the described diversity within the E. manto complex may therefore be old and predate the last glacial cycle (Cupedo 1997; Schmitt et al. 2014). Based on a detailed morphological study, Cupedo (1997) proposed to recognize the manto, the bubastis and the vogesiaca lineages, which substantially differ in genital structure. In particular, manto has substantially more spines on the valve than bubastis, while vogesiaca shows an intermediate phenotype (Cupedo 1997; Sonderegger 2005; Cupedo and Doorenweerd 2020; Fig. 2). Such marked differentiation in genital morphology in butterflies and in the genus Erebia in particular, is frequently used to delineate species (Warren 1936; Sonderegger 2005). Difference in male genital morphology may act as a reproductive barrier through lock-and-key mechanisms that reduce or prevent interspecific gene flow (Hollander et al. 2018). However, the lineages of the manto complex have so far mostly been treated as infraspecific units. Cupedo (1997) suggested that the phenotypic differences among the three lineages were as important as those observed between other Erebia species, but refrained from formally recognizing them as distinct species because evidence from cross experiments was lacking. By contrast, Cupedo and Doorenweerd (2020) recommended to treat the three lineages as conspecific due to one event of mitochondrial introgression between manto and bubastis. As far as it is known, the three lineages currently never occur in sympatry. While the manto lineage is widespread throughout Europe, the distribution of the other two lineages is more restricted. The vogesiaca lineage, in particular, is found in the Vosges (ssp. vogesiaca), possibly with an isolated population in the French Jura (Cupedo and Doorenweerd 2020), and in the Carpathian Mountains (ssp. trajanus). The bubastis lineage is restricted to narrow areas in the southwestern (ssp. valmaritima), western (ssp. willieni) and central Alps (ssp. bubastis), where it shows a disjunct distribution intermixed with populations of the manto lineage, which is widely distributed in Europe, including in the Alps (Fig. 1; Sonderegger 2005; Cupedo and Doorenweerd 2020).
Distribution of the three lineages of the Erebia manto species complex across Europe. Each dot depicts an individual occurrence based on data from GBIF.org (accessed 16th of March 2022 with additional data from Lelo 2000; Burnaz 2008; Sanz et al. 2017; Jakšic 2019). A few doubtful occurences, as well as a few occurrences in the French Alps south of Grenoble, which could not be attributed unambiguously to one subspecies, have been omitted. Colors depict the different recognized E. manto subspecies (see Cupedo & Doorenweerd 2020). The inset depicts the Alps with numbers highlighting the populations sampled (see Supplementary Information 1 for details). The attribution to some occurrences to the different subspecies have been made following Cupedo (1997), Schmitt et al. (2014) and Cupedo & Doorenweerd (2020). For simplicity, manto and mantoides are merged and treated as "manto" given the uncertainties regarding the type locality of manto (see Cupedo & Doorenweerd 2020). Map sources: European Comission GISCO: https://ec.europa.eu/eurostat/web/gisco; EU-DEM https://www.eea.europa.eu/data-and-maps/data/eu-dem
Phylogram artificially rooted on the node that separates E. eriphyle from the E. manto complex comprising manto, bubastis and vogesiaca. Numbers depict bootstrap support for nodes with > 90% support. Colors indicate taxa. Numbers at the terminal branches indicate sampling location as given in Fig. 1. For each E. manto lineage, a picture of the male genital valve is given
The status of the lineages of the manto complex has only partially been examined using genetic data. Based on allozymes, Schmitt et al. (2014) demonstrated the presence of six genetic clusters in Europe, one in the Pyrenees and the Massif Central (ssp. constans belonging to the manto lineage), one in the Vosges (vogesiaca), three in the Alps and the Slovakian Carpathians (including ssp. mantoides and ssp. manto; both are subsequently merged and referred to as ssp. manto for simplicity due to uncertainties regarding the type locality of manto; see Cupedo and Doorenweerd 2020), and one in the Carpathians mountains (ssp. trajanus). Unfortunately, no bubastis sample was included, preventing conclusions on the status of the bubastis lineage. They nonetheless showed that the divergence between the vogesiaca and the other forms were as deep as those between the manto complex and the closely-related species E. eriphyle. These results suggested that the separation between the lineages of the manto complex could be far more divergent than initially assumed. More recent research based on mitochondrial DNA indicate that manto and bubastis exhibit minimal, but constant differences in their barcode sequences, except for one site in the French Alps; the manto lineage was though paraphyletic with respect to bubastis (Litman et al. 2018; Cupedo and Doorenweerd 2020). While mitochondrial barcodes may often suffice to delineate European butterfly species (Dincă et al. 2021), they may not have the resolution to do so in evolutionary young species or in the case of cytonuclear discordance as a result of incomplete lineage sorting and/or introgression (Toews and Brelsford 2012; Gueuning et al. 2020). Here, we used restriction-site associated DNA (RAD) markers to study the genetic relationship within part of the manto complex. We focused on the Alps and the Vosges and first assessed if and to which degree bubastis and vogesiaca may be genetically isolated from manto. Because the Alps were often recolonized by different glacial lineages of Erebia (Schmitt et al. 2006, 2016; Lucek et al. 2020), we further tested for genetic differentiation within manto and bubastis across the Alps. We discuss our findings for the manto complex in the broader context of Erebia systematics and their implications for conservation.
Methods
Sampling
We included a total of 95 butterflies collected during summer 2019 encompassing specimens of the three Central European E. manto lineages (following Cupedo and Doorenweerd 2020): manto (N = 65), bubastis (N = 21) and vogesiaca (N = 5). We further refer to these lineages simply as manto, bubastis and vogesiaca respectively. Sampling was conducted at 24 sites that were at least 3 km apart from each other and covered the Alps and the Vosges mountain ranges (Fig. 1; see Supplementary Information 1 for details). At each site, we collected one to six individuals (Supplementary Information 1). For bubastis, we specifically sampled the three locations in the Swiss Alps for which the presence of bubastis is documented (Sonderegger 2005; Litman et al. 2018). We further included a disjunct population of the bubastis lineage from the French Alps. This population belongs to the subspecies willieni and was collected at its type locality, located 180 km south-west from the nearest Swiss bubastis population (Cupedo 1997; Fig. 1); manto populations are found between this French and the Swiss bubastis populations. We also included four individuals of the closely related species E. eriphyle as an outgroup for our phylogenomic analysis. All individuals were captured with an insect net, their bodies stored in 100% ethanol and the wings kept separately. We a priori assigned each individual to an E. manto lineage based on geography combined with wing morphology for females and genital morphology for males. DNA extractions were performed using the Qiagen Dneasy Blood and Tissue kit (Qiagen, Zug, Switzerland) from the thorax. Wings and abdomens are deposited at the Natural History Museum of Neuchâtel, Switzerland.
Because manto and bubastis have been suggested to primarily occur on calcareous and siliceous substrates, respectively (Sonderegger 2005), we visualized their occurrence across the Swiss Alps in regard to broad scale geological substrate. For this we obtained location data with a precision of < 100 m from the database of the Swiss zoological record centre (www.infospecies.ch) on the 2nd of April 2022 for a total of 3886 E. manto and 81 E. bubastis. We further extracted the lithological-petrographic information for each occurrence based on geological layers of Switzerland (https://opendata.swiss/en/dataset/lithologisch-petrografische-karte-der-schweiz-gesteinklassierung-1-500000).
Genetic data processing
We genotyped all 95 individuals using single-end restriction-site associated DNA (RAD) sequencing with the restriction enzyme SbfI. Library preparation and sequencing on one Illumina HiSeq 4000 lane was outsourced to Floragenex (Portland, OR, USA).
We filtered all obtained genomic data following Lucek et al. (2020), i.e., we only retained reads with an intact SbfI restriction site, followed by de-multiplexing and barcode-trimming with process_radtags from Stacks 1.48 (Catchen et al. 2013). Using the FASTX toolkit (http://hannonlab.cshl.edu/fastx_toolkit/), we then removed reads containing bases with a Phred quality score < 10 or more than 5% of base pairs with quality < 30. This approach yielded ~ 370 million high quality reads in total for our analysis. In a next step, we mapped the reads of each individual against a genome assembly of E. cassioides with BWA MEM 0.7.17 (Li 2013) and genotyped all specimens with BCFtools 1.10.2 (Danecek and McCarthy 2017). We filtered the genotypes with VCFtools 0.1.16 (Danecek et al. 2011) to include only bi-allelic polymorphic sites with a minimal depth of six and a minimal genotype quality of 28, employing a minor allele frequency filter of 0.03 and allowing up to 20% of missing data per site. Due to high rates of missing data, two manto specimens were filtered out. The overall filtering resulted in 3′994 SNP sites available for our downstream analyses.
Population genetic analyses
To infer the phylogenomic structure across all retained specimens, we first used RAxML 8.2.11 (Stamatakis 2014) implementing a generalised time-reversible (GTR) model with optimised substitution rates and a gamma model of rate heterogeneity. We further applied an ascertainment bias correction to account for the fact that we only used polymorphic SNP positions with the ASC_GTRGAMMA function implemented in RAxML. Significance was assessed using 1′000 bootstrap replicates followed by a thorough maximum likelihood search.
We inferred population structure in a first step with Admixture 1.3.0, which implements a likelihood approach to estimate ancestry (Alexander et al. 2009). We ran Admixture first across all individuals, excluding E. eriphyle, and then separately for manto and bubastis, to test for further intraspecific population structure. In each case, we varied the values for K, i.e., the number of assumed populations, from 1 to 10 and performed a cross-validation test to determine the optimal value of K. In a second step we used a principal component (PC) analysis as implemented in GenoDive 3.0.5 (Meirmans 2020) to visualize the genetic relationship among individuals. As for Admixture we performed the PC analysis including either all individuals or separately for manto and bubastis. Subsequent statistical analyses on the resulting PC scores were done using linear models in R 4.1.1 (R Core Team 2021).
Next, we estimated the level of pairwise genetic differentiation (FST) among species and lineages using GenoDive, pooling individuals from across the distribution range for manto and bubastis, respectively. Significance was estimated based on 1′000 bootstrap iterations. We also estimated FST between individuals of two identified clusters within the manto lineage, excluding individuals that showed admixture.
Lastly, we assessed recent migration rates between species with BA3-SNPs V 3.0.4 (Mussmann et al. 2019), a modification of BayesAss (Wilson and Rannala 2003) that allows handling of large SNP datasets. First, we assessed the optimal mixing parameters for migration rates (deltaM = 0.1563), allele frequencies (delta = 0.5500), and inbreeding coefficients (deltaF = 0.0750) by running ten repetitions in BA3-SNP-autotune V 3.0.4 as recommended by Mussmann et al. (2019). Subsequently, BA3-SNPs was run with the predefined mixing parameters for 50 million generations, sampling every 100th generation. The first million generations were discarded as burn-in and chain convergence was assessed in Tracer V 1.7.1 (Rambaut et al. 2018).
Results
Consistent with three independent taxonomic entities, the RAXML analysis resolved the manto, bubastis and vogesiaca lineages as distinct phylogenetic clades with 100% bootstrap support each (Fig. 2). Here, vogesiaca split first from the two other taxa, which formed a monophyletic group with 100% bootstrap support. Some phylogenetic structuring was observed among bubastis individuals, each population representing a distinct clade. In manto, the phylogenetic structuring was less pronounced, except for some populations; in particular, the four individuals from the easternmost part of the Swiss Alps (population 24 in Figs. 1 & 2), which constituted a sister clade to all other individuals.
Four genetic clusters (K = 4) were the best fitting number as inferred with Admixture when manto, bubastis and vogesiaca were jointly analysed (Fig. 3a, b & Supplementary Information 2). Two clusters represent the vogesiaca and bubastis lineages respectively and two additional clusters occur within the manto lineage. The overall PC analysis suggests three main clusters corresponding to bubastis, vogesiaca and manto respectively, where the two manto clusters were grouped together (Fig. 3c). Both bubastis and vogesiaca formed a distinct genetic cluster that showed no evidence for gene flow from manto. Interspecific gene flow seems indeed to have primarily occurred from bubastis into manto, but its current extent seems limited (Fig. 3a). Contrasting with this result, the two genetic clusters within the manto lineage showed substantial genetic admixture in the central Swiss Alps (Fig. 3a, b), which was also true when Admixture was run on manto samples only (Supplementary Information 2). The leading axis of the PC decomposition, accounting for 8.8% of the total variation, similarly separated the two manto clusters, where subsequent statistical analyses showed a significant correlation between PC1 scores and longitude (F1,61 = 428.0, p < 0.001; R2 = 0.875; Fig. 3d). The PC1 axis for bubastis individuals only (Fig. 3e), accounting for 13.8% of the total variation, similarly showed a geographic clustering. However, a linear relationship between PC scores and longitude was only significant when the individuals from the French Alps were excluded (all bubastis: F1,19 = 0.6, p = 0.448; R2 = 0.031; Fig. 3e; Swiss Alps: F1,16 = 252.5, p < 0.001; R2 = 0.940). Despite the geographic clustering of the PC scores, Admixture did not detect any additional clusters when bubastis was run separately (Supplementary Information 2).
Summary of the individual based clustering analysis and their spatial relationship. A Individual-based assignments using Admixture for samples belonging to the E. manto complex. The case for four genetic clusters (K = 4) is shown as a bar diagram. B Map of all genotyped E. manto complex individuals (map modified from Bing Maps https://www.bing.com). C Principal component (PC) scores for all genotyped E. manto complex individuals. D–E Relationship between longitude and the PC1 scores for PC analyses done seperatenly on manto and bubastis individuals only. For B-E pie charts depict the Admixture assignment for each individual as shown in A
The clear separation of the three lineages with very limited gene flow in Admixture is further substantiated in our assessment of contemporary migration. The latter revealed very low migration rates (m) between the three lineages, often being close to zero (Fig. 4). Lastly, the overall degree of genetic differentiation was substantial among our three focal lineages (FST manto-bubastis = 0.544, FST manto-vogesiaca = 0.629, FST bubastis-vogesiaca = 0.823, all p < 0.001; Fig. 4). This contrasts with the level of intraspecific differentiation, i.e. between individuals of the two manto clusters that showed no admixture (FST = 0.116, p < 0.001).
Summary of the degree of genetic differentiation (FST) and recent migration (m) as estimated by BA3-SNPs. Black arrows indicate the direction of migration with the respective estimate of m ± 1 SD. For E. manto the FST between un-admixed individuals of the two inferred genomic clusters is given. All FST values are significant (p < 0.001)
The substrates on which manto and bubastis occurred in Switzerland differed: manto occurs predominantly on calcareous sedimentary rock (79.6%, N = 3'094, Fig. 5) and to a lesser degree on unconsolidated rock (14.0%, N = 543) and only rarely on siliceous crystalline rock (6.4%, N = 249). By contrast, bubastis did not show a preference for either siliceous crystalline (40.7%, N = 33; Fig. 5) or calcareous sedimentary rock (49.4%, N = 81); a small proportion of the observations were made on unconsolidated rock (9.9%, N = 8).
Discussion
A species complex represents a conundrum for both taxonomy and conservation as it combines putatively cryptic species or lineages into a single taxon (Bickford et al. 2007). Classic genetic markers often do not provide the resolution to resolve such complexes (Wagner et al. 2013). Using several thousand SNPs, we examined the genetic relationships of three lineages of the Erebia manto complex from central Europe, i.e., the manto, bubastis and vogesiaca lineages, respectively. Overall, we found that the three lineages are genetically strongly differentiated (Figs. 2, 3, 4) with very limited evidence for current gene flow (Fig. 4). Gene flow between lineages seems to have moreover occurred unidirectionally from bubastis into manto (Fig. 3). The level of genetic differentiation (FST) between these lineages is similar to interspecific comparisons in other and taxonomically resolved Erebia species in the Alps (Lucek et al. 2020) and greatly exceeds differentiation within a lineage, as found for manto (Fig. 4). A previous study based on analyses of the same genomic dataset found that the three E. manto lineages have a high prevalence of the endosymbiotic bacterium Wolbachia and share a similar Wolbachia strain (Lucek et al. 2021). Given the genomic differentiation of the hosts, this could implicate that they acquired a widely distributed Wolbachia strain only recently. Different to other Erebia species (Lucek et al. 2020, 2021), Wolbachia seems therefore unlikely to have significantly contributed to the differentiation of the three lineages of the manto complex.
The strong genomic differentiation that we found between manto and vogesiaca is consistent with former genetic inferences (Schmitt et al. 2014). With a much broader sampling, the aforementioned study further recovered three distinct genetic clusters within the manto lineage across the Alps, which have been interpreted to originate from distinct glacial refugia. The two genetic clusters that we identified for manto (Fig. 3) likely correspond to the Western and the Northern clusters described by Schmitt et al. (2014). However, our denser sampling for manto across the Swiss Alps revealed substantial admixture between these two clusters over a relatively large geographic range (Fig. 3b). This pattern suggests that the two manto clusters are not reproductively isolated and thus evolutionary less differentiated than either vogesiaca or bubastis from manto, confirming previous hypotheses that several levels of differentiation underlie the observed variation in this complex (Cupedo 1997). Inter-lineage gene flow seems to have primarily occurred from bubastis into manto (Figs. 3 & 4), which is consistent with previously reported introgression of mitochondrial haplotypes in the same direction (Cupedo and Doorenweerd 2020). Such asymmetries may occur when the degree of selection against gene flow differs between species, promoting unidirectional introgression (Pickup et al. 2019).
In Europe, the diversification of Erebia has been shaped by differentiation in distinct glacial refugia during the Quaternary glacial cycles, and the current distributions emerged through postglacial range expansions (Sonderegger 2005; Schmitt et al. 2006, 2016; Cupedo and Doorenweerd 2020). Distantly related Erebia species can often coexist and exploit different microhabitats (Kleckova et al. 2014). However, more closely related species or lineages may rather exclude each other to different degrees, which for Erebia falls into three broad scenarios. The first occurs when speciation is nearly completed, but rare events of introgressions may still be possible. An example is E. nivalis, which is found in near sympatry with E. tyndarus and E. cassioides but only exhibits very limited gene flow, possibly due to distinct chromosomal numbers, differing phenologies and micro-habitat preferences (Gratton et al. 2016; Ehl et al. 2018; Lucek et al. 2020). Other species pairs that fall under this scenario include E. ligea and E. euryale or E. eriphyle and E. manto (Sonderegger 2005; Cupedo 2014; Litman et al. 2018). The second scenario occurs in species pairs presenting a parapatric distribution, often only connected by very narrow contact zones and no morphological intergradation. This pattern is observed in E. tyndarus and E. cassioides in the Alps, both of which only meet in very restricted zones of contact (Sonderegger 2005; Lucek et al. 2020), where they form bimodal hybrid zones (Jiggins and Mallet 2000). Only rare first generation hybrids occur in such contact zones, suggesting a reduced hybrid fertility, possibly mediated by Wolbachia-induced incompatilibities also resulting in nearly completely interrupted gene flow (Lucek et al. 2020, 2021; Augustijnen et al. 2022). A similar situation is probably observed between E. melampus and E. sudetica in Switzerland, although no sympatric occurrence of these two taxa has been reported, at least in the Central Alps (Cupedo 1996); a genomic investigation of this complex is so far lacking (but see Haubrich and Schmitt 2007). A few additional species pairs of Erebia likely fall in these categories, such E. mnestra and E. aethiopella (Descimon and Mallet 2009) or E. montana and E. styx (Sonderegger 2005). The third scenario consists of more or less narrow contact zones, but not completely interrupted gene flow, leading to some morphological intergradation over a unimodal zone of secondary contact. Examples include contact zones between E. euryale subspecies that often have parapatric distributions, but show morphological intergradation (Sonderegger 2005; Cupedo 2014; Cupedo and Doorenweerd 2022) or between subspecies of E. melampus (Cupedo 1996).
Taxonomists have the somehow ungrateful task of translating this continuum into nomenclatural actions, an arbitrary, but necessary duty: delineated taxonomic units without names are de facto inexistant in biodiversity surveys or in conservation (Mace 2004). Current practices in Switzerland for the three scenarios outlined above mostly recognize species-level differentiations for cases that fall within the first or second scenario (SwissLepTeam 2010; Wermeille et al. 2014). The concept of "subspecies" similarly remains controversial, however, it seems to be best applied to well-delineated geographic units, either because of vicariance, or, as under the third abovementioned scenarios, narrow contact zone with limited, but existing morphological intergradation. Given our findings, the two E. manto lineages from the Alps, i.e., manto and bubastis, clearly fall under the second scenario given the absence of large-scale spatial overlap (Fig. 1; Sonderegger 2005) and the limited evidence for past interspecific gene flow (Figs. 3, 4).
The absence of coexistence in sympatry in the manto complex may on the one hand suggest that the two lineages lack enough differentiation in their ecology (Leibold and McPeek 2006) and/or phenotypic traits linked to mate choice (M’Gonigle et al. 2012) and thus occurrence in sympatry could result in interspecific mating with low offspring fitness. On the other hand, the three lineages could be ecologically differentiated and therefore exclude each other due to pronounced differences in habitat preferences. Ecological differentiation is likely limited given that in southwestern Switzerland E. manto and E. bubastis are separated by only 7–8 km (Fig. 1), without sharp geographic or climatic boundaries. Nevertheless, the vast majority of Swiss populations of manto seem to occur on calcareous substrate, while some populations of bubastis are located on or nearby siliceous substrate (Sonderegger 2005; Fig. 5). Our limited observations in France suggest that E. manto is similarly restricted to calcareous substrate while E. bubastis (at least the subspecies willieni) is found on siliceous substrate. Differences in geological substrate is a commonly used proxy to describe species distributions of Alpine butterflies (Illán et al. 2010; Augustijnen et al. 2022), however, what aspects of the environment may be causal in shaping the actual distributions is unknown. For example, larval development on Poaceae (the main host plants of Erebia) on specific substrates may require particular adaptations. The resulting distribution of E. manto and E. bubastis in the Alps could thus reflect the outcome of competitive exclusion to different substrates. Potential substrate association in E. vogesiaca requires though further investigation: while the Vosges populations are found on siliceous substrates, the possibly Jura populations are likely on calcareous substrate. In the Vosges, E. vogesiaca is ecologically highly specialized and restricted, living exclusively in the narrow area of the upper timberline, which in the Vosges is most formed by Sorbus shrubs. E. vogesiaca is however absent from the open pastures on the mountain ridges, as well as from the beech forests found at lower elevations.
Conclusions and implications for conservation
Taken together, our analyses show that the three lineages of the E. manto complex that we studied are genetically strongly isolated, supporting their status as distinct species, in agreement with the treatment in other similar cases (Gratton et al. 2016; Lucek et al. 2020). Speciation is an evolutionary process whereby barriers to gene flow accumulate through time until gene flow becomes impossible or strongly selected against (Seehausen et al. 2014; Stankowski and Ravinet 2021). The limited and unidirectional gene flow that we observed (Fig. 3) suggests that differentiation of the three lineages characterises an advanced stage of speciation (Kulmuni et al. 2020). Such limited gene flow is also possible in other, taxonomically well resolved Alpine butterfly species (Presgraves 2002; Descimon and Mallet 2009), that in some cases have diverged since the mid-Pleistocene (Ebdon et al. 2021). Although we did not perform cross experiments, our genomic inferences provide a surrogate to estimate the potential for hybridization in the wild, showing that the latter is absent and if still possibly, likely unidirectional and selected against. Based on our findings, we therefore recommend to treat the three lineages as distinct species, especially for conservation purposes. Climate change is predicted to significantly reduce the available habitat for E. manto in Central Europe (Schmitt et al. 2014). Given the restricted and disjunct distributions of E. bubastis and E. vogesiaca, these species may be especially vulnerable. Indeed, some populations of both E. bubastis (Wermeille et al. 2014) and especially E. vogesiaca (IMAGO 2014) are considered to be endangered, but taxonomic uncertainties have effectively precluded conservation measures so far. Our results also suggest that future conservation measures require to integrate fine scale ecology, given the possible difference in substrates on which E. manto and E. bubastis occur. Finally, our study highlights how genomic data may be used to overcome current taxonomic uncertainties that remain in several Alpine butterflies (Litman et al. 2018). This is especially true for the genus Erebia, which is known for its often cryptic diversity (Sonderegger 2005), where several candidates for potentially cryptic species have been identified (Tschudin et al. 2017).
Data availability
Sequence data is available from NCBI (BioProject PRJNA909280).
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664. https://doi.org/10.1101/gr.094052.109
Augustijnen H, Patsiou T, Lucek K (2022) Secondary contact rather than coexistence - Erebia butterflies in the Alps. Evolution 76:2669–2686
Bickford D, Lohman DJ, Sodhi NS, Ng PKL, Meier R, Winker K, Ingram KK, Das I (2007) Cryptic species as a window on diversity and conservation. Trends Ecol Evol 22:148–155. https://doi.org/10.1016/j.tree.2006.11.004
Burnaz S (2008) Endemits and rare species in the Lepidoptera collection of the Museum of Dacian and Roman Civilisation (Hunedoara County, Romania). Olten Stud Comun, Ştiinţ Nat 24:130–138
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) Stacks: an analysis tool set for population genomics. Mol Ecol 22:3124–3140. https://doi.org/10.1111/mec.12354
Ceballos G, Ehrlich PR, Dirzo R (2017) Biological annihilation via the ongoing sixth mass extinction signaled by vertebrate population losses and declines. Proc Natl Acad Sci USA 114:E6089–E6096. https://doi.org/10.1073/pnas.1704949114
Cupedo F (1996) Die morphologische Gliederung des Erebia melampus- Komplexes, nebst Beschreibung zweier neuer Unterarten: Erebia melampus semisudetica ssp.n. und Erebia sudetica belledonnae ssp.n. (Lepidoptera, Satyridae). Nota Lepi 18:95–125
Cupedo F (1997) Die geographische Variabilität und der taxonomische Status der Erebia manto bubastis-Gruppe, nebst Beschreibung einer neuen Unterart (Nymphalidae : Satyrinae). Nota Lepi 20:3–22
Cupedo F (2014) Reproductive isolation and intraspecific structure in Alpine populations of Erebia euryale (Esper, 1805) (Lepidoptera, Nymphalidae, Satyrinae). Nota Lepi 37:19–36. https://doi.org/10.3897/nl.37.7960
Cupedo F, Doorenweerd C (2020) The intraspecific structure of the Yellow-spotted ringlet Erebia manto (Denis & Schiffermüller, [1775]), with special reference to the bubastis group: an integration of morphology, allozyme and mtDNA data (Lepidoptera, Nymphalidae, Satyrinae). Nota Lepi 43:43–60. https://doi.org/10.3897/nl.43.47409
Cupedo F, Doorenweerd C (2022) Mitochondrial DNA-based phylogeography of the large ringlet Erebia euryale (Esper, 1805) suggests recurrent Alpine-Carpathian disjunctions during Pleistocene (Nymphalidae, Satyrinae). Nota Lepi 45:65–86. https://doi.org/10.3897/nl.45.68138
Danecek P, McCarthy SA (2017) BCFtools/csq: haplotype-aware variant consequences. Bioinformatics 33:2037–2039. https://doi.org/10.1093/bioinformatics/btx100
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Descimon H, Mallet J (2009) Bad species. In: Settele J, Shreeve T, Konvicka M, van Dyck H (eds) Ecology of butterflies in Europe, 1st edn. Cambridge University Press, pp 219–249
Dincă V, Dapporto L, Somervuo P et al (2021) High resolution DNA barcode library for European butterflies reveals continental patterns of mitochondrial genetic diversity. Commun Biol 4:315. https://doi.org/10.1038/s42003-021-01834-7
Dunn RR (2005) Modern insect extinctions, the neglected majority. Cons Biol 19:1030–1036. https://doi.org/10.1111/j.1523-1739.2005.00078.x
Ebdon S, Laetsch DR, Dapporto L, Hayward A, Ritchie MG, Dinca V, Vila R, Lohse K (2021) The Pleistocene species pump past its prime: evidence from European butterfly sister species. Mol Ecol 30:3575–3589. https://doi.org/10.1111/mec.15981
Ehl S, Dalstein V, Tull F, Gross P, Schmitt T (2018) Specialized or opportunistic-how does the high mountain endemic butterfly Erebia nivalis survive in its extreme habitats?: Population ecology of Erebia nivalis. Insect Sci 25:161–171. https://doi.org/10.1111/1744-7917.12400
Forister ML, Pelton EM, Black SH (2019) Declines in insect abundance and diversity: we know enough to act now. Conserv SciPrac 1:e80. https://doi.org/10.1111/csp2.80
Gratton P, Trucchi E, Trasatti A, Riccarducci G, Marta S, Allegrucci G, Cesaroni D, Sbordoni V (2016) Testing classical species properties with contemporary data: how “bad species” in the brassy ringlets (Erebia tyndarus complex, Lepidoptera) turned good. Syst Biol 65:292–303. https://doi.org/10.1093/sysbio/syv087
Gueuning M, Frey JE, Praz C (2020) Ultraconserved yet informative for species delimitation: ultraconserved elements resolve long-standing systematic enigma in Central European bees. Mol Ecol 29:4203–4220. https://doi.org/10.1111/mec.15629
Hallmann CA, Sorg M, Jongejans E, Siepel H, Hofland N, Schwan H, Stenmans W, Müller A, Sumser H, Hörren T, Goulson D, de Kroon H (2017) More than 75 percent decline over 27 years in total flying insect biomass in protected areas. Plos one. 12: e0185809. https://doi.org/10.1371/journal.pone.0185809
Haubrich K, Schmitt T (2007) Cryptic differentiation in alpine-endemic, high-altitude butterflies reveals down-slope glacial refugia. Mol Ecol 16:3643–3658. https://doi.org/10.1111/j.1365-294X.2007.03424.x
Hollander J, Montaño-Rendón M, Bianco G, Yang X, Westram AM, Duvaux L, Reid DG, Butlin RK (2018) Are assortative mating and genital divergence driven by reinforcement? Evol Lett 2:557–566. https://doi.org/10.1002/evl3.85
Illán JG, Gutiérrez D, Wilson RJ (2010) The contributions of topoclimate and land cover to species distributions and abundance: fine-resolution tests for a mountain butterfly fauna: determinants of butterfly distribution and abundance. Glob Ecol Biogeogr 19:159–173. https://doi.org/10.1111/j.1466-8238.2009.00507.x
Jakšic P (2019) A critical review of the current checklist of butterflies of Serbia. Univ Thought Publ Nat Sci 9:1–7. https://doi.org/10.5937/univtho9-19910
Jiggins CD, Mallet J (2000) Bimodal hybrid zones and speciation. Trends Ecol Evol 15:250–255. https://doi.org/10.1016/S0169-5347(00)01873-5
Kleckova I, Konvicka M, Klecka J (2014) Thermoregulation and microhabitat use in mountain butterflies of the genus Erebia: importance of fine-scale habitat heterogeneity. J Therm Biol 41:50–58. https://doi.org/10.1016/j.jtherbio.2014.02.002
Kudrna O, Pennerstorfer J, Lux K (2015) Distribution atlas of European butterflies and skippers. Wissenschaftlicher Verlag Peks, Schwanfeld
Kulmuni J, Butlin RK, Lucek K, Savolainen V, Westram AM (2020) Towards the completion of speciation: the evolution of reproductive isolation beyond the first barriers. Phil Trans R Soc B 375:20190528. https://doi.org/10.1098/rstb.2019.0528
Leibold MA, McPeek MA (2006) Coexistence of the niche and neutral perspectives in community ecology. Ecology 87:1399–1410
Lelo S (2000) Revised inventory of the butterflies of Bosnia and Herzegovina (Insecta: Lepidoptera: Hesperioidea: Papilionidea). Nat Croat 9:139–156
Litman J, Chittaro Y, Birrer S, Praz C, Wermeille E, Fluri M, Stalling T, Schmid S, Wyler S, Gonseth Y (2018) A DNA barcode reference library for Swiss butterflies and forester moths as a tool for species identification, systematics and conservation. PLoS ONE 13:e0208639. https://doi.org/10.1371/journal.pone.0208639
Lucek K, Butlin RK, Patsiou T (2020) Secondary contact zones of closely-related Erebia butterflies overlap with narrow phenotypic and parasitic clines. J Evol Biol 33:1152–1163. https://doi.org/10.1111/jeb.13669
Lucek K, Bouaouina S, Jospin A, Grill A, de Vos J (2021) Prevalence and relationship of endosymbiotic Wolbachia in the butterfly genus Erebia. BMC Ecol Evo 21:95. https://doi.org/10.1186/s12862-021-01822-9
Mace GM (2004) The role of taxonomy in species conservation. Phil Trans R Soc Lond B 359:711–719. https://doi.org/10.1098/rstb.2003.1454
Meirmans PG (2020) Genodive version 3.0: easy-to-use software for the analysis of genetic data of diploids and polyploids. Mol Ecol Resour 20:1126–1131. https://doi.org/10.1111/1755-0998.13145
M’Gonigle LK, Mazzucco R, Otto SP, Dieckmann U (2012) Sexual selection enables long-term coexistence despite ecological equivalence. Nature 484:506–509. https://doi.org/10.1038/nature10971
Mussmann SM, Douglas MR, Chafin TK, Douglas ME (2019) BA3-SNPs: contemporary migration reconfigured in BayesAss for next-generation sequence data. Methods Ecol Evol 10:1808–1813. https://doi.org/10.1111/2041-210X.13252
Pickup M, Brandvain Y, Fraïsse C et al (2019) Mating system variation in hybrid zones: facilitation, barriers and asymmetries to gene flow. New Phytol 224:1035–1047. https://doi.org/10.1111/nph.16180
Presgraves DC (2002) Patterns of postzygotic isolation in Lepidoptera. Evolution 56:1168–1183
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Sanz TS, Jiménez MM, Maestre MÁP, González JÁA (2017) Nueva cita de Erebia manto (Denis y Schiffermüller, 1775) (Lepidoptera: Nymphalidae) en la vertiente leonesa de los Picos de Europa (Cordillera Cantábrica, Norte de España). Aquivos Entomolóxicos 18:133–135
Schmitt T, Hewitt GM, Muller P (2006) Disjunct distributions during glacial and interglacial periods in mountain butterflies: Erebia epiphron as an example. J Evolution Biol 19:108–113. https://doi.org/10.1111/j.1420-9101.2005.00980.x
Schmitt T, Habel JC, Rödder D, Louy D (2014) Effects of recent and past climatic shifts on the genetic structure of the high mountain yellow-spotted ringlet butterfly Erebia manto (Lepidoptera, Satyrinae): a conservation problem. Glob Change Biol 20:2045–2061. https://doi.org/10.1111/gcb.12462
Schmitt T, Louy D, Zimmermann E, Habel JC (2016) Species radiation in the Alps: multiple range shifts caused diversification in Ringlet butterflies in the European high mountains. Org Divers Evol 16:791–808. https://doi.org/10.1007/s13127-016-0282-6
Seehausen O, Butlin RK, Keller I et al (2014) Genomics and the origin of species. Nat Rev Genet 15:176–192. https://doi.org/10.1038/nrg3644
Settele J, Kudrna O, Harpke A et al (2008) Climatic risk atlas of European butterflies. BioRisk 1:1–712. https://doi.org/10.3897/biorisk.1
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stankowski S, Ravinet M (2021) Defining the speciation continuum. Evolution 75:1256–1273. https://doi.org/10.1111/evo.14215
SwissLepTeam (2010) Die Schmetterlinge (Lepidoptera) der Schweiz: eine kommentierte, systematisch-faunistische Liste. Fauna Helvetica 25:1–350
Toews DPL, Brelsford A (2012) The biogeography of mitochondrial and nuclear discordance in animals. Mol Ecol 21:3907–3930. https://doi.org/10.1111/j.1365-294X.2012.05664.x
Vodă R, Dapporto L, Dincă V, Vila R (2015) Why do cryptic species tend not to co-occur? A case study on two cryptic pairs of butterflies. PLoS ONE 10:e0117802. https://doi.org/10.1371/journal.pone.0117802
Wagner CE, Keller I, Wittwer S, Selz OM, Mwaiko S, Greuter L, Sivasundar A, Seehausen O (2013) Genome-wide RAD sequence data provide unprecedented resolution of species boundaries and relationships in the Lake Victoria cichlid adaptive radiation. Mol Ecol 22:787–798. https://doi.org/10.1111/mec.12023
Warren BCS (1936) Monograph of the genus Erebia. British Museum (Natural History), London
Warren MS, Maes D, van Swaay CAM, Goffart P, van Dyck H, Bourn NAD, Wynhoff I, Ellis S (2021) The decline of butterflies in Europe: problems, significance, and possible solutions. Proc Natl Acad Sci USA 118:e2002551117. https://doi.org/10.1073/pnas.2002551117
Wiemers M, Balletto E, Dincă V et al (2018) An updated checklist of the European Butterflies (Lepidoptera, Papilionoidea). ZooKeys 811:9–45. https://doi.org/10.3897/zookeys.811.28712
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191. https://doi.org/10.1093/genetics/163.3.1177
IMAGO (2014) La Liste rouge des Rhopalocères et Zygènes menacés en Alsace. IMAGO, ODONAT
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. https://arxiv.org/13033997
R Core Team (2021) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Seibold S, Gossner MM, Simons NK, Blüthgen N, Müller J, Ambarli D, Ammer C, Bauhaus J, Fischer M, Habel JC, Linsenmair KE, Nauss T, Penone C, Prati D, Schall P, Schulze ED, Vogt J, Wöllauer S, Weisser WW (2019) Arthropod decline in grasslands and forests is associated with landscape-level drivers. Nature 574: 671–674. https://doi.org/10.1038/s41586-019-1684-3
Sonderegger P (2005) Die Erebien der Schweiz. P. Sonderegger, Brügg bei Biel
Tschudin P, Eggenberg S, Fivaz S, Jutzi M, Sanchez A, Schnyder N, Senn-Irlet B, Gonseth Y (2017) Endemiten der Schweiz – Methode und Liste 2017. Schlussbericht im Auftrag des Bundesamts für Umwelt (BAFU), Bern. p 37
van Klink R, Bowler DE. Gongalsky KB, Swengel AB, Gentile A, Chase JM (2020) Meta-analysis reveals declines in terrestrial but increases in freshwater insect abundances. Science. 368(6489): 417–420. https://doi.org/10.1126/science.aax9931
Wermeille E, Chittaro Y, Gonseth Y (2014) Rote Liste der Tagfalter und Widderchen: Papilionidea, Hesperioidea und Zygaenidae: Gefährdete Arten der Schweiz, Stand 2012. Bundesamt für Umwelt BAFU, Bern
Acknowledgements
We thank Luna Sartori and Emmanuel Rey for their help preparing the maps and Vincent Baudraz for helpful comments. Daniel Croll kindly provided bioinformatic support and allowed us to use his lab infrastructure. Sequencing was funded by info fauna and the Swiss Federal Office for the Environment (FOEN). KL was funded by the University of Basel and the Swiss National Science Foundation (Grant: PCEFP3_202869). Field work was funded by the Wüthrich & Matthey-Dupraz Foundation. Ulrich Hiermann kindly provided specimens of Liechtenstein and Austria. We thank the Amt für Umwelt Liechtenstein, the Bezirkshauptmannschaft Feldkirch, the Conservatoire d'Espaces Naturels de Lorraine, the services for fauna and nature of the cantons Bern, Grisons, Ticino, Uri, Valais and Vaud for granting collection permits.
Funding
Open access funding provided by University of Neuchâtel. This work was supported by the info fauna, the Swiss Federal Office for the Environment (FOEN), the Wüthrich & Matthey-Dupraz Foundation and the Swiss National Science Foundation (Grant PCEFP3_202869).
Author information
Authors and Affiliations
Contributions
AJ, YC, DB, DD, KG, JH, AS and CP: collected specimens; AJ and KL: analysed the data; AJ, YC, CP and KL: wrote the paper with input from all co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Confict of interest
The authors have no relevant financial or non-financial interest to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jospin, A., Chittaro, Y., Bolt, D. et al. Genomic evidence for three distinct species in the Erebia manto complex in Central Europe (Lepidoptera, Nymphalidae). Conserv Genet 24, 293–304 (2023). https://doi.org/10.1007/s10592-023-01501-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-023-01501-w