Abstract
Iron is an essential nutrient for bacteria, however its propensity to form toxic hydroxyl radicals at high intracellular concentrations, requires its acquisition to be tightly regulated. Ferric uptake regulator (Fur) is a metal-dependent DNA-binding protein that acts as a transcriptional regulator in maintaining iron metabolism in bacteria and is a highly interesting target in the design of new antibacterial drugs. Fur mutants have been shown to exhibit decreased virulence in infection models. The protein interacts specifically with DNA at binding sites designated as ‘Fur boxes’. In the present study, we have investigated the interaction between Fur from the fish pathogen Aliivibrio salmonicida (AsFur) and its target DNA using a combination of biochemical and in silico methods. A series of target DNA oligomers were designed based on analyses of Fur boxes from other species, and affinities assessed using electrophoretic mobility shift assay. Binding strengths were interpreted in the context of homology models of AsFur to gain molecular-level insight into binding specificity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Iron is an essential nutrient for all living organisms and many key biological processes are dependent on its abundance. For bacteria, iron is crucial for growth and host colonization. Iron mostly exists in the insoluble Fe3+ form under aerobic conditions at physiological pH and availability of the soluble reduced form, Fe2+, is restricted. Due to the ability of free iron to form toxic hydroxyl radicals through the Fenton reaction (Guerinot 1994), the essential high-affinity uptake systems of iron and iron homeostasis in bacteria must be tightly regulated, and in most bacteria, these processes are under control of the global metalloregulator Ferric uptake regulator (Fur) (Hantke 2001). Fur was first described in Escherichia coli (Hantke 1981), where it controls the expression of more than 90 genes, and its chemical properties and role in homeostasis has since been studied in homologs from multiple bacteria, including Mycobacterium smegmatis (Gao et al. 2019), Acidovorax citrulli (Liu et al. 2019), Campylobacter jejuni (Sarvan et al. 2019), Porphyromonas gingivalis (Smiga et al. 2019) and Salmonella enterica serovar Typhimurium (Wang et al. 2019). Although Fur was originally described as a repressor of genes coding for components of the ferric uptake systems found in the cell membrane, it is now understood to control the expression of toxins such as hemolysin and exotoxins, as well as proteins involved in iron-scavenging and uptake systems (Prince et al. 1991; Vasil and Ochsner 1999).
The typical model of action states that when intracellular levels of iron are high, dimeric Fur will act as a repressor by complexing Fe2+, binding specific Fur recognition sites in the promoter region and preventing transcription of associated genes involved in iron uptake, storage and metabolism. Similarly, when iron is limiting, the Fe-Fur complex dissociates from the promoter and allows gene expression. Recent studies however, have broadened our understanding of Fur-mediated regulation, indicating that Fur also may function as an activator and act in an iron-independent manner (Butcher et al. 2012; Miles et al. 2010), for example in Helicobacter pylori, all four combinations of Fur regulation have been characterized: repression and activation, with or without cofactor (Carpenter et al. 2009; Danielli and Scarlato 2010). Further, apo-Fur repression has been described in Staphylococcus aureus (Deng et al. 2012) and Campylobacter jejuni (Grabowska et al. 2011).
In addition to iron, which is the primary functional metal bound in vivo, DNA-binding by Fur can be activated by other divalent metals in vitro; Mn2+, Cu2+, Cd2+, Ni2+, Co2+ or Zn2+ (Bagg and Neilands 1987; de Lorenzo et al. 1987; Gao et al. 2008; Mills and Marletta 2005; Ochsner et al. 1995). Mn2+ is considered a suitable physiological mimic of Fe2+ for in vitro studies as it is bound with a similar affinity to Fe2+ by E. coli Fur and adopts the same hexacoordinated octahedral geometry using conserved residues, as seen in recent metal-bound crystal structures of Fur from Francisella tularensis (Deng et al. 2015; Mills and Marletta 2005; Perard et al. 2018). In contrast Zn2+ is bound with lower affinity and in a tetrahedral geometry (Deng et al. 2015; Perard et al. 2018).
Fur enacts its biological DNA-binding function as a dimer (Michaud-Soret et al. 1997), but may exist in several oligomeric states in solution, depending on protein concentration, salt concentration and pH (D'Autreaux et al. 2007). Each Fur monomer consists of two domains; an N-terminal winged helix-shaped domain involved in DNA binding (DNA binding domain; DBD) and a C-terminal α/β domain involved in protein dimerization (Dimerization domain; DD) (Supplementary Figure S1) (Hernandez et al. 2005; Pohl et al. 2003; Stojiljkovic and Hantke 1995). Crystal structures of apo- and holo-Fur have been available for some time from several bacterial species including Pseudomonas aeruginosa Fur (Pohl et al. 2003), F. tularensis Fur (Perard et al. 2018), Vibrio cholerae Fur (Sheikh and Taylor 2009), H. pylori Fur (Dian et al. 2011), C. jejuni Fur (Butcher et al. 2012), as well as a crystal structure of the DBD of E. coli Fur (Pecqueur et al. 2006); however, only with the recent structures of Magnetospirillum gryphiswaldense MSR-1 Fur (MgFur) in complex with DNA have structural details of Fur-DNA interactions become clear (Deng et al. 2015). A series of crystal structures, which include apo-Fur, holo-Fur and two different Fur-DNA complexes, gave a better understanding of issues regarding metal-binding, molecular mechanisms and structural basis of Fur-DNA interaction, at least for that organism.
The Fur-DNA interaction site, generally referred to as the “Fur box”, is a conserved sequence motif represented by a 19 base pair (bp) palindrome, located between the -35 and -10 sites at the promoters of Fur-regulated genes. The classical Fur box (Supplementary Figure S2a) originates from DNase I protection- and footprinting-experiments on E. coli Fur, where a Fur dimer recognizes a 19 bp inverted repeat sequence: 5′-GATAATGATAATCATTATC-3′ (Escolar et al. 1999), although this exact sequence is not found in the E. coli genome. This inverted repeat operator site was confirmed by binding of Fur to oligonucleotides inserted into a plasmid (Calderwood and Mekalanos 1988). In addition, Fur boxes from other genera have also been characterized and described (Baichoo and Helmann 2002; Escolar et al. 1998, 1999, 2000; Pedersen et al. 2010; Pich et al. 2012). While a recapture of the alternative arrangements of the Fur box and its interaction with Fur can be found in Supplementary text and Supplementary Figure S2, the binding modes observed from crystal structures of MgFur in complex with Fur box mimics are consistent with: a 9-1-9 inverted repeat model (Supplementary Figure S2a), where one MgFur dimer interacts; and a 7-1-7 heptamer inverted repeat model (Supplementary Figure S2e), where two MgFur dimers interact with DNA. The latter is consistent with a slightly extended Fur consensus sequence of 21 bp (Supplementary Figure S2e). Crystal structures determined for complexes of DtxR bound to its operator site show a similar binding model (Pohl et al. 1999; White et al. 1998).
In the search for the shortest recognition unit by Fur, the 7-1-7 inverted repeat was found to be the minimum in B. subtilis Fur (Baichoo and Helmann 2002). Single 6-mer or 7-mer oligonucleotides showed no affinity to Fur and Fur boxes with two 6-mers bound weakly. Similar results were obtained for E. coli Fur, where a minimum of three repeats of the hexameric motif GATAAT was required for Fur binding (Escolar et al. 2000). Thus, in the search for a Fur box consensus, the focus is shifting towards the functional pattern within the sequence, rather than the specific sequence or length. The consensus hexamer NATA/TAT appears to be the main unit of interaction with Fur, regardless of orientation and number. In addition, Fur boxes typically have a high content of A/T bases. Experimentally and computationally determined Fur boxes in various bacteria showed consensus sequence identity ranging from 50 to 80%. (Ahmad et al. 2009a; Baichoo et al. 2002; Pedersen et al. 2010; Sebastian et al. 2002; Thompson et al. 2002), and Fur appears to have a rather broad substrate-binding ability.
The published crystal structures of MgFur in complex with two different DNA targets demonstrate the lack of a well-defined sequence specificity, and a high degree of degeneration in the Fur box (Deng et al. 2015). DNase I footprinting with the feoAB1 operator showed a protected region without the typical arrays of GATAAT hexamers. However, for successful co-crystallization, the feoAB1 operator was mutated to a near-perfect inverted repeat, which bound one dimer of MgFur with similar binding affinities to the original feoAB1 operator. Gel shift-based assays showed that MgFur also specifically binds the P. aeruginosa Fur box, and furthermore, that two dimers of MgFur co-crystallized with the P. aeruginosa Fur box sequence (identical in sequence to the E. coli Fur box, which we will use throughout) (Deng et al. 2015). These MgFur-DNA complex structures are the first to demonstrate the ability to bind DNA at different ratios.
Common for these two rather different DNA targets is the way each Fur monomer formed contacts with both DNA strands using its DNA-binding domain (DBD), which interacted with a 10–11 bp sequence containing an important G base, conserved T base and an AT-rich region characterised by a narrow minor groove. In vivo experiments indicated that specific Fur-DNA contacts may be directly connected to DNA shape instead of being base specific. The positively charged Lys15 in MgFur bound this narrow minor groove with enhanced negative electrostatic potential. The narrow minor groove of AT-rich sequences is highlighted as an essential feature for Fur interaction (Deng et al. 2015).
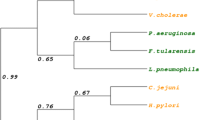
The Gram-negative Vibrionaceae family of gamma-proteobacteria include many mammalian pathogens, and the role of Fur and iron homeostasis in infection has received much attention due to its potential as a drug target (Jones and Oliver 2009; León-Sicairos et al. 2015; Mey et al. 2005; Wright et al. 1981). Amino acid alignments and phylogenetic analysis shows that the Fur protein is highly conserved within the Vibrionaceae, and in the present study we have investigated Fur from the Vibrio fish pathogen Aliivibrio salmonicida, the causative agent of cold-water vibriosis in Atlantic salmon and cod (Egidius et al. 1986). Previously, Thode et al. demonstrated a key role of A. salmonicida Fur (hereafter AsFur) in iron homeostasis (Thode et al. 2017) where construction of a fur null mutant strain severely affected fitness and growth of the bacteria, caused oxidative stress and a general reduced ability to cope with low-iron conditions. Furthermore, evaluation of expression levels compared to the wild-type identified up-regulation of numerous genes encoding for iron uptake and storage and down-regulation of potential targets for RyhB and other sRNAs involved in iron homeostasis (Thode et al. 2017). AsFur and its DNA target (Fur box) have previously been studied in vitro and in silico, with emphasis on identification of residues of importance for protein-DNA interactions (Ahmad et al. 2009b; Pedersen et al. 2010). A 19 bp inverted repeat Vibrio Fur box consensus, 5′-AATGATAATAATTATCATT-3′, was identified by computational methods (Ahmad et al. 2009b) and later shown to be specifically recognized and bound by AsFur in vitro in EMSA experiments with strong affinity (Pedersen et al. 2010). Additionally, specific individual nucleotides and amino acid residues possibly interacting in the AsFur-Vibrio Fur box complex have been predicted, some species-specific for AsFur. In the Vibrio Fur box, A14, C16 and T13 were suggested to contribute directly to the AsFur-DNA complex, on one or both strands. By homology modelling, the C16 nucleotide was predicted to be in close proximity to the amino acids Tyr56, Arg57 and Arg70, identified by binding free energy calculations (Pedersen et al. 2010). However, the base-specificity of these interactions remained elusive.
While these previous studies have mainly focused on investigating the effect of amino acid substitutions on DNA interaction, the present study aimed to elucidate the effect of nucleotide substitutions in the target DNA in an attempt to establish the binding mode of AsFur on Fur box-DNA.
In this study, we have characterized AsFur with respect to activity, thermal stability and its binding capability on a range of oligonucleotides in order to investigate the importance of key nucleotides in AsFur-DNA interaction.
Materials and methods
Cloning, expression and purification of AsFur
AsFur was overexpressed and purified with some changes from the previously described protocol (Pedersen et al. 2010). Following cloning of the fur gene from A. salmonicida into the pDEST14 Gateway expression vector (Invitrogen™, USA), AsFur was overexpressed at 20ºC overnight in E. coli BL21-CodonPlus® (DE3)-RIL competent cells, grown in LB broth with 100 µg/ml ampicillin and 34 µg/ml chloramphenicol. Harvested cells were resuspended in lysis buffer (Buffer A; 300 mM NaCl, 50 mM Tris–HCl, 5 mM β-mercaptoethanol, 10 mM Imidazole, pH 7.5). The histidine-rich AsFur was purified by Immobilized metal ion affinity chromatography (IMAC) on a 5 ml HisTrap HP column (GE Healthcare). Buffer A was used as the running buffer and Buffer B (300 mM NaCl, 50 mM Tris–HCl, 5 mM β-mercaptoethanol, 500 mM Imidazole, pH 7.5) as the elution buffer. The second purification step was performed using size-exclusion chromatography (SEC) on a Superdex 200 16/60 gel filtration column (GE Healthcare) equilibrated with Buffer A without Imidazole added. AsFur purity was verified by SDS-PAGE and protein concentration was determined by NanoDrop 2000c (Thermo Scientific) using the theoretical extinction coefficient.
Thermofluor
In order to improve the stability of the purified AsFur, thermal stability in various buffer systems and salt concentrations was investigated by a Thermofluor assay (Ericsson et al. 2006). Protein unfolding and its melting temperature (Tm) is monitored by using the fluoroprobe SYPRO Orange dye which emits fluorescence upon binding to exposed hydrophobic regions.
The buffer screen contained 24 buffers covering a pH range from 4.5 to 9.0. Briefly, 5 µl protein (2.5 mg/ml), 12.5 µl 2 × buffer solution (100 mM) and 7.5 µl 300 × SYPRO® Orange (Sigma Aldrich) were mixed and added to the wells of a 48-well PCR-plate (Bio-Rad). To assess the effect of various salts, 15 µl of protein (0.8 mg/ml) diluted in the appropriate buffer (Tris pH 7.5) were mixed with 7.5 µl of 300 × SYPRO® Orange (Sigma Aldrich) and 2.5 µl different salts in concentrations ranging from 0.1–2.0 M. The plates were sealed with Microseal® 'B' Adhesive Seals (Bio-Rad) and heated in a MiniOpticon Real-Time PCR System from 20 to 80 °C in increments of 1 °C per sec. Melting curves were monitored with a charge-coupled device (CCD) camera with wavelengths for excitation and emission at 490 and 575 nm, respectively. Tm, corresponding to the midpoint of the transition curve, was determined using the supplied instrument software and monitoring the fluorescence of the HEX channel.
DNA protection assay and the effect of metals on Fur binding
The capability of purified AsFur to bind DNA in the presence of various metals was investigated using a restriction site protection assay. The aerobactin plasmid pDT10, (kindly provided by Isabelle Michaud-Soret, Grenoble, France) carries four restriction enzyme sites, with the E. coli Fur box incorporated into one of the HinfI sites (D'Autreaux et al. 2002). Based on the method developed by Bagg and Neilands (Bagg and Neilands 1987), activated Fur binds the Fur box and thereby makes the restriction site unavailable for digestion by HinfI. Fur activity is confirmed by observing digestion patterns on gel electrophoresis. Fur is active if a 1781 bp band is observed, while two bands, respectively 1530 bp and 251 bp are observed if the protein is inactive. Supplementary Figure S3 (modified from (Cisse et al. 2014)) summarizes the principle behind the assay.
AsFur (20 µM) was incubated with two equivalents of a range of metals (40 µM) in binding buffer (100 mM BisTrisPropane pH 7.5, 100 mM KCl, 5 mM MgSO4) for 10 min at room temperature, followed by addition of pDT10 plasmid at 10 nM final concentration and 20 min additional incubation. Restriction enzyme digestion was carried out by adding 4 units per µl of HinfI to the mixture and incubating for 1 h at 37 °C before quenching with 0.5 mM of EDTA. The samples were run for 30 min at 100 V on 1% agarose gel in TAE and visualized under UV light.
Design of synthetic Fur box-containing oligonucleotides
To investigate the effect of length and specific base-substitutions to the Vibrio and E. coli respective Fur boxes, a range of oligonucleotides of various lengths (15–24 nt) were designed. In short, single-stranded DNA (Sigma-Aldrich) were diluted in Buffer C (50 mM HEPES pH 8, 50 mM NaCl) to 1 mM and annealed to double-stranded DNA by boiling for 5 min and cooling slowly to room temperature. Annealed oligonucleotides were separated by anion-exchange liquid-chromatography column (Mono-Q), with Buffer C as running buffer and Buffer D (50 mM HEPES pH 8, 1 M NaCl) as elution buffer, followed by dialyses in Slide-A-Lyzer Dialysis Cassettes (3.5 k MWCO; Thermo Scientific) overnight back to Buffer C. DNA concentrations were measured by Nanodrop 2000c (Thermo Scientific).
Electrophoretic gel mobility shift assay (EMSA)
Unlabelled Fur box-mimicking oligonucleotides were used as probes in EMSA assays, where DNA mobility is detected by the double stranded nucleic acid stain SybrGreen (Life Technologies), and slower mobility indicates that AsFur has complexed with the Fur box.
To complex AsFur with its DNA target, desired concentrations of purified AsFur were incubated with binding buffer (20 mM Tris acetate pH 8.0, 1 mM MgCl2, 50 mM KCl, 1 mM DTT and 100 µM MnCl2) at RT for 20 min. After addition of DNA (5 µM), the mixture was incubated for another 30 min before adding 10 × loading dye (30% glycerol in binding buffer). Samples were loaded on native 8% polyacrylamide/TB gels and electrophoresis was performed at 200 V for 2–2½ hours and at 6ºC with circulating 1 × TB buffer (89 mM Tris, 89 mM boric acid, pH 8.0). Finally, the gel was incubated with SybrGreen 1:10,000 in TB buffer for 20 min and band shifts were detected under UV light at ~ 254 nm. Binding strengths were examined and rated by visualization.
Analysis of AsFur compared to functional and structural homologs
Homology models of AsFur were generated using as templates the crystal structures of MgFur in complex with the E. coli Fur box (PDB4rb1) and the feoAB1 operator (PDB4rb3), respectively. The modelling tools of the Swiss-model repository were utilized in default mode to obtain the homology models. Both template structures contain Mn2+-ions in the regulatory S2 and structural S3 sites and were kept in the homology model. Although most likely present in AsFur, the structural S1 metal site generally coordinated by four Cysteine residues was not taken into account in the AsFur homology models as this site has a remote location to the DNA-interaction region. The two different interactions modes and stoichiometry of MgFur interacting with dsDNA were further analysed using WinCoot (Emsley et al. 2010) and visualized by PyMol (www.pymol.org). Conserved nucleotide base-protein interactions were highlighted from structure-based sequence alignments with homologous Fur crystal structures, rendered by ESPript 3.0 (Robert and Gouet 2014) and from the output from the NuProPlot server (Pradhan and Nam 2015).
Results and discussion
AsFur was purified to homogeneity through affinity- and size exclusion-chromatography
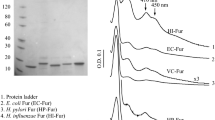
AsFur consists of 147 amino acid residues with theoretical pI and molecular weight of 5.75 and 16.6 kDa, respectively. A large-scale purification procedure of AsFur was established by Pedersen et al. (Pedersen et al. 2010). In brief, the fur gene from A. salmonicida was cloned, over-expressed in BL21-CodonPlus® (DE3)-RIL and purified to apparent homogeneity by two consecutive steps; IMAC affinity purification using HisTrap HP followed by SEC using HiLoad Superdex 200 pg. From SEC chromatography, AsFur fractions were detected at a volume corresponding to a homodimer, consistent with previous observations (Pedersen et al. 2010). The SDS-PAGE analysis of purified AsFur is shown in Fig. 1.
Thermal denaturation screening on pH and salt showed a slight effect on the stabilization of Fur
Although AsFur was purified to homogeneity as seen in Fig. 1, the initial protein batches showed a tendency to aggregate, with complete loss of binding activity within a week at standard storage conditions. To avoid protein aggregation and increase stability, a thermal shift assay (Thermofluor) was implemented to identify better buffer conditions. Screening of a range of buffer compositions (and pH) by Thermofluor only showed negligible effects on AsFur stability, however, activity assays indicated that a minor change in pH from 8.0 to 7.5 in Tris-buffer reduced aggregation and increased the storage stability of AsFur at 4 °C. Furthermore, a slight increase in NaCl concentration up to 200 mM showed a positive effect on AsFur stability (Fig. 2), in comparison to MgCl2 and KCl where only minor improvements could be seen. Although the storage stability of AsFur was improved by the above-mentioned changes in pH and NaCl concentrations, batch variations were still a frequent problem in the following characterization.
Presence of divalent metals alters DNA binding by AsFur
In the classical regulation pattern, iron is the primary functional metal that dimerizes and activates Fur in vivo. The ability of Fur to be effectively activated by a wide range of other divalent metal ions in vitro has prompted discussions about the true physiological metal responsible for Fur activation, although evaluation of Fur metal affinity by metal titration experiments suggests that only Fe2+ show sufficient affinity to activate Fur within relevant concentration ranges in vivo (Mills and Marletta 2005). However, elevated concentrations of other metals intracellularly could have implications for the normal iron regulation and the different metal bound Fur could potentially act on different DNA targets (Hantke 1987).
To measure the effect of a range of metals on AsFur-DNA binding, an in vitro assay utilising protection of a restriction site in the aerobactin promoter was used (D'Autreaux et al. 2002). Functional binding by Fur is envisaged by the absence of a fourth 251 bp band on the gel and an increase in size of the upper band to 1781 bp. Analysis showed that AsFur binds the aerobactin promoter in a metal-dependent fashion with Mn2+ present (Fig. 3a). Furthermore, the results in Fig. 3b show that AsFur also is able to bind the Fur box in presence of the divalent metal cations Mn2+, Zn2+, Cu2+ and Co2+. Fe2+ is considered the most physiologically-relevant metal ion for Fur activation, however it was omitted from this panel as its rapid oxidation precludes its use under standard assay conditions. Although plasmid protection appears weaker for Mn2+ compared to Zn2+, Cu2+ and in particular Co2+, Mn2+ was still the preferred choice for subsequent AsFur-DNA interaction studies, as Mn2+ and Fe2+ have been shown to have conserved metal coordination and similar chemical behavior in structural studies (Perard et al. 2018). The behaviour of AsFur in the presence of Cd2+ could not be interpreted, as the migration pattern resembles that of untreated plasmid, suggesting that HinfI is inhibited in the presence of Cd2+.
Plasmid protection assay verifying AsFur DNA binding in the presence of two equivalents of manganese (a) and identifying additional metals able to activate AsFur (b). The plasmid pDT10 was cleaved by HinfI in the absence or presence of active Fur and the digested migration pattern was analyzed by 1% gel electrophoresis. Active AsFur binds to the incorporated Fur box in the 1781 bp restriction fragment and protects it from being cleaved into 1530-bp and 251-bp fragments. a Lane 1: 1 kb ladder; Lane 2: plasmid pDT10; Lane 3: pDT10 + HinfI; Lane 4: pDT10 + apo AsFur + HinfI; Lane 5: pDT10 + AsFur + EDTA + HinfI; Lane 6: pDT10 + AsFur + Mn2+ + HinfI; Lane 7: pDT10 + EcFur + Mn2+ + HinfI. b Lane 1: plasmid pDT10 + apo AsFur + HinfI; Lane 2: pDT10 + As-Fur + Mn2+ + HinfI; Lane 3: pDT10 + AsFur + Zn2+ + HinfI; Lane 4: pDT10 + AsFur + Cu2+ + HinfI; Lane 5: pDT10 + AsFur + Co2+ + HinfI; Lane 6: pDT10 + AsFur + Cd2+ + HinfI
Oligonucleotides of different lengths and sequences have an effect on the Fur-DNA binding
The 19 bp inverted repeat Vibrio consensus sequence, 5′-AATGATAATAATTATCATT-3′, as well as the E. coli Fur box, 5′-GATAATGATAATCATTATC-3′, formed the templates for EMSA assays of a range of oligonucleotides (see Table 1 for oligonucleotide composition). Single bases and/or arrays of bases were mutated to examine possible important binding sites or important hexamer arrangements. Binding strengths were examined and rated.
The oligonucleotides were loosely grouped based on conservation and length. First, EMSA experiments were performed in order to verify interaction of AsFur with the consensus sequences from both Vibrio species and E. coli using the least-conserved Vibrio sequence (Pedersen et al. 2010) as a negative control (Fig. 4). Strong interaction was observed with the Vibrio consensus sequence (Fig. 4a), and moderate binding was also seen with the E. coli consensus sequence (Fig. 4b), with essentially no interaction detected with the Vibrio least-conserved (anti-consensus) sequence even at the highest AsFur concentration (Fig. 4c).
EMSA positive and negative controls. a Vibrio consensus sequence. b E. coli consensus sequence. c Vibrio anti-consensus sequence. 5 μM DNA was incubated with increasing concentrations of AsFur (0, 10, 20, 40 and 80 μM) for lanes 1–5, respectively. The experiments were performed in the presence of Mn2+
Subsequent EMSA experiments were run with oligonucleotides of varying content compared to the two different consensus sequences, in order to investigate the effect of specific base changes on interaction strength with AsFur. The primary targets for the experimental design were the AsFur-DNA interactions predicted by previous MD simulations and binding free energy calculations (Pedersen et al. 2010). In order to further investigate the existing Fur-DNA interaction models, different oligonucleotide lengths (both shorter and longer than the 19 bp Fur box) were probed, as well as variation in the oligonucleotide termini which were either blunt or included a 1 nucleotide overhang capable of forming a ‘sticky’ end with adjacent DNA substrates (Table 1). As expected, the oligonucleotides showed varying interaction strength with AsFur (Fig. 5). For most EMSA experiments, AsFur binding caused the substrate to be retained in the wells of the gel, which most likely reflects the tendency of AsFur to aggregate, possibly triggered by the initial DNA complex formation.
EMSA experiments on variants of Vibrio and E. coli consensus oligonucleotides modifying individual positions and/or length. For each experiment, 5 μM DNA was incubated with increasing concentrations of AsFur (0, 10, 20, 40 and 80 μM) for lanes 1–5, respectively. Positions labelled in red or dark green denote modifications with respect to the Vibrio species and E. coli consensus sequences, respectively. The experiments were performed in the presence of Mn2+. (Color figure online)
The T12G/T13G substitution reduces interaction of AsFur with DNA considerably (Fig. 5e) compared to the Vibrio consensus sequence (Fig. 4a), suggesting these are key positions for interaction. The corresponding nucleotide positions in the E. coli Fur box are T15 and T16 which have previously been shown to interact with E. coli Fur by crosslinking experiments (Tiss et al. 2005). Furthermore, molecular dynamics simulations with AsFur indicated T13 as an important contributor in protein interaction (Pedersen et al. 2010). We thus present the first EMSA experiments probing these positions directly in comparison to the Vibrio consensus sequence.
When the A14G/C16A substitution (Fig. 5f) is compared to the Vibrio consensus sequence (Fig. 4a), much reduced interaction capacity with AsFur is observed. These nucleotide positions have previously been shown to contribute favourably to AsFur DNA binding through binding free energy simulations (Pedersen et al. 2010), and the EMSA results further indicate them to participate in sequence-specific interactions. It is interesting to observe that the substitutions A17G/C19G (Fig. 5j) to the E. coli consensus sequence (Fig. 4b) also has a detrimental effect, although slightly less pronounced than for the Vibrio consensus sequence. Interestingly, the results highlight the importance of both DNA strands in Fur interaction, as these nucleotide positions form part of the first GATAAT hexamer repeat on the complementary strand of both the Vibrio consensus sequence and the E. coli Fur box (altered to TACAAT and CACAAT, respectively).
The A8C and T9G individual substitutions (Fig. 5b, c) also lead to much weaker interactions when compared to the Vibrio consensus sequence (Fig. 4a). AT-rich regions have previously been shown to be essential for Fur-DNA interactions (Hantke 1981, 2001; Prince et al. 1991; Vasil and Ochsner 1999). In particular, the last T base of the GATAAT (T6) unit in the hexamer repeat model described in Supplementary Figure S2b and c, corresponding to the substituted T9 in the Vibrio consensus sequence, has been highlighted for its role in DNA recognition by footprinting and missing-T assays with E. coli Fur (Escolar et al. 1998). The matching T on the complementary strand (T5) showed comparable effect in interactions. However, the combined substitutions A8C/T9G (Fig. 5d) does not show an additive effect and has slightly stronger interaction than the individual substitutions. It is interesting that the dual removal of AT-nucleotides in the core of the Vibrio recognition sequence does not appear to further reduce binding strength.
As expected, and in a similar fashion as for the Vibrio least conserved sequence (Fig. 4c) which showed almost no sign of DNA interaction with AsFur, the combined alteration of all nucleotides addressed so far (A8C/T9G/T12G/T13G/A14G/C16A; Fig. 5g) produced a much-weakened interaction with AsFur, although for this EMSA gel some trace amounts of AsFur can be seen shifted to the wells of the gel throughout.
For the A1G/A2C/T3A substitutions (Fig. 5a) compared to the Vibrio consensus sequence (Fig. 4a), substantially reduced interaction strength can be observed. This result is interesting in view of the differences in the 5′-regions of the Vibrio and E. coli consensus sequences where the Vibrio consensus sequence has a three-nucleotide ‘insertion’ (AAT) compared to the classical E. coli Fur box.
The importance of a minimum length of the Vibrio consensus sequence in AsFur interactions was demonstrated by the EMSA experiments on shortened oligonucleotides compared to the Vibrio consensus sequence (Fig. 5h, i), where both the 16-nucleotide sticky-end variant and the 15-nucleotide blunt-ended oligonucleotides displayed much-reduced binding strength compared to the Vibrio consensus sequence. This agrees well with previous EMSA studies on E. coli Fur indicating that only weak interaction is formed when oligonucleotides are considerably shorter than three hexamer repeats of the GATAAT sequence (Lavrrar and McIntosh 2003).
When somewhat similar experiments were performed on GATAAT hexamer repeats of the E. coli fur box, either with a 19-nucleotide sticky-end variant (Fig. 5k) or the 24-mer quadruple repeat of the GATAAT sequence (Fig. 5l), reduced interaction strengths were observed for both compared to the E. coli Fur box (Fig. 4b). As above, these trends correspond well with EMSA experiments on EcFur with the E. coli Fur box, as well as a range of GATAAT repeats, where the interaction strengths were rated as Fur box > 4 × GATAAT > 3 × GATAAT (Lavrrar and McIntosh 2003). The introduction of sticky ends to the triple GATAAT repeat in our study appears to improve binding slightly.
Analysis of AsFur compared to functional and structural homologs
To enable analysis of structural interactions contributing to specificity of binding between AsFur and variations on canonical Fur-box sequences, the sequence of AsFur was compared to structurally-determined homologs. A number of structurally-characterized homologs of AsFur were identified with sequence identities ranging from 86 to 30% (Table 2 and Fig. 6). The sequence alignment between these homologs and AsFur highlights several conserved sequence patches both within the DNA-binding- and dimerization domains (DBD and DD, respectively). Although most Fur proteins characterised are found to be dimers in solution, some also exist in the form of stable tetramers, exemplified by Fur from Francisella tularensis and Pseudomonas aeruginosa (Nader et al. 2019; Perard et al. 2018). While the conserved positions in the DD are mainly attributed to metal-coordination, the conserved patches in the DBD are involved in interactions with the Fur box (Fig. 6).
Structure-based sequence alignment of AsFur with known Fur structural homologs. Abbreviations are as defined in Table 2. EcFur was not included as the structure only represents the DBD. PDB identifiers are indicated between vertical lines. Secondary structure elements are shown above the alignment with spirals and arrows indicating α-helices and β-strands, respectively. Identical residues are shown in white on red background, while conserved residues are shown in red. Residues relevant for metal coordination to the regulatory site S2 and structural site S3 are indicated with triangles (coloured blue and red for S2 and S3, respectively), while residues forming base contacts are indicated with a black asterisk. (Color figure online)
Based on the sequence identity between V. cholerae Fur (VcFur) and AsFur at 86%, the crystal structure of VcFur (Sheikh and Taylor 2009) would be a preferred choice for homology modelling. However, this crystal structure represents Fur in an unbound state and structural alignments between the unbound and DNA-bound states of the MgFur structures revealed substantial movements in the DBD region upon DNA binding with VcFur and the MgFur DNA complexes having root-mean-square deviations of 2.6–3.0 Å. Thus, for homology modelling, the published structures of MgFur (Deng et al. 2015) were selected as templates despite the relatively low sequence similarity with AsFur. Sequence alignment between AsFur and MgFur revealed 37% identity for the 135 residues that could be structurally aligned and enabled reliable modelling of the entire protein including the N-terminal DNA-binding domain (DBD), which is highly flexible in the un-bound form of MgFur (Deng et al. 2015; Sarvan et al. 2018). To compare different possible binding modes of AsFur, two models were constructed: the first based on the MgFur dimer bound to a feoAB1 operator as a 9-1-9 inverted repeat (PDB4rb3; Supplementary Figure S2a) and the second based on the two MgFur dimers bound to an E. coli Fur box as a 7-1-7 inverted repeat offset by 6 nucleotides (PDB4rb1; Supplementary Figure S2e). Comparison of these models reveals a conservation in amino acids in the interacting regions of the two proteins (Fig. 7).
Homology models of AsFur in the two Fur-DNA interaction modes observed for MgFur and reported by Deng et al. (2015). a A dimer of AsFur interacting with the feoAB1 operator. b Two AsFur dimers interacting with the E. coli Fur box. Each AsFur monomer is coloured individually (monomer A, dark green; monomer B, turquoise; monomer C, purple; monomer D, red) and the DNA strands are coloured in dark yellow and blue for the primary and complementary strands, respectively. Nucleotides coloured in red indicate base contacts with AsFur. The modelled Mn2+ ions are indicated as grey spheres. (Color figure online)
Analysis of structural determinants of AsFur-DNA interaction
The homology models generated based on MgFur were analysed to structurally rationalize the variations in interaction strengths from EMSA.
These strongly indicate that AsFur Tyr56 forms base-specific major groove interactions through hydrophobic interactions with the methyl groups of both T12 and T13 (Fig. 8b), explaining the observed decrease in binding affinity in the T12G/T13G substitution. Interestingly, this interaction is conserved in both structural models, i.e. both in the forms of a 9-1-9 inverted repeat, as well as in the 7-1-7 inverted repeat offset by 6 nucleotides, highlighting the role of Tyr56 in interactions with Fur box-containing DNA.
Predicted AsFur-nucleotide base interactions from the homology model. Nucleotide base-interactions observed in the AsFur homology model based on the crystal structure of two dimers of MgFur in complex with the E. coli Fur box (PDB4rb1). Nucleotide numbering follows the numbering scheme used for the E. coli consensus sequence in Table 1. a Lys14 in monomer A interacts in the minor groove with T18 on the primary strand and T3′ on the complementary strand. b Tyr56 in monomer B forms hydrophobic interactions in the major groove with T15′ and T16′ on the complementary strand (identical interactions are formed between Tyr56 in monomer D, generated through a crystallographic symmetry operation, and T15/T16 on the primary strand). c Arg57 in monomer D interacts in the major groove with T18 on the primary strand and G1′ on the complementary strand. (Color figure online)
The homology model of AsFur interacting with the E. coli Fur box (Fig. 7b) also highlights the importance of both DNA strands in this interaction, providing a rationale for the impact of the A17G/C19G substitution (Fig. 5j) with nucleotide base-contacts formed in the 5′-end of the complementary strand. The detailed view of the interactions shown in Fig. 8a, c, illustrates that the nucleotide positions in the first hexamer repeat of the complementary strand (G1′ and T3′) form minor- and major-groove nucleotide base interactions with the conserved residues Lys14 and Arg57, respectively. Arg57 appears to form bidentate base-specific major-groove interactions, while the minor-groove interactions formed by Lys14 are base-unspecific. The corresponding Lys residue in MgFur has previously been shown to interact with the DNA target through a shape readout mechanism, where the AT-rich region in each hexamer repeat results in a narrow minor groove with enhanced negative electrostatic potential (Deng et al. 2015). Previous studies have indicated that most DNA-binding proteins use interplay between the base- and shape-readout modes to recognize their DNA binding sites (Slattery et al. 2014). This in turn allows for alterations in the specific nucleotide succession, and for the specific case of Fur, thus rationalises the degree of degeneracy found among Fur recognition sequences.
While Lys14, Tyr56 and Arg57 are also found to interact with nucleotide bases in the homology model of AsFur in complex with the feoAB1 operator (Fig. 7a), these interactions can not to the same extent justify a structural rationalisation of the above-mentioned effects from our EMSA experiments, making it less likely that AsFur interacts with the Vibrio consensus sequence and the E. coli Fur box as one dimer in the form of an 9-1-9 inverted repeat.
The AsFur homology model with the E. coli Fur box does not show direct contacts in the 5′-region of the Vibrio consensus sequence (upstream of the first GATAAT repeat), which is equivalent to the position of the substitution A1G/A2C/T3A; however, it is likely that Lys14 from the DBD of monomer B may undergo conformational changes in order to be involved in minor groove interactions. In fact, previous studies have suggested that the Fur box should be extended in the 5′-end, where Baichoo et al. (Baichoo and Helmann 2002) suggested an additional T extension in the B. subtilis Fur box and Chen et al. (Chen et al. 2007) that the E. coli Fur box should include the sequence AAT, i.e. identical to the Vibrio consensus sequence.
Conclusion
Fur has an important role in iron homeostasis and regulation of virulence mechanisms in many pathogenic bacteria. In an attempt to better understand the molecular basis behind DNA-recognition by AsFur, we have examined its DNA interaction with the combined use of interaction assays and structural modeling, which allowed for a structure/function interpretation of the biochemical results obtained.
AsFur was found to be a dimer during purification conditions. Due to protein instability issues, it was difficult to further investigate the stoichiometric rates on its interaction with different consensus sequences. However, the combined output of the homology modelling and the EMSA investigations indicate that AsFur will be able to interact in the form of two dimers.
The combined results of the EMSA experiments and homology models indicate that AsFur binding strength to DNA is stronger for longer oligonucleotides than shorter, and we observed a small increase in binding strength when sticky ends were introduced to the same oligo sequence. The results further showed that no single base mutations were crucial, and that only anti-consensus depleted binding completely. However, nucleotide positions T12 and T13 (T15 and T16 in E. coli) and A14 and C16 (A17 and T19 in E. coli) previously suggested to be in direct contact with Fur, lead to a markedly reduced binding strength between AsFur and DNA when mutated. This indicated that these bases were important for AsFur-DNA specific interaction. In addition, mutations of individual and dual AT bases in the core of the vibrio consensus sequence highlighted the importance of AT-rich regions for interaction with AsFur.
The interplay between base- and shape-readout modes, allowing degeneracy between Fur consensus sequences within and between bacteria, was also for AsFur important in binding site recognition. Similarity in Fur-DNA interaction mode between bacteria through base readout by conserved Tyrosine and Arginine residues and shape readout by conserved Lysine residue.
In summary, biochemical assays combined with structural modeling has provided further insight into the AsFur-DNA interaction mode.
References
Ahmad R, Brandsdal BO, Michaud-Soret I, Willassen NP (2009a) Ferric uptake regulator protein: binding free energy calculations and per-residue free energy decomposition. Proteins 75:373–386. https://doi.org/10.1002/prot.22247
Ahmad R, Hjerde E, Hansen GA, Haugen P, Willassen NP (2009b) Prediction and experimental testing of ferric uptake regulator regulons in vibrios. J Mol Microbiol Biotechnol 16:159–168. https://doi.org/10.1159/000128322
Bagg A, Neilands JB (1987) Ferric uptake regulation protein acts as a repressor, employing iron (II) as a cofactor to bind the operator of an iron transport operon in Escherichia coli. Biochemistry 26:5471–5477
Baichoo N, Helmann JD (2002) Recognition of DNA by Fur: a reinterpretation of the Fur box consensus sequence. J Bacteriol 184:5826–5832
Baichoo N, Wang T, Ye R, Helmann JD (2002) Global analysis of the Bacillus subtilis Fur regulon and the iron starvation stimulon. Mol Microbiol 45:1613–1629
Butcher J, Sarvan S, Brunzelle JS, Couture JF, Stintzi A (2012) Structure and regulon of Campylobacter jejuni ferric uptake regulator Fur define apo-Fur regulation. Proc Natl Acad Sci USA 109:10047–10052. https://doi.org/10.1073/pnas.1118321109
Calderwood SB, Mekalanos JJ (1988) Confirmation of the Fur operator site by insertion of a synthetic oligonucleotide into an operon fusion plasmid. J Bacteriol 170:1015–1017
Carpenter BM, Whitmire JM, Merrell DS (2009) This is not your mother's repressor: the complex role of fur in pathogenesis. Infect Immun 77:2590–2601. https://doi.org/10.1128/IAI.00116-09
Chen Z et al (2007) Discovery of Fur binding site clusters in Escherichia coli by information theory models. Nucleic Acids Res 35:6762–6777. https://doi.org/10.1093/nar/gkm631
Cisse C et al (2014) Inhibition of the ferric uptake regulator by peptides derived from anti-FUR peptide aptamers: coupled theoretical and experimental approaches. ACS Chem Biol 9:2779–2786. https://doi.org/10.1021/cb5005977
D'Autreaux B et al (2007) Reversible redox- and zinc-dependent dimerization of the Escherichia coli fur protein. Biochemistry 46:1329–1342. https://doi.org/10.1021/bi061636r
D'Autreaux B, Touati D, Bersch B, Latour JM, Michaud-Soret I (2002) Direct inhibition by nitric oxide of the transcriptional ferric uptake regulation protein via nitrosylation of the iron. Proc Natl Acad Sci USA 99:16619–16624. https://doi.org/10.1073/pnas.252591299
Danielli A, Scarlato V (2010) Regulatory circuits in Helicobacter pylori : network motifs and regulators involved in metal-dependent responses. FEMS Microbiol Rev 34:738–752. https://doi.org/10.1111/j.1574-6976.2010.00233.x
de Lorenzo V, Wee S, Herrero M, Neilands JB (1987) Operator sequences of the aerobactin operon of plasmid ColV-K30 binding the ferric uptake regulation (fur) repressor. J Bacteriol 169:2624–2630
Deng X, Sun F, Ji Q, Liang H, Missiakas D, Lan L, He C (2012) Expression of multidrug resistance efflux pump gene norA is iron responsive in Staphylococcus aureus. J Bacteriol 194:1753–1762. https://doi.org/10.1128/JB.06582-11
Deng Z et al (2015) Mechanistic insights into metal ion activation and operator recognition by the ferric uptake regulator. Nat Commun 6:7642. https://doi.org/10.1038/ncomms8642
Dian C et al (2011) The structure of the Helicobacter pylori ferric uptake regulator Fur reveals three functional metal binding sites. Mol Microbiol 79:1260–1275. https://doi.org/10.1111/j.1365-2958.2010.07517.x
Egidius E, Wiik R, Andersen K, Hoff KA, Hjeltnes B (1986) Vibrio salmonicida sp. nov., a new fish pathogen. Int J Syst Evol Microbiol 36:518–520. https://doi.org/10.1099/00207713-36-4-518
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Features and development of Coot. Acta Crystallogr D 66:486–501. https://doi.org/10.1107/S0907444910007493
Ericsson UB, Hallberg BM, Detitta GT, Dekker N, Nordlund P (2006) Thermofluor-based high-throughput stability optimization of proteins for structural studies. Anal Biochem 357:289–298. https://doi.org/10.1016/j.ab.2006.07.027
Escolar L, Perez-Martin J, de Lorenzo V (1998) Binding of the fur (ferric uptake regulator) repressor of Escherichia coli to arrays of the GATAAT sequence. J Mol Biol 283:537–547. https://doi.org/10.1006/jmbi.1998.2119
Escolar L, Perez-Martin J, de Lorenzo V (1999) Opening the iron box: transcriptional metalloregulation by the Fur protein. J Bacteriol 181:6223–6229
Escolar L, Perez-Martin J, de Lorenzo V (2000) Evidence of an unusually long operator for the fur repressor in the aerobactin promoter of Escherichia coli. J Biol Chem 275:24709–24714. https://doi.org/10.1074/jbc.M002839200
Gao CH, Wei WP, Tao HL, Cai LK, Jia WZ, Hu L, Yang M (2019) Cross-talk between the three furA orthologs in Mycobacterium smegmatis and the contribution to isoniazid resistance. J Biochem 166:237–243. https://doi.org/10.1093/jb/mvz030
Gao H et al (2008) The iron-responsive Fur regulon in Yersinia pestis. J Bacteriol 190:3063–3075. https://doi.org/10.1128/jb.01910-07
Grabowska AD et al (2011) Campylobacter jejuni dsb gene expression is regulated by iron in a Fur-dependent manner and by a translational coupling mechanism. BMC Microbiol 11:166. https://doi.org/10.1186/1471-2180-11-166
Guerinot ML (1994) Microbial iron transport. Annu Rev Microbiol 48:743–772. https://doi.org/10.1146/annurev.mi.48.100194.003523
Hantke K (1981) Regulation of ferric iron transport in Escherichia coli K12: isolation of a constitutive mutant. Mol Gen Genet 182:288–292
Hantke K (1987) Selection procedure for deregulated iron transport mutants (fur) in Escherichia coli K 12: fur not only affects iron metabolism. Mol Gen Genet 210:135–139
Hantke K (2001) Iron and metal regulation in bacteria. Curr Opin Microbiol 4:172–177
Hernandez JA et al (2005) The conformational stability and thermodynamics of Fur A (ferric uptake regulator) from Anabaena sp. PCC 7119. Biophys J 89:4188–4200. https://doi.org/10.1529/biophysj.105.065805
Jones MK, Oliver JD (2009) Vibrio vulnificus: disease and pathogenesis. Infect Immun 77:1723–1733. https://doi.org/10.1128/IAI.01046-08
Lavrrar JL, McIntosh MA (2003) Architecture of a fur binding site: a comparative analysis. J Bacteriol 185:2194–2202
León-Sicairos N, Angulo-Zamudio U, de la Garza M, Velazquez-Roman J, Flores-Villaseñor H, Canizalez-Roman A (2015) Strategies of Vibrio parahaemolyticus to acquire nutritional iron during host colonization. Front Microbiol 6: https://doi.org/10.3389/fmicb.2015.00702
Liu J, Tian Y, Zhao Y, Zeng R, Chen B, Baishi H, Walcott RRP (2019) Ferric uptake regulator (FurA) is required for Acidovorax citrulli virulence on watermelon. Phytopathology. https://doi.org/10.1094/PHYTO-05-19-0172-R
Mey AR, Wyckoff EE, Kanukurthy V, Fisher CR, Payne SM (2005) Iron and fur regulation in Vibrio cholerae and the role of fur in virulence. Infect Immun 73:8167–8178. https://doi.org/10.1128/IAI.73.12.8167-8178.2005
Michaud-Soret I, Adrait A, Jaquinod M, Forest E, Touati D, Latour JM (1997) Electrospray ionization mass spectrometry analysis of the apo- and metal-substituted forms of the Fur protein. FEBS Lett 413:473–476
Miles S, Carpenter BM, Gancz H, Merrell DS (2010) Helicobacter pylori apo-Fur regulation appears unconserved across species. J Microbiol 48:378–386. https://doi.org/10.1007/s12275-010-0022-0
Mills SA, Marletta MA (2005) Metal binding characteristics and role of iron oxidation in the ferric uptake regulator from Escherichia coli. Biochemistry 44:13553–13559. https://doi.org/10.1021/bi0507579
Nader S, Perard J, Carpentier P, Arnaud L, Crouzy S, Michaud-Soret I (2019) New insights into the tetrameric family of the Fur metalloregulators. Biometals 32:501–519. https://doi.org/10.1007/s10534-019-00201-8
Ochsner UA, Vasil AI, Vasil ML (1995) Role of the ferric uptake regulator of Pseudomonas aeruginosa in the regulation of siderophores and exotoxin A expression: purification and activity on iron-regulated promoters. J Bacteriol 177:7194–7201. https://doi.org/10.1128/jb.177.24.7194-7201.1995
Pecqueur L et al (2006) Structural changes of Escherichia coli ferric uptake regulator during metal-dependent dimerization and activation explored by NMR and X-ray crystallography. J Biol Chem 281:21286–21295. https://doi.org/10.1074/jbc.M601278200
Pedersen HL et al (2010) Experimental and computational characterization of the ferric uptake regulator from Aliivibrio salmonicida (Vibrio salmonicida). J Microbiol 48:174–183. https://doi.org/10.1007/s12275-010-9199-5
Perard J et al (2018) Structural and functional studies of the metalloregulator Fur identify a promoter-binding mechanism and its role in Francisella tularensis virulence. Commun Biol 1:93. https://doi.org/10.1038/s42003-018-0095-6
Pich OQ, Carpenter BM, Gilbreath JJ, Merrell DS (2012) Detailed analysis of Helicobacter pylori Fur-regulated promoters reveals a Fur box core sequence and novel Fur-regulated genes. Mol Microbiol 84:921–941. https://doi.org/10.1111/j.1365-2958.2012.08066.x
Pohl E, Holmes RK, Hol WG (1999) Crystal structure of a cobalt-activated diphtheria toxin repressor-DNA complex reveals a metal-binding SH3-like domain. J Mol Biol 292:653–667. https://doi.org/10.1006/jmbi.1999.3073
Pohl E, Haller JC, Mijovilovich A, Meyer-Klaucke W, Garman E, Vasil ML (2003) Architecture of a protein central to iron homeostasis: crystal structure and spectroscopic analysis of the ferric uptake regulator. Mol Microbiol 47:903–915
Pradhan L, Nam HJ (2015) NuProPlot: nucleic acid and protein interaction analysis and plotting program. Acta Crystallogr D 71:667–674. https://doi.org/10.1107/S1399004715000139
Prince RW, Storey DG, Vasil AI, Vasil ML (1991) Regulation of toxA and regA by the Escherichia coli fur gene and identification of a Fur homologue in Pseudomonas aeruginosa PA103 and PA01. Mol Microbiol 5:2823–2831
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42:W320–324. https://doi.org/10.1093/nar/gku316
Sarvan S, Butcher J, Stintzi A, Couture JF (2018) Variation on a theme: investigating the structural repertoires used by ferric uptake regulators to control gene expression. Biometals 31:681–704. https://doi.org/10.1007/s10534-018-0120-8
Sarvan S, Yeung A, Charih F, Stintzi A, Couture JF (2019) Purification and characterization of Campylobacter jejuni ferric uptake regulator. Biometals 32:491–500. https://doi.org/10.1007/s10534-019-00177-5
Sebastian S, Agarwal S, Murphy JR, Genco CA (2002) The gonococcal fur regulon: identification of additional genes involved in major catabolic, recombination, and secretory pathways. J Bacteriol 184:3965–3974
Sheikh MA, Taylor GL (2009) Crystal structure of the Vibrio cholerae ferric uptake regulator (Fur) reveals insights into metal co-ordination. Mol Microbiol 72:1208–1220. https://doi.org/10.1111/j.1365-2958.2009.06718.x
Slattery M, Zhou T, Yang L, Dantas Machado AC, Gordan R, Rohs R (2014) Absence of a simple code: how transcription factors read the genome. Trends Biochem Sci 39:381–399. https://doi.org/10.1016/j.tibs.2014.07.002
Smiga M, Bielecki M, Olczak M, Olczak T (2019) Porphyromonas gingivalis PgFur is a member of a novel fur subfamily with non-canonical function. Front Cell Infect Microbiol 9:233. https://doi.org/10.3389/fcimb.2019.00233
Stojiljkovic I, Hantke K (1995) Functional domains of the Escherichia coli ferric uptake regulator protein (Fur). Mol Gen Genet 247:199–205
Thode SK, Baekkedal C, Soderberg JJ, Hjerde E, Hansen H, Haugen P (2017) Construction of a fur null mutant and RNA-sequencing provide deeper global understanding of the Aliivibrio salmonicida Fur regulon. PeerJ 5:e3461. https://doi.org/10.7717/peerj.3461
Thompson DK et al (2002) Transcriptional and proteomic analysis of a ferric uptake regulator (fur) mutant of Shewanella oneidensis: possible involvement of fur in energy metabolism, transcriptional regulation, and oxidative stress. Appl Environ Microbiol 68:881–892
Tiss A, Barre O, Michaud-Soret I, Forest E (2005) Characterization of the DNA-binding site in the ferric uptake regulator protein from Escherichia coli by UV crosslinking and mass spectrometry. FEBS Lett 579:5454–5460. https://doi.org/10.1016/j.febslet.2005.08.067
Vasil ML, Ochsner UA (1999) The response of Pseudomonas aeruginosa to iron: genetics, biochemistry and virulence. Mol Microbiol 34:399–413
Wang S et al (2019) The ferric uptake regulator represses type VI secretion system function by binding directly to the clpV promoter in Salmonella enterica serovar Typhimurium. Infect Immun 87: https://doi.org/10.1128/IAI.00562-19
White A, Ding X, van der Spek JC, Murphy JR, Ringe D (1998) Structure of the metal-ion-activated diphtheria toxin repressor/tox operator complex. Nature 394:502–506. https://doi.org/10.1038/28893
Wright AC, Simpson LM, Oliver JD (1981) Role of iron in the pathogenesis of Vibrio vulnificus infections. Infect Immun 34:503–507
Acknowledgements
Open Access funding provided by UiT The Arctic University of Norway. We would like to acknowledge Dr. Isabelle Michaud-Soret for donating the pDT10 plasmid. We are grateful to Dr. Marcin Pierechod for his assistance in biophysical characterisation attempts and Dr. Adele Williamson for proofreading the manuscript.
Author information
Authors and Affiliations
Contributions
IL had the original project idea. IL and HLP have been involved in the overall planning of the study. KB has been primarily responsible in planning and performing the experiments, such as protein purification, plasmid protection assay, design of oligonucleotides, EMSA assays and thermal stability screenings. KB, IL and HLP have contributed to analysis of biochemical data and writing the paper. IL has been responsible for homology modelling and structural analysis thereof.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Berg, K., Pedersen, H.L. & Leiros, I. Biochemical characterization of ferric uptake regulator (Fur) from Aliivibrio salmonicida. Mapping the DNA sequence specificity through binding studies and structural modelling. Biometals 33, 169–185 (2020). https://doi.org/10.1007/s10534-020-00240-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10534-020-00240-6