Abstract
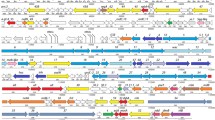
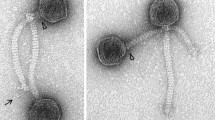
This study characterizes a cryptic (pro)phage-related sequence within the Caldibacillus debilis GB1 genome, designated CBP1.CBP1 is a Siphoviridae-like genome highly related to GBVS1 from Geobacillus sp. 6k51. The CBP1genome is a 37,315 bp region containing 69 putative ORFs with a GC content of 42% flanked on both sides by host DNA integrated into the main bacterial chromosome (contig 16). Bioinformatic analyses identified cassettes of genes within the CBP1 genome that were similar in function, yet distinct in sequence, from genes previously identified in GBVS1. All of CBP1 genes had less than 60% amino acid sequence identity with GBVS1by tBLASTx, with the exception of the TMP repeat gene. CBP1 possessed all the necessary genes to undergo a temperate/lytic phage life cycle, including excision, replication, structural genes, DNA packaging, and cell lyses. Proteomic analysis of CBP1 revealed the expression of 5 proteins. One of the expressed proteins was a transcriptional regulator protein homologous to the bacteriophage λ repressor protein (cI) expressed in high amounts from the CBP1 region, consistent with a lysogenic phage in a repressed state. The CBP1 protein expression profile during host growth provides unique insight into thermophilic Siphoviridae-like phages in the repressed state within their host cells.

Similar content being viewed by others
References
Ackers GK, Johnson AD, Shea MA (1982) Quantitative model for gene regulation by lambda phage repressor. Proc Nat Acad Sci 79:1129–1133
Banat I, Marchant R, Rahman T (2004) Geobacillus debilis sp. nov., a novel obligately thermophilic bacterium isolated from a cool soil environment, and reassignment of Bacillus pallidus to Geobacillus pallidus comb. nov. Int J Syst Evol Microbiol 54:2197–2201. https://doi.org/10.1099/ijs.0.63231-0
Bohannan BJ, Lenski RE (2000) Linking genetic change to community evolution: insights from studies of bacteria and bacteriophage. Ecol Lett 3:362–377
Brüssow H, Desiere F (2001) Comparative phage genomics and the evolution of Siphoviridae: insights from dairy phages. Mol Microbiol 39:213–223. https://doi.org/10.1046/j.1365-2958.2001.02228.x
Coorevits A, Dinsdale AE, Halket G, Lebbe L, De Vos P, Van Landschoot A, Logan NA (2012) Taxonomic revision of the genus Geobacillus: emendation of Geobacillus, G. stearothermophilus, G. jurassicus, G. toebii, G. thermodenitrificans and G. thermoglucosidans (nom. corrig., formerly ‘thermoglucosidasius’); transfer of Bacillus thermantarcticus to the genus as G. thermantarcticus comb. nov.; proposal of Caldibacillus debilis gen. nov., comb. nov.; transfer of G. tepidamans to Anoxybacillus as A. tepidamans comb. nov.; and proposal of Anoxybacillus caldiproteolyticus sp. nov. Int J Syst Evol Microbiol 62:1470–1485. https://doi.org/10.1099/ijs.0.030346-0
Doi K, Mori K, Martono H, Nagayoshi Y, Fujino Y, Tashiro K, Ohshima T (2013) Draft genome sequence of Geobacillus kaustophilus GBlys, a lysogenic strain with bacteriophage ϕOH2. Gen Ann 1:e00634-13. https://doi.org/10.1128/genomea.00634-13
Hargreaves KR, Colvin HV, Patel KV, Clokie JJP, Clokie MR (2013) Genetically diverse Clostridium difficile strains harboring abundant prophages in an estuarine environment. Appl Environ Microbiol 79:6236–6243
Horgan M, O’Sullivan O, Coffey A, Fitzgerald GF, van Sinderen D, McAuliffe O, Ross RP (2010) Genome analysis of the Clostridium difficile phage ΦCD6356, a temperate phage of the Siphoviridae family. Gene 462:34–43. https://doi.org/10.1016/j.gene.2010.04.010
Islam R, Cicek N, Sparling R, Levin D (2006) Effect of substrate loading on hydrogen production during anaerobic fermentation by clostridium thermocellum 27405. Appl Microbiol Biotechnol 72:576–583. https://doi.org/10.1007/s00253-006-0316-7
Liu B, Zhang X (2008) Deep-sea thermophilic Geobacillus bacteriophage GVE2 transcriptional profile and proteomic characterization of virions. Appl Microbiol Biotechnol 80:697–707. https://doi.org/10.1007/s00253-008-1575-2
Liu B, Suijie W, Qing S, Xiaobo Z, Lianhui X (2006) Two novel bacteriophages of thermophilic bacteria isolated from deep-sea hydrothermal fields. Curr Microbiol 53:163–166. https://doi.org/10.1007/s00284-005-0509-9
Liu B, Zhou F, Wu S, Xu Y, Zhang X (2009) Genomic and proteomic characterization of a thermophilic Geobacillus bacteriophage GBSV1. Res Microbiol 160:166–171. https://doi.org/10.1016/j.resmic.2008.12.005
Lucchini S, Desiere F, Brüssow H (1998) The structural gene module in Streptococcus thermophilus bacteriophage φSfi11 shows a hierarchy of relatedness to Siphoviridae from a wide range of bacterial hosts. Virology 246:63–73. https://doi.org/10.1006/viro.1998.9190
Nariya H, Miyata S, Tamai E, Sekiya H, Maki J, Okabe A (2011) Identification and characterization of a putative endolysin encoded by episomal phage phiSM101 of Clostridium perfringens. Appl Microbiol Biotechnol 90:1973–1979
Ochman H, Lawrence JG, Groisman EA (2000) Lateral gene transfer and the nature of bacterial innovation. Nature 405:299–304
Sidhu SS (2000) Phage display in pharmaceutical biotechnology. Curr Opin Biotechnol 11:610–616. https://doi.org/10.1016/S0958-1669(00)00152-X
Sutherland IW, Hughes KA, Skillman LC, Tait K (2004) The interaction of phage and biofilms. FEMS Microbiol Lett 232:1–6. https://doi.org/10.1016/S0378-1097(04)00041-2
Wang Y, Zhang X (2010) Genome analysis of deep-sea thermophilic phage D6E. Appl Environ Microbiol 76:7861–7866
Weitz JS, Wilhelm SW (2012) Ocean viruses and their effects on microbial communities and biogeochemical cycles. Biology Rep 4:17
Wushke S, Levin DB, Cicek N, Sparling R (2013) Characterization of enriched aerotolerant cellulose-degrading communities for biofuels production using differing selection pressures and inoculum sources. Can J Microbiol 59:679–683. https://doi.org/10.1139/cjm-2013-0430
Wushke S, Levin DB, Cicek N, Sparling R (2015) Characterization of the facultative anaerobe Caldibacillus debilis GB1 and its use in a designed aerotolerant, cellulose degrading, co-culture with Clostridium thermocellum. Appl Environ Microbiol. https://doi.org/10.1128/AEM.00735-15
Wushke S, Spicer V, Zhang X, Fristensky B, Krokhin OV, Levin DB, Cicek N, Sparling R (2017) Understanding aerobic/anaerobic metabolism in Caldibacillus debilis through a comparison with model organisms. Syst Appl Microbiol 40(5):245–253. https://doi.org/10.1016/j.syapm.2017.03.004
Yoon BH, Hyo IC (2011) Complete genomic sequence of the Lactobacillus temperate phage LF1. Arch Virol 156(10):1909
Zdobnov EM, Apweiler R (2001) InterProScan—an integration platform for the signature recognition methods in InterPro. Bioinformatics 17:847–848
Zhou Y, Liang Y, Lynch KH, Dennis JJ, Wishart DS (2011) PHAST: a fast phage search tool. Nucleic Acids Res. https://doi.org/10.1093/nar/gkr485
Acknowledgements
This work was funded by Genome Canada, through the Applied Genomics Research in Bioproducts or Crops (ABC) program for the grant titled, “Microbial Genomics for Biofuels and Co-Products from Biorefining Processes” as well as a Natural Science and Engineering Research Council (NSERC) Discovery grant to RS. SW was also supported through a University of Manitoba Graduate Fellowship (UMGF) award.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Atomi.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wushke, S., Jin, Z., Spicer, V. et al. Description of a cryptic thermophilic (pro)phage, CBP1 from Caldibacillus debilis strain GB1. Extremophiles 22, 203–209 (2018). https://doi.org/10.1007/s00792-017-0988-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-017-0988-1