Abstract
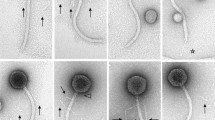
The Siphoviridae phage PMBT6 was identified by transmission electron microscopy in the supernatant of Bifidobacterium thermophilum MBT94004 bioreactor fermentation culture, where it occurred at a moderately high titer. Genome analysis of the bacterial DNA confirmed the presence of this prophage within the genome of the lysogenic host. Under laboratory conditions, the prophage could not be induced by mitomycin C, ultraviolet C irradiation or hydrogen peroxide, suggesting that the prophage was released by spontaneous induction under (yet unknown) bioreactor conditions. Genome sequencing of the virion resulted in a linear, double-stranded DNA molecule of 36,561 bp with a mol% G + C content of 61.7 and 61 predicted open reading frames with low similarity to other Bifidobacterium spp. genomes, confirming that PMBT6 represents a novel temperate phage for this genus.


Similar content being viewed by others
References
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NewYork
Kearse M, Moir R, Wilson A et al (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28(12):1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Nurk S, Bankevich A, Antipov D et al. (2013) Assembling genomes and mini-metagenomes from highly chimeric reads. In: Deng M, Jiang R, Sun F et al. (eds) Research in computational molecular biology: 17th Annual International Conference, RECOMB 2013, Beijing, China, April 7–10, 2013. Proceedings, vol 7821. Springer Berlin Heidelberg; Imprint; Springer, Berlin, Heidelberg, pp 158–170
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Aziz RK, Bartels D, Best AA et al (2008) The RAST server: rapid annotations using subsystems technology. BMC Genom 9:75. https://doi.org/10.1186/1471-2164-9-75
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Letunic I, Bork P (2018) 20 years of the SMART protein domain annotation resource. Nucleic Acids Res 46(D1):D493–D496. https://doi.org/10.1093/nar/gkx922
Besemer J, Lomsadze A, Borodovsky M (2001) GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res 29(12):2607–2618
Zhou Y, Liang Y, Lynch KH et al (2011) PHAST: a fast phage search tool. Nucleic Acids Res 39((Web Server issue)):W347-52. https://doi.org/10.1093/nar/gkr485
Lugli GA, Milani C, Turroni F et al (2016) Prophages of the genus Bifidobacterium as modulating agents of the infant gut microbiota. Environ Microbiol 18(7):2196–2213. https://doi.org/10.1111/1462-2920.13154
Mahony J, Lugli GA, van Sinderen D et al (2018) Impact of gut-associated bifidobacteria and their phages on health: two sides of the same coin? Appl Microbiol Biotechnol 102(5):2091–2099. https://doi.org/10.1007/s00253-018-8795-x
Ventura M, Lee J-H, Canchaya C et al (2005) Prophage-like elements in bifidobacteria: insights from genomics, transcription, integration, distribution, and phylogenetic analysis. Appl Environ Microbiol 71(12):8692–8705. https://doi.org/10.1128/AEM.71.12.8692-8705.2005
Ventura M, Turroni F, Foroni E et al (2010) Analyses of bifidobacterial prophage-like sequences. Antonie Van Leeuwenhoek 98(1):39–50. https://doi.org/10.1007/s10482-010-9426-4
Ventura M, Turroni F, Lima-Mendez G et al (2009) Comparative analyses of prophage-like elements present in bifidobacterial genomes. Appl Environ Microbiol 75(21):6929–6936. https://doi.org/10.1128/AEM.01112-09
Mavrich TN, Casey E, Oliveira J et al (2018) Characterization and induction of prophages in human gut-associated Bifidobacterium hosts. Sci Rep 8(1):12772. https://doi.org/10.1038/s41598-018-31181-3
Jans C, Lacroix C, Follador R et al (2013) Complete genome sequence of the probiotic Bifidobacterium thermophilum strain RBL67. Genome Announc 1(3):e00191-13–e00191-13. https://doi.org/10.1128/genomea.00191-13
Groth AC, Calos MP (2004) Phage integrases: biology and applications. J Mol Biol 335(3):667–678
Koberg S, Mohamed MDA, Faulhaber K et al (2015) Identification and characterization of cis- and trans-acting elements involved in prophage induction in Streptococcus thermophilus J34. Mol Microbiol 98(3):535–552. https://doi.org/10.1111/mmi.13140
Ptashne M (2004) A genetic switch: phage lambda revisited, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25(5):955–964
Kala S, Cumby N, Sadowski PD et al (2014) HNH proteins are a widespread component of phage DNA packaging machines. Proc Natl Acad Sci USA 111(16):6022–6027. https://doi.org/10.1073/pnas.1320952111
Merrill BD, Ward AT, Grose JH et al (2016) Software-based analysis of bacteriophage genomes, physical ends, and packaging strategies. BMC Genomics 17:679. https://doi.org/10.1186/s12864-016-3018-2
Acknowledgements
We kindly thank Angela Back, Inka Lammertz and Gesa Gehrke for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None of the authors has any conflict of interest to declare.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Handling Editor: T. K. Frey.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sprotte, S., Bockelmann, W., Brinks, E. et al. Genome analysis of the temperate bacteriophage PMBT6 residing in the genome of Bifidobacterium thermophilum MBT94004. Arch Virol 165, 233–236 (2020). https://doi.org/10.1007/s00705-019-04448-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-019-04448-x