Abstract
Ultraviolet (UV) light irradiation has serious consequences for cell survival, including DNA damage by formation of cyclobutane pyrimidine dimers (CPD) and pyrimidine (6,4) pyrimidone photoproducts. In general, the Nucleotide Excision Repair pathway repairs these lesions; however, all living forms, except placental mammals and some marsupials, produce a flavoprotein known as photolyase that directly reverses these lesions. The aim of this work was the isolation and identification of Antarctic UVC-resistant bacteria, and the search for novel photolyases. Two Antarctic water samples were UVC-irradiated (254 nm; 50–200 J m− 2) and 12 UVC-resistant bacteria were isolated and identified by 16S rDNA amplification/analysis as members of the genera Pseudomonas, Janthinobacterium, Flavobacterium, Hymenobacter and Sphingomonas. The UVC 50% lethal dose and the photo-repair ability of isolates were analyzed. The occurrence of photolyase coding sequences in Pseudomonas, Hymenobacter and Sphingomonas isolates were searched by PCR or by searching in the draft DNA genome. Results suggest that Pseudomonas and Hymenobacter isolates produce CDP-photolyases, and Sphingomonas produces two CPD-photolyases and a 6,4-photolyase. Results suggest that the Antarctic environment is an important source of genetic material for the identification of novel photolyase genes with potential biotechnological applications.


Similar content being viewed by others
References
Albarracin VH, Pathak GP, Douki T, Cadet J, Borsarelli CD, Gärtner W (2012) Extremophilic Acinetobacter strains from High-Altitude lakes in Argentinean puna: remarkable UV-B resistance and efficient DNA damage repair. Life Evol Biosph 42: 201. doi:10.1007/s11084-012-9276-3
Albarracin VH, Simon J, Pathak GP, Valle L, Douki T, Cadet J, Borsarelli CD, Farias ME, Gärtner W (2014) First characterisation of a CPD-class I photolyase from a UV-resistant extremophile isolated from High-Altitude Andean Lakes. Photochem Photobiol Sci 13:739–750. doi:10.1039/C3PP50399B
Albarracin VH, Gärtner W, Farias ME (2016) Forged under the sun: life and art of extremophiles from Andean lakes. Photochem Photobiol 92:14–28. doi:10.1111/php.12555
Aziz RK, Bartels D, Best AA et al (2008) The RAST server: rapid annotations using subsystems technology. BMC Genom 9:75. doi:10.1186/1471-2164-9-75
Benjdia A (2012) DNA photolyases and SP lyase: structure and mechanism of light-dependent and independent DNA lyases. Curr Opin Struct Biol 22:711–720 doi:10.1016/j.sbi.2012.10.002
Berardesca E, Bertona M, Altabas K, Altabas V, Emanuele E (2012) Reduced ultraviolet-induced DNA damage and apoptosis in human skin with topical application of a photolyase-containing DNA repair enzyme cream: clues to skin cancer prevention. Mol Med Rep 5: 570–574 doi:10.3892/mmr.2011.673
Brash DE, Franklin WA, Sancar GB, Sancar A, Haseltine WA (1985) Escherichia coli DNA photolyase reverses cyclobutane pyrimidine dimers but not pyrimidine-pyrimidone (6–4) photoproducts. J Biol Chem 260:11438–11441
Brettel K, Byrdin M (2010) Reaction mechanisms of DNA photolyase. Curr Opin Struct Biol 20:693–701 doi:10.1016/j.sbi.2010.07.003
Budden T, Bowden NA (2013) The role of altered nucleotide excision repair and UVB-induced DNA damage in melanomagenesis. Int J Mol Sci 14:1132–1151. doi:10.3390/ijms14011132
Dai J, Wang Y, Zhang L, Tang Y, Luo X, An H, Fang C (2009) Hymenobacter tibetensis sp. nov., a UV-resistant bacterium isolated from Qinghai–Tibet plateau. Syst Appl Microbiol 32:543–548. doi:10.1016/j.syapm.2009.09.001
Dillon JG, Castenholz RW (1999) Scytonemin, a cyanobacterial sheath pigment, protects against UVC radiation: implications for early photosynthetic life. J Phycol 35:673–681. doi:10.1046/j.1529-8817.1999.3540673.x
Garinis GA, Mitchell JR, Moorhouse MJ et al (2005) Transcriptome analysis reveals cyclobutane pyrimidine dimers as a major source of UV-induced DNA breaks. EMBO J 24:3952–3962. doi:10.1038/sj.emboj.7600849
Goosen N, Moolenaar GF (2008) Repair of UV damage in bacteria. DNA Repair 7: 353–379 doi:10.1016/j.dnarep.2007.09.002
Graf D, Wesslowski J, Ma H, Scheerer P, Krauß N, Oberpichler I, Lamparter T (2015) Key Amino Acids in the Bacterial (6–4) Photolyase PhrB from Agrobacterium fabrum. PloS One 10:e0140955. doi:10.1371/journal.pone.0140955
Kato R, Hasegawa K, Hidaka Y, Kuramitsu S, Hoshino T (1997) Characterization of a thermostable DNA photolyase from an extremely thermophilic bacterium, Thermus thermophilus HB27. J Bacteriol 179:6499–6503. doi:10.1128/jb.179.20.6499-6503.1997
Klassen JL, Foght JM (2008) Differences in carotenoid composition among Hymenobacter and related strains support a tree-like model of carotenoid evolution. Appl Environ Microbiol 74:2016–2022. doi:10.1128/AEM.02306-07
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. doi:10.1093/molbev/msw054
Lee J-J, Joe ES, Kim EB, Jeon SH, Srinivasan S, Jung H-Y, Kim MM (2016) Hymenobacter rubidus sp. nov., bacterium isolated from a soil. Antonie Van Leeuwenhoek 109:457–466. doi:10.1007/s10482-016-0652-2
Lehtola MJ, Miettinen IT, Vartiainen T, Rantakokko P, Hirvonen A, Martikainen PJ (2003) Impact of UV disinfection on microbially available phosphorus, organic carbon, and microbial growth in drinking water. Water Res 37:1064–1070. doi:10.1016/S0043-1354(02)00462-1
Mageswari A, Subramanian P, Srinivasan R, Karthikeyan S, Gothandam KM (2015) Astaxanthin from psychrotrophic Sphingomonas faeni exhibits antagonism against food-spoilage bacteria at low temperatures. Microbiol Res 179:38–44. doi:10.1016/j.micres.2015.06.010
Martínez-Rosales C, Castro-Sowinski S (2011) Antarctic bacterial isolates that produce cold-active extracellular proteases at low temperature but are active and stable at high temperature. Polar Res 30:7123. doi:10.3402/polar.v30i0.7123
Miller JH (1972) Experiments in Molecular Genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor
Morel MA, Braña V, Martinez-Rosales C, Cagide C, Castro-Sowinski S (2015) Five-year bio-monitoring of aquatic ecosystems near Artigas Antarctic Scientific Base, King George Island. Adv Polar Sci 26: 102–106 doi:10.13679/j.advps.2015.1.00102
Morel MA, Iriarte A, Jara E, Musto H, Castro-Sowinski S (2016) Revealing the biotechnological potential of Delftia sp. JD2 by a genomic approach. AIMS Bioeng, 3: 156–175 doi:10.3934/bioeng.2016.2.156
Narayanan DL, Saladi RN, Fox JL (2010) Review: ultraviolet radiation and skin cancer. Int J Dermat 49:978–986. doi:10.1111/j.1365-4632.2010.04474.x
Oberpichler I, Pierik AJ, Wesslowski J, Pokorny R, Rosen R, Vugman M, Zhang F, Neubauer O, Ron EZ, Batschauer A, Lamperter T (2011) A photolyase-like protein from Agrobacterium tumefaciens with an iron-sulfur cluster. PLoS One 6:e26775. doi:10.1371/journal.pone.0026775
Ordoñez OF, Flores MR, Dib JR, Paz A, Farías ME (2009) Extremophile culture collection from Andean Lakes: extreme pristine environments that host a wide diversity of microorganisms with tolerance to UV radiation. Microb Ecol 58:461–473. doi:10.1007/s00248-009-9527-7
Puig-Butillé JA, Malvehy J, Potrony M, Trullas C, Garcia-García F, Dopazo J, Puig S (2013) Role of CPI-17 in restoring skin homoeostasis in cutaneous field of cancerization: effects of topical application of a film-forming medical device containing photolyase and UV filters. Exp Dermatol 22:482–501. doi:10.1111/exd.12177
Sancar A, Smith FW, Sancar GB (1984) Purification of Escherichia coli DNA photolyase. J Biol Chem 259:6028–6032
Santos AL, Lopes S, Baptista I, Henriques I, Gornes NCM, Almeida A, Correia A, Cunha A (2011) Diversity of UV sensitivity and recovery potential among bacterioneuston and bacterioplankton isolates. Lett Appl Microbiol 52:360–366. doi:10.1111/j.1472-765X.2011.03011.x
Santos AL, Oliveira V, Baptista I, Henriques I, Gomes NCM, Almeida A, Correia A, Cunha A (2013) Wavelength dependence of biological damage induced by UV radiation on bacteria. Arch Microbiol 195:63–74. doi:10.1007/s00203-012-0847-5
Scheerer P, Zhang F, Kalms J, von Stetten D, Krauβ N, Oberpichler I, Lamparter T (2015) The Class III cyclobutane pyrimidine dimer photolyase structure reveals a new antenna chromophore binding site and alternative photoreduction pathways. J Biol Chem 290:11504–11514. doi:10.1074/jbc.M115.637868
Stege H, Roza L, Vink AA, Grewe M, Ruzicka T, Grether-Beck S, Krutmann J (2000) Enzyme plus light therapy to repair DNA damage in ultraviolet-B-irradiated human skin. PNAS 97:1790–1795. doi:10.1073/pnas.030528897
Su S, Chen M, Teng C, Jiang S, Zhang C, Lin M, Zhang W (2014) Hymenobacter kanuolensis sp. Nov., a novel radiation-resistant bacterium. Int J Sys Evol Microbiol 64:2108–2112. doi:10.1099/ijs.0.051680-0
Wang J, Du X, Pan W, Wang X, Wu W (2015) Photoactivation of the cryptochrome/photolyase superfamily. J Photochem Photobiol C Photochem Rev 22: 84–102 doi:10.1016/j.jphotochemrev.2014.12.001
Yang W (2011) Surviving the sun: Repair and bypass of DNA UV lesions. Protein Sci 20:1781–1789 doi:10.1002/pro.723
Yasui A, Takao M, Oikawa A, Kiener A, Walsh CT, Eler APM (1988) Cloning and characterization of a photolyase gene from the cyanobacterium Anacystis nidulans. Nucl Acids Res 16:4447–4463. doi:10.1093/nar/16.10.4447
Yasui A, Eker AP, Yasuhira S, Yajima H, Kobayashi, T, Takao M, Oikawa A (1994) A new class of DNA photolyases present in various organisms including aplacental mammals. The EMBO J 13:6143–6151
Zhang F, Scheerer P, Oberpichler I, Lamparter T, Krauβ N (2013) Crystal structure of a prokaryotic (6,4) photolyase with an Fe-S cluster and a 6,7-dimethyl-8-ribityllumazine antenna chromophore. Proc Nat Acad Sci 110: 7217–7222 doi:10.1073/pnas.1302377110
Acknowledgements
This work was partially supported by PEDECIBA (Programa de Desarrollo de las Ciencias Básicas) and Celsius Laboratory (http://www.celsius.uy/). The work of JJM was supported by the National Agency of Investigation and Innovation (ANII, Agencia Nacional de Investigación e Innovación). The authors thank the Uruguayan Antarctic Institute for the logistic support during the stay in the Artigas Base. S. Castro-Sowinski, M. A. Morel and W. Martínez-López are members of the National Research System (SNI, Sistema Nacional de Investigadores, of ANII).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. da Costa.
Electronic supplementary material
Below is the link to the electronic supplementary material.
792_2016_914_MOESM1_ESM.pptx
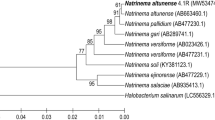
Supplementary material 1 Fig. S1 Phylogenetic analyses—Multiple alignments of 16S rRNA gene sequences, including sequences of reference strains, were performed using ClustalW. Gaps and missing data were eliminated and the molecular evolutionary analyses were conducted using MEGA version 7 (Kumar et al. 2016). The evolutionary history was inferred using the neighbor-joining (NJ) and maximum parsimony methods (bootstrap analyses of 500 replicates). Branches with less than 50 % bootstrap replicates were collapsed. The trees showed similar topology; thus, only the NJ-based tree is shown (PPTX 54 KB)
Rights and permissions
About this article
Cite this article
Marizcurrena, J.J., Morel, M.A., Braña, V. et al. Searching for novel photolyases in UVC-resistant Antarctic bacteria. Extremophiles 21, 409–418 (2017). https://doi.org/10.1007/s00792-016-0914-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-016-0914-y