Abstract
Karyotype evolution in species with non-localised centromeres (holocentric chromosomes) is usually very dynamic and associated with recurrent fission and fusion (also termed agmatoploidy/symploidy) events. In Rhynchospora (Cyperaceae), one of the most species-rich sedge genera, all analysed species have holocentric chromosomes and their numbers range from 2n = 4 to 2n = 84. Agmatoploidy/symploidy and polyploidy were suggested as the main processes in the reshuffling of Rhynchospora karyotypes, although testing different scenarios of chromosome number evolution in a phylogenetic framework has not been attempted until now. Here, we used maximum likelihood and model-based analyses, in combination with genome size estimation and ribosomal DNA distribution, to understand chromosome evolution in Rhynchospora. Overall, chromosome number variation showed a significant phylogenetic signal and the majority of the lineages maintained a karyotype of 2n = 10 (~48% of the species), the most likely candidate for the ancestral number of the genus. Higher and lower chromosome numbers were restricted to specific clades, whilst polyploidy and/or fusion/fission events were present in specific branches. Variation in genome size and ribosomal DNA site number showed no correlation with ploidy level or chromosome number. Although different mechanisms of karyotype evolution (polyploidy, fusion and fission) seem to be acting in distinct lineages, the degree of chromosome variation and the main mechanisms involved are comparable to those found in some monocentric genera and lower than expected for a holocentric genus.


Similar content being viewed by others
References
Ahola V, Lehtonen R, Somervuo P, Salmela L, Koshinen P, Rastas P et al (2014) The Glanville fritillary genome retains an ancient karyotype and reveals selective chromosomal fusions in Lepidoptera. Nat Commun 5:4337. doi:https://doi.org/10.1038/ncomms5737
Arguelho EG, Michelan VS, Nogueira FM, Da Silva CRM, Rodriguez C, Trevisan R, Vanzela ALL (2012) New chromosome counts in Brazilian species of Rhynchospora (Cyperaceae). Caryologia 65:140–146. doi:https://doi.org/10.1080/00087114.2012.711675
Bennett MD, Leitch IJ (2005) Genome size evolution in plants. In: Gregory TR (ed) The evolution of the genome, 1st edn. Elsevier Academic Press, The Netherlands, pp 89-162
Božek M, Leitch AR, Leitch IJ, Drábková LZ, Kuta E (2012) Chromosome and genome size variation in Luzula (Juncaceae), a genus with holocentric chromosomes. Bot J Linn Soc 170:529–541. doi:https://doi.org/10.1111/j.1095-8339.2012.01314.x
Buddenhagen CE (2016) A view of Rhynchosporeae (Cyperaceae) diversification before and after the application of anchored phylogenomics across the angiosperms. Dissertation, Florida State University
Buddenhagen CE, Thomas WW, Mast AR (2017) A First Look at Diversification of Beaksedges (Tribe Rhynchosporeae; Cyperaceae) in Habitat, Pollination, and Photosynthetic Features. Mem N Y Bot Gard 118:111–124. doi:https://doi.org/10.21135/893275341.002
Cabral G, Marques A, Schubert V, Pedrosa-Harand A, Schlögelhofer P (2014) Chiasmatic and achiasmatic inverted meiosis of plants with holocentric chromosomes. Nat Commun 5:5070. doi:https://doi.org/10.1038/ncomms6070
Carvalho CR, Saraiva LS (1993) An air dry technique for maize chromosomes without enzymatic maceration. Biotech Histochem 68:142–145
da Silva CRM, González-Elizondo MS, Vanzela ALL (2005) Reduction of chromosome number in Eleocharis subarticulata (Cyperaceae) by multiple translocations. Bot J Linn Soc 149:457–464
da Silva CRM, González-Elizondo MS, Vanzela ALL (2008) Chromosome reduction in Eleocharis maculosa (Cyperaceae). Cytogenet Genome Res 122:175–180
Doležel J, Greilhuber J, Suda J (2007) Estimation of nuclear DNA content in plants using flow cytometry. Nat Protoc 2:2233–2244
Drábková LZ (2013) A survey of karyological phenomena in the Juncaceae with emphasis on chromosome number variation and evolution. Bot Rev 79:401–446. doi:https://doi.org/10.1007/s12229-013-9127-6
Escudero M, Hipp AL, Luceño M (2010) Karyotype stability and predictors of chromosome number variation in sedges: a study in Carex section Spirostachyae (Cyperaceae). Mol Phylogenet Evol 57:353–363. doi:https://doi.org/10.1016/j.ympev.2010.07.009
Escudero M, Hipp AL, Hansen TF, Voje KL, Luceño M (2012) Selection and inertia in the evolution of holocentric chromosomes in sedges (Carex, Cyperaceae). New Phytol 195:237–247. doi:https://doi.org/10.1111/j.1469-8137.2012.04137.x
Escudero M, Martín-Bravo S, Mayrose I, Fernández-Mazuecos M, Fiz-Palacios O, Hipp AL, Pimentel M, Jiménez-Mejías P, Valcárcel V, Vargas P, Luceño M (2014) Karyotypic changes through dysploidy persist longer over evolutionary time than polyploid changes. PLoS One 9:e85266. doi:https://doi.org/10.1371/journal.pone.0085266
Fonsêca A, Ferreira J, Santos TRB, Mosiolek M, Belluci E, Kami J, Gepts P, Geffroy V, Schweizer D, Santos KGB, Pedrosa-Harand A (2010) Cytogenetic map of common bean (Phaseolus vulgaris L.) Chrom Res 18:487–502. doi:https://doi.org/10.1007/s10577-010-9129-8
Gerlach WL, Bedbrook JR (1979) Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 7:1869–1885
Goldblatt P, Johnson DE (1979) Index to plant chromosome numbers. Missouri Botanical Garden, St. Louis. http://mobot.mobot.org/W3T/Search/ipcn.html. Accessed 22 July 2016
Guerra M (2016) Agmatoploidy and symploidy: a critical review. Genet Mol Biol 39:492–496. doi:https://doi.org/10.1590/1678-4685-GMB-2016-0103
Heckmann S, Houben A (2013) Holokinetic centromeres. In: Jiang J, Bichler JA (eds) Plant centromere Biology, 1st edn. John Willey and Sons, New York, pp 83–94
Hipp AL (2007) Nonuniform processes of chromosome evolution in sedges (Carex: Cyperaceae). Evolution 61:2175–2194
Hipp AL, Rothrock PE, Roalson EH (2009) The evolution of chromosome arrangements in Carex (Cyperaceae). Bot Rev 75:96–109. doi:https://doi.org/10.1007/s12229-008-9022-8
Jankowska M, Fuchs J, Klocke E, Fojtová M, Polanská P, Fajkus J, Schubert V, Houben A (2015) Holokinetic centromeres and efficient telomere healing enable rapid karyotype evolution. Chromosoma 124:519–528. doi:https://doi.org/10.1007/s00412-015-0524-y
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Mentjies P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. doi:https://doi.org/10.1093/bioinformatics/bts199
Kolano B, McCann J, Orzechowska M, Siwinska D, Temsch E, Weiss-Schneeweiss H (2016) Molecular and cytogenetic evidence for an allotetraploid origin of Chenopodium quinoa and C. berlandieri (Amaranthaceae). Mol Phylogenet Evol 100:109–123. doi:https://doi.org/10.1016/j.ympev.2016.04.009
Lipnerová I, Bureš P, Horová L, Šmarda P (2012) Evolution of genome size in Carex (Cyperaceae) in relation to chromosome number and genomic base composition. Ann Bot 111:79–94. doi:https://doi.org/10.1093/aob/mcs239
Loureiro J, Rodriguez E, Dolezel J, Santos C (2007) Two new nuclear isolation buffers for plant DNA flow cytometry: a test with 37 species. Ann Bot 100:875–888. doi:https://doi.org/10.1093/aob/mcm152
Luceño M, Martín J (1986) Cyperaceae Vahl. In: Castroviejo S, Luceño M, Galán A, Jiménez Mejías P, Cabezas F, Medina L (eds) Flora Iberica. Plantas vasculares de la Península Ibérica e Islas Baleares, vol 18. Real Jardín Botánico, Madrid, pp 99–102
Luceño M, Mendes AP, Vanzela ALL, Alves MV (1998a) Agmatoploidy and symploidy in Rhynchospora cephalotes (L.) Vahl (Cyperaceae). Cytologia 63:79–81
Luceño M, Vanzela ALL, Guerra M (1998b) Cytotaxomonic studies in Brazilian Rhynchospora (Cyperaceae), a genus exhibiting holocentric chromosomes. Can J Bot 76:440–449
Maddison WP, Maddison DR (2011) Mesquite: a modular system for evolutionary analysis. Version 2.75 http://mesquiteproject.org
Marques A, Ribeiro T, Neumann P, Macas J, Novák P, Schubert V, Pellino M, Fuchs J, Ma W, Kuhlmann M, Brandt R, Vanzela ALL, Beseda T, Šimková H, Pedrosa-Harand A, Houben A (2015) Holocentromeres in are associated with genome-wide centromere-specific repeat arrays interspersed among euchromatin. PNAS 112:13633–13638. doi:https://doi.org/10.1073/pnas.1512255112
Mayrose I, Barker MS, Otto SP (2010) Probabilistic models of chromosome number evolution and the inference of polyploidy. Syst Biol 59:132–144. doi:https://doi.org/10.1093/sysbio/syp083
Melters DP, Paliulis LV, Korf IF, Chan SWL (2012) Holocentric chromosomes: convergent evolution, meiotic adaptations and genomic analysis. Chrom Res 20:579–593. doi:https://doi.org/10.1007/s10577-012-9292-1
Pagel M (1999) Inferring the historical patterns of biological evolution. Nature 401:877–884
Pellicer J, Kelly LJ, Magdalena C, Leitch IJ (2013) Insights into the dynamics of genome size and chromosome evolution in the early diverging angiosperm lineage Nymphaeales (water lilies). Genome 56:437–449. doi:https://doi.org/10.1139/gen-2013-0039
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256. doi:https://doi.org/10.1093/molbev/msn083
R Core Team (2015) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/
Revell LJ (2012) Phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol Evol 3:217–223. doi:https://doi.org/10.1111/j.2041-210X.2011.00169.x
Ribeiro T, Marques A, Novák P, Schubert V, Vanzela ALL, Macas J, Houben A, Pedrosa-Harand A (2016) Centromeric and non-centromeric satellite DNA organisation differs in holocentric Rhynchospora species. Chromosoma 126:325–335. doi:https://doi.org/10.1007/s00412-016-0616-3
Roa F, Guerra M (2012) Distribution of 45S rDNA sites in chromosomes of plants: structural and evolutionary implications. BMC Evol Biol 12:225. doi:https://doi.org/10.1186/1471-2148-12-225
Roalson EH (2008) A synopsis of chromosome number variation in the Cyperaceae. Bot Rev 74:209–393. doi:https://doi.org/10.1007/s12229-008-9011-y
Roalson EH, McCubbin AG, Whitkus R (2007) Chromosome evolution in Cyperales. Aliso 23:62–71. doi:https://doi.org/10.5642/aliso.20072301.08
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Ruban A, Fuchs J, Marques A, Schubert V, Soloviev A, Raskina O, Badaeva E, Houben A (2014) B chromosomes of Aegilops speltoides are enriched in organelle genome-derived sequences. PLoS One 9:2. doi:https://doi.org/10.1371/journal.pone.0090214
Schubert I (2007) Chromosome evolution. Curr Opin Plant Biol 10:109–115
Schubert I, Lysak MA (2011) Interpretation of karyotype evolution should consider chromosome structural constraints. TIG 27:207–216. doi:https://doi.org/10.1016/j.tig.2011.03.004
Sousa A, Barros e Silva AE, Cuadrado A, Loarce Y, Alves MV, Guerra M (2011) Distribution of 5S and 45S rDNA sites in plants with holokinetic chromosomes and the “chromosome field” hypothesis. Micron 42:625–631. doi:https://doi.org/10.1016/j.micron.2011.03.002
Sousa A, Cusimano N, Renner S (2014) Combining FISH and model-based predictions to understand chromosome evolution in Typhonium (Araceae). Ann Bot 113:669–680. doi:https://doi.org/10.1093/aob/mct302
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Vanzela ALL, Colaço W (2002) Mitotic and meiotic behavior of γ irradiated holocentric chromosomes of Rhynchospora pubera (Cyperaceae). Acta Sci 24:611–614
Vanzela ALL, Guerra M, Luceño M (1996) Rhynchospora tenuis Link (Cyperaceae), a species with the lowest number of holocentric chromosomes. Cytobios 88:219–228
Vanzela ALL, Cuadrado A, Jouve L, Luceño M, Guerra M (1998) Multiple locations of the rDNA sites in holocentric chromosomes of Rhynchospora (Cyperaceae). Chrom Res 6:345–350
Vanzela ALL, Luceño M, Guerra M (2000) Karyotype evolution and cytotaxonomy in Brazilian species of Rhynchospora Vahl (Cyperaceae). Bot J Linn Soc 134:557–566
Vanzela ALL, Cuadrado A, Guerra M (2003) Localization of 45S rDNA and telomeric sites on holocentric chromosomes of Rhynchospora tenuis Link (Cyperaceae). Genet Mol Biol 26:199–201
Yano O, Hoshino T (2006) Phylogenetic relationships and chromosomal evolution of Japanese Fimbristylis (Cyperaceae) using nrDNA ITS and ETS 1f sequence data. Acta Phytotax Geobot 57:205–217
Yoshido A, Yasukochi Y, Sahara K (2011) Samia Cynthia versus Bombyx mori: comparative gene mapping between a species with a low-number karyotype and the model species of Lepidoptera. Insect Biochem Molec Biol 41:370–377. doi:https://doi.org/10.1016/j.ibmb.2011.02.005
Zedek F, Šmerda J, Šmarda P, Bureš P (2010) Correlated evolution of LTR retrotransposons and genome size in the genus Eleocharis. BMC Plant Biol 10:265
Acknowledgements
We thank the Brazilian agencies Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) for the scholarship to T. Ribeiro, Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for the fellowship to A. Pedrosa-Harand and Fundação de Amparo à Ciência e Tecnologia do Estado de Pernambuco (FACEPE) for the financial support (APQ-2008-2.02/12). We are also grateful to Aretuza Sousa (University of Munich, Germany) for the help with the plotting of ChromEvol output in R and André Vanzela (State University of Londrina, Paraná, Brazil) and Ana Carolina Galindo da Costa (Federal University of Pernambuco, Brazil) for collecting some species in the field.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Heiti Paves
Electronic supplementary material
Online Resource 1
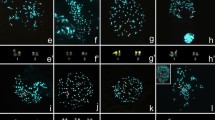
Mitotic metaphases of R. globosa (2n = 50), R. elatior (2n = 12) and R. radicans (2n = 10) stained with DAPI. Bar = 5 μm. (GIF 190 kb)
Online Resource 2
Ancestral character estimation of chromosome number (log-transformed maximum chromosome number count) along the branches and nodes of the phylogeny of the Rhynchospora (Cyperaceae). The colour of edges in the tree represents observed and reconstructed values for chromosome number on the tree. Red colours correspond with relatively low chromosome number, whereas dark blue colours represent larger observed and reconstructed chromosome sets. (PDF 1824 kb)
Online Resource 3
Ancestral state reconstruction of chromosome number (estimated in Mesquite) for the genus Rhynchospora. Haploid chromosome numbers are coloured according to legend. (PDF 463 kb)
Rights and permissions
About this article
Cite this article
Ribeiro, T., Buddenhagen, C., Thomas, W. et al. Are holocentrics doomed to change? Limited chromosome number variation in Rhynchospora Vahl (Cyperaceae). Protoplasma 255, 263–272 (2018). https://doi.org/10.1007/s00709-017-1154-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-017-1154-4