Abstract
Purpose
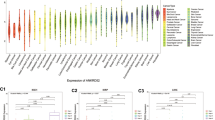
KLHDC7B is a member of Kelch family, with a Kelch domain in the C-terminal half, which plays a role in various cellular events, such as cytoskeletal arrangement, protein degradation, gene expression. Although there is increasing evidence supporting KLHDC7B's vital role in tumorigenesis, a systematic analysis of KLHDC7B in cancers remains lacking. Therefore, we intended to investigate the prognostic value for KLHDC7B across 33 cancer types and explore its potential immunological function.
Methods
GEO (Gene Expression Omnibus database) and TCGA (The Cancer Genome Atla) database were used to explore the role of KLHDC7B in 33 cancers. TIMER2, GEPIA2 and Kaplan–Meier plotter were utilized to explore the KLHDC7B expression level and prognostic value in different cancers. The pan cancer genetic variation and DNA methylation of KLHDC7B were analyzed by cBioPortal and MEXPRESS. TIMER2 was employed to investigate the correlation between KLHDC7B expression and immune infiltration. The relationship of KLHDC7B expression with TMB (tumor mutational burden) and MSI (microsatellite instability) were evaluated using Spearman correlation analysis. Finally, by GO and KEGG enrichment analysis, the underlying mechanisms of KLHDC7B in tumor pathophysiology were further investigated.
Results
KLHDC7B expression level was related to pathological stages, MSI, TMB, immune checkpoint and immune cell infiltration in most cancers. Especially, we found that the KLHDC7B expression was negatively correlated with the immune infiltration of Myeloid derived suppressor cells into TGCT and GBM. Additionally, survival analysis showed that the expression of KLHDC7B was connected with overall survival (OS) in 3 cancers and disease-free survival (DFS) in 5 cancers. Furthermore, the enrichment analysis revealed that the KLHDC7B collecting genes and binding proteins are related to the function of proteins and immune response.
Conclusion
KLHDC7B demonstrates strong clinical utility as markers of prognostic and immune response in pan-cancer.









Similar content being viewed by others
Data Availability
The authors certify that all the original data in this research could be obtained from the public databases.
References
Andre F, Mardis E, Salm M et al (2014) Prioritizing targets for precision cancer medicine. Ann Oncol 25:2295–2303. https://doi.org/10.1093/annonc/mdu478
Bardou P, Mariette J, Escudié F et al (2014) jvenn: an interactive Venn diagram viewer. BMC Bioinformatics 15:293. https://doi.org/10.1186/1471-2105-15-293
Baretti M, Le DT (2018) DNA mismatch repair in cancer. Pharmacol Ther 189:45–62. https://doi.org/10.1016/j.pharmthera.2018.04.004
Bonneville R, Krook MA, Kautto EA et al (2017) Landscape of microsatellite instability across 39 cancer types. JCO Precis Oncol. https://doi.org/10.1200/po.17.00073
Chan TA, Yarchoan M, Jaffee E et al (2019) Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol 30:44–56. https://doi.org/10.1093/annonc/mdy495
Chen X, Song E (2019) Turning foes to friends: targeting cancer-associated fibroblasts. Nat Rev Drug Discov 18:99–115. https://doi.org/10.1038/s41573-018-0004-1
Cui G, Wang C, Lin Z et al (2021) Prognostic and immunological role of Ras-related protein Rap1b in pan-cancer. Bioengineered 12:4828–4840. https://doi.org/10.1080/21655979.2021.1955559
Dan H, Zhang S, Zhou Y et al (2019) DNA methyltransferase inhibitors: catalysts for antitumour immune responses. Onco Targets Ther 12:10903–10916. https://doi.org/10.2147/ott.S217767
Dudley JC, Lin MT, Le DT et al (2016) Microsatellite instability as a biomarker for PD-1 blockade. Clin Cancer Res 22:813–820. https://doi.org/10.1158/1078-0432.Ccr-15-1678
Fishel R (2015) Mismatch repair. J Biol Chem 290:26395–26403. https://doi.org/10.1074/jbc.R115.660142
Fridman WH, Galon J, Dieu-Nosjean MC et al (2011) Immune infiltration in human cancer: prognostic significance and disease control. Curr Top Microbiol Immunol 344:1–24. https://doi.org/10.1007/82_2010_46
Gao J, Aksoy BA, Dogrusoz U et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6:pl1. https://doi.org/10.1126/scisignal.2004088
Guo P, Wang D, Wu J et al (2015) The landscape of alternative splicing in cervical squamous cell carcinoma. Onco Targets Ther 8:73–79. https://doi.org/10.2147/ott.S72832
Gupta VA, Beggs AH (2014) Kelch proteins: emerging roles in skeletal muscle development and diseases. Skelet Muscle 4:11. https://doi.org/10.1186/2044-5040-4-11
Györffy B (2021) Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput Struct Biotechnol J 19:4101–4109. https://doi.org/10.1016/j.csbj.2021.07.014
Györffy B, Lanczky A, Eklund AC et al (2010) An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat 123:725–731. https://doi.org/10.1007/s10549-009-0674-9
Hause RJ, Pritchard CC, Shendure J et al (2016) Classification and characterization of microsatellite instability across 18 cancer types. Nat Med 22:1342–1350. https://doi.org/10.1038/nm.4191
Houthuijzen JM, Jonkers J (2018) Cancer-associated fibroblasts as key regulators of the breast cancer tumor microenvironment. Cancer Metast Rev 37:577–597. https://doi.org/10.1007/s10555-018-9768-3
Huang H, Du J, Jin B et al (2021) Combination of urine exosomal mRNAs and lncRNAs as novel diagnostic biomarkers for bladder cancer. Front Oncol 11:667212. https://doi.org/10.3389/fonc.2021.667212
Jeong G, Bae H, Jeong D et al (2018) A Kelch domain-containing KLHDC7B and a long non-coding RNA ST8SIA6-AS1 act oppositely on breast cancer cell proliferation via the interferon signaling pathway. Sci Rep 8:12922. https://doi.org/10.1038/s41598-018-31306-8
Kent WJ, Sugnet CW, Furey TS et al (2002) The human genome browser at UCSC. Genome Res 12:996–1006. https://doi.org/10.1101/gr.229102
Khirade MF, Lal G, Bapat SA (2015) Derivation of a fifteen gene prognostic panel for six cancers. Sci Rep 5:13248. https://doi.org/10.1038/srep13248
Kim TW, Kim YJ, Lee HJ et al (2010) Hs.137007 is a novel epigenetic marker hypermethylated and up-regulated in breast cancer. Int J Oncol 36:1105–1111. https://doi.org/10.3892/ijo_00000592
Koch A, De Meyer T, Jeschke J et al (2015) MEXPRESS: visualizing expression, DNA methylation and clinical TCGA data. BMC Genom 16:636. https://doi.org/10.1186/s12864-015-1847-z
Kwa MQ, Herum KM, Brakebusch C (2019) Cancer-associated fibroblasts: how do they contribute to metastasis? Clin Exp Metast 36:71–86. https://doi.org/10.1007/s10585-019-09959-0
Lánczky A, Győrffy B (2021) Web-Based Survival Analysis Tool Tailored for Medical Research (KMplot): Development and Implementation. J Med Internet Res 23:e27633. https://doi.org/10.2196/27633
Lei Y, Yu T, Li C et al (2021) Expression of CAMK1 and its association with clinicopathologic characteristics in pancreatic cancer. J Cell Mol Med 25:1198–1206. https://doi.org/10.1111/jcmm.16188
Lonsdale J, Thomas J, Salvatore M, Phillips R, Lo E, Shad S, Hasz R, Walters G, Garcia F, Young N, Foster B (2013) The Genotype-Tissue Expression (GTEx) project. Nat Genet 45:580–585. https://doi.org/10.1038/ng.2653
Ma X, Liu Y, Liu Y et al (2018) Pan-cancer genome and transcriptome analyses of 1,699 paediatric leukaemias and solid tumours. Nature 555:371–376. https://doi.org/10.1038/nature25795
Malouf GG, Su X, Zhang J et al (2016) DNA methylation signature reveals cell ontogeny of renal cell carcinomas. Clin Cancer Res 22:6236–6246. https://doi.org/10.1158/1078-0432.Ccr-15-1217
Paauwe M, Schoonderwoerd MJA, Helderman R et al (2018) Endoglin expression on cancer-associated fibroblasts regulates invasion and stimulates colorectal cancer metastasis. Clin Cancer Res 24:6331–6344. https://doi.org/10.1158/1078-0432.Ccr-18-0329
Papic N, Maxwell CI, Delker DA et al (2012) RNA-sequencing analysis of 5’ capped RNAs identifies many new differentially expressed genes in acute hepatitis C virus infection. Viruses 4:581–612. https://doi.org/10.3390/v4040581
Praveen K, Dobbyn L, Gurski L et al (2022) Population-scale analysis of common and rare genetic variation associated with hearing loss in adults. Commun Biol 5:540. https://doi.org/10.1038/s42003-022-03408-7
Priestley P, Baber J, Lolkema MP et al (2019) Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 575:210–216. https://doi.org/10.1038/s41586-019-1689-y
Russo M, Crisafulli G, Sogari A et al (2019) Adaptive mutability of colorectal cancers in response to targeted therapies. Science 366:1473–1480. https://doi.org/10.1126/science.aav4474
Saghafinia S, Mina M, Riggi N et al (2018) Pan-cancer landscape of aberrant DNA methylation across human tumors. Cell Rep 25:1066-1080.e1068. https://doi.org/10.1016/j.celrep.2018.09.082
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Steven A, Seliger B (2018) The Role of Immune Escape and Immune Cell Infiltration in Breast Cancer. Breast Care (basel) 13:16–21. https://doi.org/10.1159/000486585
Szklarczyk D, Franceschini A, Wyder S et al (2015) STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43:D447-452. https://doi.org/10.1093/nar/gku1003
Tang Z, Kang B, Li C et al (2019) GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res 47:W556-w560. https://doi.org/10.1093/nar/gkz430
Van Velzen MJM, Derks S, Van Grieken NCT et al (2020) MSI as a predictive factor for treatment outcome of gastroesophageal adenocarcinoma. Cancer Treat Rev 86:102024. https://doi.org/10.1016/j.ctrv.2020.102024
Yan T, Zhu S, Shi Y et al (2021) Pan-cancer analysis of atrial-fibrillation-related innate immunity gene ANXA4. Front Cardiovasc Med 8:713983. https://doi.org/10.3389/fcvm.2021.713983
Yarchoan M, Hopkins A, Jaffee EM (2017) Tumor mutational burden and response rate to PD-1 inhibition. N Engl J Med 377:2500–2501. https://doi.org/10.1056/NEJMc1713444
Yu G, Wang LG, Han Y et al (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287. https://doi.org/10.1089/omi.2011.0118
Zhang J, Wu LY, Zhang XS et al (2014) Discovery of co-occurring driver pathways in cancer. BMC Bioinformatics 15:271. https://doi.org/10.1186/1471-2105-15-271
Zhang G, Fan E, Yue G et al (2019) Five genes as a novel signature for predicting the prognosis of patients with laryngeal cancer. J Cell Biochem. https://doi.org/10.1002/jcb.29535
Acknowledgements
The authors would also like to thank Ningbo Institute for Medicine & Biomedical Engineering Combined Innovation, The First Affiliated Hospital of Nanchang University for their critical support of this study.
Funding
The work was supported by the National Natural Science Foundation of China, 81860342, 81972445, and 81772083 (GF) for funding this project.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Data collection and analysis were performed by JD, XJ, LL, D-ZC, NL, X-TY and FG. The first draft of the manuscript was written by XJ and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ding, J., Ji, X., Liu, L. et al. A prognostic and immunological analysis of 7B-containing Kelch structural domain (KLHDC7B) in pan-cancer: a potential target for immunotherapy and survival. J Cancer Res Clin Oncol 149, 7857–7876 (2023). https://doi.org/10.1007/s00432-023-04738-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04738-7