Abstract
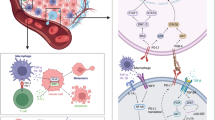
Myeloid-derived suppressor cells (MDSC) populate the peripheral blood and contribute to immune regulation in cancer. However, there is limited knowledge on the myeloid cell types with proinflammatory capacities that may serve as opponents of MDSC. In the circulation of cancer patients, a monocyte subpopulation was identified with a specific immunophenotype and transcriptomic signature. They were predominantly CD14+CD33hiCD16−/+HLA-DR+/hi cells that typically expressed CD66b. In accordance with the transcriptomics data, NALP3, LOX-1 and PAI-1 levels were also significantly upregulated. The CD66b+ monocytes displayed high phagocytic activity, matrix adhesion and migration, and provided costimulation for T cell proliferation and IFN-γ secretion; thus, they did not suppress T cell responses. Irrespective of clinical stage, they were identified in various cancers. In conclusion, the CD66b+ monocytes represent a novel myeloid subpopulation which is devoid of immune regulatory influences of cancer and displays enhanced proinflammatory capacities.




Similar content being viewed by others
References
Nagl M, Kacani L, Mullauer B et al (2002) Phagocytosis and killing of bacteria by professional phagocytes and dendritic cells. Clin Vaccine Immunol 9:1165–1168. https://doi.org/10.1128/cdli.9.6.1165-1168.2002
Jakubzick CV, Randolph GJ, Henson PM (2017) Monocyte differentiation and antigen-presenting functions. Nat Rev Immunol 17:349–362. https://doi.org/10.1038/nri.2017.28
Boyette LB, MacEdo C, Hadi K et al (2017) Phenotype, function, and differentiation potential of human monocyte subsets. PLoS ONE 12:1–20. https://doi.org/10.1371/journal.pone.0176460
Ginhoux F, Jung S (2014) Monocytes and macrophages: developmental pathways and tissue homeostasis TL—14. Nat Rev Immunol 14 VN-r:392–404. https://doi.org/10.1038/nri3671
Guilliams M, Mildner A, Yona S (2018) Review developmental and functional heterogeneity of monocytes. Immunity 49:595–613. https://doi.org/10.1016/j.immuni.2018.10.005
Patel AA, Zhang Y, Fullerton JN et al (2017) The fate and lifespan of human monocyte subsets in steady state and systemic inflammation. J Exp Med 214:1913–1923
Gordon S, Taylor PR (2005) Monocyte and macrophage heterogeneity. Nat Rev Immunol 5:953–964. https://doi.org/10.1038/nri1733
Ong SM, Hadadi E, Dang TM et al (2018) The pro-inflammatory phenotype of the human non-classical monocyte subset is attributed to senescence article. Cell Death Dis 9:1–12. https://doi.org/10.1038/s41419-018-0327-1
Wildgruber M, Aschenbrenner T, Wendorff H et al (2016) The “Intermediate” CD14++CD16+ monocyte subset increases in severe peripheral artery disease in humans. Sci Rep 6:39483. https://doi.org/10.1038/srep39483
Marimuthu R, Francis H, Dervish S et al (2018) Characterization of human monocyte subsets by whole blood flow cytometry analysis. J Vis Exp 140:1–10. https://doi.org/10.3791/57941
Bruger AM, Dorhoi A, Esendagli G et al (2019) How to measure the immunosuppressive activity of MDSC: assays, problems and potential solutions. Cancer Immunol Immunother 68:631–644. https://doi.org/10.1007/s00262-018-2170-8
Gabrilovich DI, Nagaraj S (2009) Myeloid-derived suppressor cells as regulators of the immune system. Nat Rev Immunol 9:162–174. https://doi.org/10.1038/nri2506
Gabrilovich DI (2017) Myeloid-derived suppressor cells. Cancer Immunol Res 5:3–8. https://doi.org/10.1158/2326-6066.CIR-16-0297
Onicescu G, Rosato A, Mandruzzato S et al (2011) A human promyelocytic-like population is responsible for the immune suppression mediated by myeloid-derived suppressor cells. Blood 118:2254–2265. https://doi.org/10.1182/blood-2010-12-325753
Brandau S, Trellakis S, Bruderek K et al (2011) Myeloid-derived suppressor cells in the peripheral blood of cancer patients contain a subset of immature neutrophils with impaired migratory properties. J Leukoc Biol 89:311–317. https://doi.org/10.1189/jlb.0310162
Filipazzi P, Valenti R, Huber V et al (2007) Identification of a new subset of myeloid suppressor cells in peripheral blood of melanoma patients with modulation by a granulocyte-macrophage colony-stimulation factor-based antitumor vaccine. J Clin Oncol 25:2546–2553. https://doi.org/10.1200/JCO.2006.08.5829
Kumar V, Patel S, Tcyganov E, Gabrilovich DI (2016) The nature of myeloid-derived suppressor cells in the tumor microenvironment. Trends Immunol 37:208–220. https://doi.org/10.1016/j.it.2016.01.004
Umansky V, Blattner C, Gebhardt C, Utikal J (2016) The role of myeloid-derived suppressor cells (MDSC) in cancer progression. Vaccines 4:36. https://doi.org/10.3390/vaccines4040036
Moses K, Brandau S (2016) Human neutrophils: their role in cancer and relation to myeloid-derived suppressor cells. Semin Immunol 28:187–196. https://doi.org/10.1016/j.smim.2016.03.018
Qu P, Wang L, Lin PC (2016) Expansion and functions of myeloid-derived suppressor cells in the tumor microenvironment. Cancer Lett 380:253–256. https://doi.org/10.1016/j.canlet.2015.10.022
Bronte V, Brandau S, Chen S-H et al (2016) Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat Commun 7:12150. https://doi.org/10.1038/ncomms12150
Ge SX, Son EW, Yao R (2018) iDEP: An integrated web application for differential expression and pathway analysis of RNA-Seq data. BMC Bioinform 19:1–24. https://doi.org/10.1186/s12859-018-2486-6
Szklarczyk D, Morris JH, Cook H et al (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45:D362–D368. https://doi.org/10.1093/nar/gkw937
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Ziegler-heitbrock L, Ancuta P, Crowe S et al (2010) Nomenclature of monocytes and dendritic cells in blood. Blood 116:5–7. https://doi.org/10.1182/blood-2010-02-258558
Yoon BR, Yoo S, Choi Y et al (2014) Functional phenotype of synovial monocytes modulating inflammatory T-cell responses in rheumatoid arthritis ( RA ). PLoS ONE 9:e109775. https://doi.org/10.1371/journal.pone.0109775
Aicher A, Hayden-Ledbetter M, Brady WA et al (2000) Characterization of human inducible costimulator ligand expression and function. J Immunol 164:4689–4696. https://doi.org/10.4049/jimmunol.164.9.4689
Ruth JH, Rottman JB, Kingsbury GA et al (2007) ICOS and B7 costimulatory molecule expression identifies activated cellular subsets in rheumatoid arthritis. Cytom Part A 71:317–326. https://doi.org/10.1002/cyto.a.20383
Galligan CL, Matsuyama W, Matsukawa A et al (2004) Up-regulated expression and activation of the orphan chemokine receptor, CCRL2, in rheumatoid arthritis. Arthritis Rheum 50:1806–1814. https://doi.org/10.1002/art.20275
Gren ST, Rasmussen TB, Janciauskiene S (2015) A single-cell gene-expression profile reveals inter-cellular heterogeneity within human monocyte subsets. PLoS ONE 10:1–20. https://doi.org/10.1371/journal.pone.0144351
Bucala R, Spiegel LA, Chesney J et al (1994) Circulating fibrocytes define a new leukocyte subpopulation that mediates tissue repair. Mol Med 1:71–81
Maurer D (1994) Expression of functional high affinity immunoglobulin E receptors (Fc epsilon RI) on monocytes of atopic individuals. J Exp Med 179:745–750. https://doi.org/10.1084/jem.179.2.745
Clanchy FIL (2006) Detection and properties of the human proliferative monocyte subpopulation. J Leukoc Biol 79:757–766. https://doi.org/10.1189/jlb.0905522
Komano Y, Nanki T, Hayashida K et al (2006) Identification of a human peripheral blood monocyte subset that differentiates into osteoclasts. Arthritis Res Ther 8:1–14. https://doi.org/10.1186/ar2046
Villani A-C, Satija R, Reynolds G, et al (2017) Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356: eaah4573. https://doi.org/10.1126/science.aah4573
Mildner A, Schönheit J, Giladi A et al (2017) genomic characterization of murine monocytes reveals C/EBPβ transcription factor dependence of Ly6C− cells. Immunity 46:849–862.e7. https://doi.org/10.1016/j.immuni.2017.04.018
Venneri MA, De Palma M, Ponzoni M et al (2007) Identification of proangiogenic TIE2-expressing monocytes (TEMs) in human peripheral blood and cancer. Blood 109:5276–5285. https://doi.org/10.1182/blood-2006-10-053504
Menezes S, Melandri D, Anselmi G et al (2016) The heterogeneity of Ly6Chi monocytes controls their differentiation into iNOS+ macrophages or monocyte-derived dendritic cells. Immunity 45:1205–1218. https://doi.org/10.1016/j.immuni.2016.12.001
Satoh T, Nakagawa K, Sugihara F et al (2017) Identification of an atypical monocyte and committed progenitor involved in fibrosis. Nature 541:96–101. https://doi.org/10.1038/nature20611
Yáñez A, Coetzee SG, Olsson A et al (2017) Granulocyte-monocyte progenitors and monocyte-dendritic cell progenitors independently produce functionally distinct monocytes. Immunity 47:890–902.e4. https://doi.org/10.1016/j.immuni.2017.10.021
Youn JI, Kumar V, Collazo M et al (2013) Epigenetic silencing of retinoblastoma gene regulates pathologic differentiation of myeloid cells in cancer. Nat Immunol 14:211–220. https://doi.org/10.1038/ni.2526
Mastio J, Condamine T, Dominguez G, et al (2019) Identification of monocyte-like precursors of granulocytes in cancer as a mechanism for accumulation of PMN-MDSCs. J Exp Med jem.20181952. https://doi.org/10.1084/jem.20181952
Mandruzzato S, Brandau S, Britten CM et al (2016) Toward harmonized phenotyping of human myeloid-derived suppressor cells by flow cytometry: results from an interim study. Cancer Immunol Immunother 65:161–169. https://doi.org/10.1007/s00262-015-1782-5
Wright HL, Moots RJ, Bucknall RC, Edwards SW (2010) Neutrophil function in inflammation and inflammatory diseases. Rheumatology 49:1618–1631. https://doi.org/10.1093/rheumatology/keq045
Wittmann S, Rothe G, Schmitz G, Fröhlich D (2004) Cytokine upregulation of surface antigens correlates to the priming of the neutrophil oxidative burst response. Cytom Part A 57:53–62. https://doi.org/10.1002/cyto.a.10108
Wagner C, Deppisch R, Denefleh B et al (2003) Expression patterns of the lipopolysaccharide receptor CD14, and the FCgamma receptors CD16 and CD64 on polymorphonuclear neutrophils: data from patients with severe bacterial infections and lipopolysaccharide-exposed cells. Shock 19:5–12. https://doi.org/10.1097/00024382-200301000-00002
Sharpe AH, Freeman GJ (2002) The B7-CD28 superfamily. Nat Rev Immunol 2:116–126. https://doi.org/10.1038/nri727
Chistiakov DA, Killingsworth MC, Myasoedova VA et al (2017) CD68/macrosialin: not just a histochemical marker. Lab Investig 97:4–13. https://doi.org/10.1038/labinvest.2016.116
Sabroe I, Jones EC, Usher LR et al (2002) Toll-like receptor (TLR)2 and TLR4 in human peripheral blood granulocytes: a critical role for monocytes in leukocyte lipopolysaccharide responses. J Immunol 168:4701–4710. https://doi.org/10.4049/jimmunol.168.9.4701
Sarmadi P, Tunali G, Esendagli-Yilmaz G et al (2015) CRAM-A indicates IFN-γ-associated inflammatory response in breast cancer. Mol Immunol 68:692–698. https://doi.org/10.1016/j.molimm.2015.10.019
Hayashida K, Kume N, Minami M, Kita T (2002) Lectin-like oxidized low-density lipoprotein receptor-1 (LOX-1) supports adhesion of leukocytes under both static and flow conditions. Circ J 66:432
Kedde M, Strasser MJ, Boldajipour B et al (2007) RNA-binding protein Dnd1 inhibits MicroRNA access to target mRNA. Cell 131:1273–1286. https://doi.org/10.1016/j.cell.2007.11.034
Ingersoll MA, Platt AM, Potteaux S, Randolph GJ (2011) Monocyte trafficking in acute and chronic inflammation. Trends Immunol 32:470–477. https://doi.org/10.1016/j.it.2011.05.001
Sha W, Mitoma H, Hanabuchi S et al (2014) Human NLRP3 inflammasome senses multiple types of bacterial RNAs. Proc Natl Acad Sci 111:16059–16064. https://doi.org/10.1073/pnas.1412487111
Pavón MA, Arroyo-Solera I, Téllez-Gabriel M et al (2015) Enhanced cell migration and apoptosis resistance may underlie the association between high SERPINE1 expression and poor outcome in head and neck carcinoma patients. Oncotarget 6:29016–29033. https://doi.org/10.18632/oncotarget.5032
Zhang Y, Venkatraj V, Fisher S et al (1997) Genomic cloning and chromosomal assignment of the E2F dimerization partner TFDP gene family. Genomics 39:95–98. https://doi.org/10.1006/geno.1996.4473
Bostick M, Kim JK, Esteve P-O et al (2007) UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science 317:1760–1765
Li XB, Chen J, Deng MJ et al (2011) Zinc finger protein HZF1 promotes K562 cell proliferation by interacting with and inhibiting INCA1. Mol Med Rep 4:1131–1137. https://doi.org/10.3892/mmr.2011.564
Teng T, Tsai JH, Puyang X et al (2017) Splicing modulators act at the branch point adenosine binding pocket defined by the PHF5A-SF3b complex. Nat Commun 8:1–16. https://doi.org/10.1038/ncomms15522
Trimarchi JM, Fairchild B, Verona R et al (2002) E2F–6, a member of the E2F family that can behave as a transcriptional repressor. Proc Natl Acad Sci 95:2850–2855. https://doi.org/10.1073/pnas.95.6.2850
Nonaka K, Saio M, Suwa T et al (2008) Skewing the Th cell phenotype toward Th1 alters the maturation of tumor-infiltrating mononuclear phagocytes. J Leukoc Biol 84:679–688. https://doi.org/10.1189/jlb.1107729
Acknowledgements
This work was partially supported by The Scientific and Technological Research Council of Turkey, (TÜBİTAK; Project No. 115S636) and Hacettepe University Scientific Research and Coordination Unit, (Grant No. TSA-2018-17239), and covered under the European Cooperation in Science and Technology (COST-EU) Action BM1404 (Mye-EUNITER) (https://www.myeeuniter.eu). COST is supported by the EU Framework Program Horizon 2020. We acknowledge the technical support by Beren Karaosmanoglu, PhD. The authors thank all patients and nurses who contributed to the study, especially Nuraydın Sahin for providing blood samples.
Author information
Authors and Affiliations
Contributions
G.E. conceptualized the project and designed the experiments. U.H. and D.Y.-E. performed experiments. G.E., U.H., and D.Y.-E. interpreted data. U.H. performed bioinformatics analysis of RNA-seq data and contributed to preparation of the figures. E.Z.T performed assays related to transcriptomics. K.B.Y., E.H., and D.K. assisted with histopathological and clinical evaluation of the patients. G.E. and U.H. wrote the manuscript. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Horzum, U., Yoyen-Ermis, D., Taskiran, E.Z. et al. CD66b+ monocytes represent a proinflammatory myeloid subpopulation in cancer. Cancer Immunol Immunother 70, 75–87 (2021). https://doi.org/10.1007/s00262-020-02656-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-020-02656-y