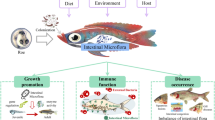
Abstract
There is a huge quantity of microorganisms in the gut of fish, which exert pivotal roles in maintaining host intestinal and general health. The fish immunity can sense and shape the intestinal microbiota and maintain the intestinal homeostasis. In the meantime, the intestinal commensal microbes regulate the fish immunity, control the extravagant proliferation of pathogenic microorganisms, and ensure the intestinal health of the host. This review summarizes developments and progress on the known interactions between host immunity and intestinal microorganisms in fish, focusing on the recent advances in zebrafish (Danio rerio) showing the host immunity senses and shapes intestinal microbiota, and intestinal microorganisms tune host immunity. This review will offer theoretical references for the development, application, and commercialization of intestinal functional microorganisms in fish.
Key points
• The interactions between the intestinal microorganisms and host immunity in zebrafish
• Fish immunity senses and shapes the microbiota
• Intestinal microbes tune host immunity in fish
Similar content being viewed by others
Introduction
In recent years, the world aquaculture is facing the challenge of increasing production, reducing emissions, and saving energy. Therefore, the aquaculture industry has gradually embarked on the road of intensive development. The problems that come with it are environmental deterioration and frequent outbreaks of diseases, causing huge economic losses every year (Assefa and Abunna 2018). In order to control the occurrence of bacterial diseases in fish, large amounts of antibiotics are used, but the effect is not satisfactory due to increasing antibiotic resistance problems (Liu et al. 2021b). At the same time, high stocking density will greatly decrease water quality, reduce function of intestinal barrier, and impair the intestinal health of aquatic animals (Sundha et al. 2019). Intestinal health has developed into one of the bottleneck constraints, restricting the healthy and sustainable development of world aquaculture. It is urgent to improve the fish intestinal health through nutritional intervention measures in intensive culture mode.
In the intestine, there exists a large number of commensal microbes (Xue et al. 2020), which provide an excellent microecological environment for fish (Liu et al. 2021a). The gut microbes play an important role in gut development, metabolism, immunity, and host health (Ou et al. 2021; Filardy et al. 2023). Recently, growing numbers of studies have begun to focus on the association of intestinal microorganisms with fish immunity and disease resistance (Xiong et al. 2019; Amenyogbe et al. 2022). Interactions between host and intestinal microbiota are the bases for the development of the immunity (Ruff et al. 2020). The host immune system coevolves with host microorganisms, which are very important to host physiology (Kogut et al. 2020), and mucosal and systemic immunity (Gutierrez et al. 2022). The host immune system monitors intestinal microorganisms, maintains the balance of intestinal microbiota and the health of intestinal mucosa, and thus strengthens the resistance to potential pathogens (Kinnebrew and Pamer 2012). The host and microorganisms interact with each other and gradually form a dynamic and stable microecological homeostasis in the gut of fish (Diwan et al. 2023). Therefore, understanding the interactions between the host immunity and intestinal microorganisms is very meaningful for maintaining the fish health.
The field on the interactions between the host immunity and microorganisms in fish is interesting and developing so quickly, and several review papers related to this topic have been published in recent years (Langlois et al. 2021; Yu et al. 2021; Diwan et al. 2023; Wang et al. 2023). In 2021, Yu et al. summarized the microbiome composition of teleost mucosa, commensal bacterial dysbiosis in teleost mucosa, crosstalk between commensals and mucosal innate and adaptive immunity, and probiotics contribute to fish immune response (Yu et al. 2021). In 2021, Langlois et al. summarized the host–microbe interactions and the functions of probiotics in salmonid, including salmonid intestinal microbiome, probiotic-mediated regulation of host health, methods of probiotic encapsulation, and delivery (Langlois et al. 2021). In 2023, Wang et al. reviewed the teleost skin microbiome-host immunity focused on the teleost external surface microbiome, including teleost skin microbiome, its modulation of host physiology and immunity, host immune modulation of skin microbiome, and models of skin microbiome-host interaction (Wang et al. 2023). In 2023, Diwan et al. summarized host-microbiome interaction in fish and shellfish, including host-microbiome interaction in fish and shellfish, manipulation of the gut microbiome, and host-microbiome interaction and aquatic environment (Diwan et al. 2023). However, there still exist huge gaps in knowledge of commensal microorganisms and their functions in fish immunity, the molecular mechanisms of the host immunity that can sense and shape intestinal microbiota. Furthermore, the zebrafish has many advantages to explore host-microbe interactions in fish, but previous reviews have lacked a systematic summary of it. Therefore, to complement previous reviews, we summarize developments and progress on the known of interactions between host immunity and intestinal microorganisms in fish, paying specific attention to recent findings in zebrafish. This review will supply theoretical support for the exploitation and application of intestinal functional microorganisms in aquaculture.
The interactions between the host immunity and intestinal microorganisms in zebrafish (Danio rerio)
Zebrafish larvae have an innate immune system, while the functional adaptive immune system is not completed until 4 weeks age (Lam et al. 2004), and the adult zebrafish possess the adaptive immune system (Stream and Madigan 2022). The body of zebrafish is transparent at the stage of embryonic development, the fish gut and morphology can be easily observed with a microscope (Teame et al. 2019). It is feasible to study the innate and adaptive immunity of zebrafish by using different developmental periods of zebrafish. Furthermore, zebrafish has several other unique advantages, the ability to image the intestinal microorganisms colonization in vivo for a long time, and the ability to present different phenotype individuals (Ganz 2018). Therefore, zebrafish have become an ideal model for revealing the interactions between the host immunity and intestinal microorganisms (Teame et al. 2019; Zhang et al. 2020b; Nadal et al. 2020; Jia et al. 2021). The summary of interactions between the host immunity and intestinal microorganisms in zebrafish is shown in Table 1.
In zebrafish, the host immunity can effectively sense intestinal microorganisms, for example, the innate immune system mediates toll-like receptor 2 (TLR2) sensing the intestinal colonization of Exiguobacterium and Chryseobacterium through negative control of myeloid differentiation factor 88 (MyD88) (Koch et al. 2018). Similarly, MyD88 mutant zebrafish are more sensitive to be infected with pathogenic Edwardsiella tarda and Mycobacterium marinum, and its pro-inflammatory cytokines interleukin-1 beta (IL-1β), activator protein-1 (AP-1), and nuclear factor kappa-B (NF-κB) are dependent on MyD88 during infections (Vaart et al. 2013). Nucleotide-binding oligomerization domain 2 (NOD2) in zebrafish embryonic fibroblasts can sense the muramyl dipeptide of Gram-negative and Gram-positive bacteria, activate NF-κB signaling, and induce antiviral defense response in spring viremia of carp virus infection (Zou et al. 2016).
The host immunity has a close relationship with the gut microbiota in zebrafish, when zebrafish are infected by Aeromonas hydrophila, the innate immune response is activated and correspondingly, the intestinal microbial structure is changed, i.e., the proportions of beneficial microorganisms such as Nitratireductor, Enterococcus, and Brevundimonas decrease, while the proportions of harmful microorganisms such as Halomonas, Pelagibacterium, and Aeromonas increase (Yang et al. 2017). When the early adaptive immunity in wild-type zebrafish has not been established, pathogenic Vibrio overgrows in the gut. For example, recombination activating gene 1 (Rag1)-deficient zebrafish lack adaptive immunity and have high abundance of Vibrio in the intestine, but transferring T lymphocytes rather than B lymphocytes to Rag1-deficient zebrafish can inhibit the growth of intestinal Vibrio (Brugman et al. 2014). The intestinal microbiota of 61 rag1− zebrafish (adaptive immunity deficiency, lacking B and T cell receptors) and 68 wild-type zebrafish (functional adaptive immunity) were surveyed to analyze the function of adaptive immunity on the intestinal microbiota. It was found that the existence of adaptive immunity makes the intestinal microbiota of the zebrafish more personalized, but this may be influenced by other conditions, including transmission of microorganisms among hosts (Stagaman et al. 2017). The intestinal microbiota is shaped by intestinal macrophages through interferon regulatory factor 8 (IRF8) in zebrafish. A reduced number of macrophages was showed with reduced expressions of complement genes c1ql, c1qa, c1qb, and c1qc, and severely dysfunctional intestinal symbiotic microbes, Fusobacteria, α-Proteobacteria, and γ-Proteobacteria, decreased, while δ-Proteobacteria increased in IRF8-deficient adult zebrafish (Earley et al. 2018).
Studies using specific microorganisms to colonize gnotobiotic zebrafish have shown that symbiotic microorganisms are important for the activation and development of immunity (Murdoch and Rawls 2019; Rawls et al. 2004, 2007). In zebrafish, the intestinal immune system was regulated by Exiguobacterium and Chryseobacterium, with the amounts of macrophages decreased and the amounts of neutrophils increased (Koch et al. 2018). In transparent gnotobiotic zebrafish, the colonization of Pseudomonas aeruginosa was enough to induce NF-κB and attenuate the expressions of both serum amyloid A (SAA) and complement factor B (CFB) (Kanther et al. 2011). Gnotobiotic zebrafish studies show that intestinal microbiota can downregulate the expression of suppressor of cytokine signaling-3 (Rawls et al. 2004) and upregulate the expressions of innate immune-related genes, such as SAA1, C-reactive protein, complement component 3 (C3), angiogenesis 4, myeloperoxidase, and glutathione peroxidase (Rawls et al. 2004, 2007). Intestinal symbiotic microorganisms colonize the gut of newly hatched zebrafish, then launch innate immunity through TLRs and MyD88 signaling (Galindo-Villegas et al. 2012). Bacillus subtilis WB800N can promote the expressions of tumor necrosis factor-α (TNF-α), IL-10, CFB, IL-1β, SAA, and MyD88 in gnotobiotic zebrafish (Tan et al. 2019a). Probiotic Lactobacillus rhamnosus could upregulate the expressions of TNF-α, IL-1β, and Beclin-1 in the gut of adult zebrafish (Gioacchini et al. 2014). Probiotic Lactobacillus plantarum ST-III enhanced the expressions of immune genes TNF-α, IL-1β, NF-κB, TLR4α, lysozyme, and IL-10, increased the diversity of gut microbiota, and attenuated the toxic impacts of triclosan through intestinal microbiota modulation in zebrafish (Zang et al. 2019). Bacillus amyloliquefaciens R8 increased the expressions of IL-1β, IL-6, IL-21, TLR1, TLR3, TLR4, and TNF-α and enhanced the innate immunity of zebrafish (Lin et al. 2018). Streptomyces sp. SH5 could enhance the immune responses and increase the expressions of TLR3, lysozyme, and nitric oxide synthase 2α in zebrafish (Liang et al. 2022). Through the above studies in zebrafish, we can initially obtain some basic information about the complex interactions of host immunity and intestinal microorganisms, which give us some reference in commercial fish species.
Host immunity senses gut microbiota
Under evolutionary selective forces, fish have formed innate immunity and an adaptive immune system consisting of B cells and T cells to distinguish opportunistic and commensal microorganisms (Yu et al. 2021). The host immune system exerts a vital function in the recognition and resistance to the invasion of pathogenic microorganisms (Liu et al. 2020b; Lv et al. 2020). The immune mucus layers of intestine, skin, and gill are rich in a large number of immune cells (including lymphocytes, plasma cells, goblet cells, granulocytes, macrophages, and antibody secretory cells) and immune effectors (including lectins, mucins, antimicrobial peptides, and immunoglobulin (Ig)), all of which jointly resist the invasion of pathogenic bacteria (Shephard 1994).
Except for extracellular metabolites, the structural components of microbial cells, especially the cell wall components, are the top layer positions that host immune cells first get in touch with and modulate immune activities (Chapot-Chartier et al. 2010). Host immune cells sense gut microorganisms through pattern recognition receptors (PRRs), which can recognize several microbe-associated molecular patterns (MAMPs), including lipopolysaccharide (LPS), microbial nucleic acid, peptidoglycan (PGN), and flagellin (Lazado and Caipang 2014), and differentially regulate downstream expression of immune molecules and maintain intestinal homeostasis. The host PRRs are microbial sensing elements that have evolved in company with symbiotic commensal microbiota and pathogens, so a similar mechanism might exist in the host immune system sensing gut beneficial microorganisms and pathogens (Murdoch and Rawls 2019). Currently, TLRs, NOD-like receptors (NLRs), C-type lectin receptors (CLRs), and peptidoglycan recognition proteins (PGRPs) are the four PRRs which were found in fish (Boltana et al. 2011).
TLRs are the most largely researched PRRs in fish, which play an important role in identifying the unique molecular patterns of microbes and modulating the innate and adaptive immune system. Fish TLRs can identify pathogen-associated molecular patterns (PAMPs), such as PGN, lipophosphoric acid (LTA), flagellin components, and LPS, activating the downstream expressions of cytokines and immune effector molecules (Ribeiro et al. 2010). For example, Staphylococcus aureus-derived PGN activates the expressions of MyD88, TLR2, TNF receptor-associated factor 6, IL-8, and NF-κB in heart cells of Labeo rohita, suggesting that TLR2-mediated microbial recognition may depend on MyD88 (Samanta et al. 2012). Recent researchers have found that fish TLRs can mediate immune system recognizing commensal microbes (Sun et al. 2014). Our previous research showed that live autochthonous Psychrobacter sp. SE6 could increase intestinal TLR2, TLR5, MyD88, transforming growth factor-β1 (TGF-β1), IL-1β, and IL-8 expressions in Epinephelus coioides, while heat-inactived SE6 only increased the TLR2 expression but did not significantly change the expressions of cytokines and MyD88. This suggests that the intestinal TLR2 signaling pathway is associated with the recognition of Psychrobacter sp. SE6, as a probiotic recognition mechanism might be independent of MyD88 (Sun et al. 2014). Hence, there may be some differences in the recognition mechanism of different symbiotic microorganisms based on TLRs in different fish species.
NLRs have crucial functions in identifying fish pathogens and activating innate immune signaling pathways. Studies have demonstrated that fish NLRs can recognize microbial ligands and perceive PAMPs or G-D-glutimyl-meso-diaminopimelic acid (iE-DAP) to activate downstream molecular pathways (Bi et al. 2017). The in vivo and in vitro studies have showed that bacteria or viruses can induce the expression of multiple NLR subfamily receptors in teleost, including NLR-X1, NLR-C3, NLR-C5, NOD1, and NOD2 (Zhang et al. 2018). When Indian major carp (Cirrhinus mrigala) were infected with pathogenic Streptococcus uberis and A. hydrophila, the NOD1 and NOD2 signaling were activated, and the downstream effector molecules IL-1β, IL-8, and interferon-γ (IFN-γ) were induced (Swain et al. 2013). The above studies suggest that NLRs may also play important functions in fish intestinal microbial recognition and immune regulation.
CLRs are produced by dendritic cells, which can recognize microbial antigens and deliver antigen information to T and B lymphocytes, thereby triggering multiple immune responses. It is found that the CLR of tilapia (Oreochromis niloticus) is homologous to mammalian natural killer cell receptor (Kikuno et al. 2004). The expression of C-type lectin (Ctl) molecules in gills, spleen, liver, and head kidney was increased significantly after Poly (I: C) and A. hydrophila challenge in Qihe crucian carp (Carassius auratus) (Wang et al. 2017). In carp (Cyprinus carpio) macrophages, CLR receptors can recognize β-dextran and mediate immunomodulatory effects (Petit et al. 2019). However, as far as we know, there are no reports about gut microbes sensed by fish CLRs, which needs further investigation.
PGRPs are the PRRs of the innate immune system and can particularly sense PGN, a peculiar component of the microbial cell wall (Hu et al. 2020). On the basis of the length of the amino acid sequence, PGRPs could be classified into short-PGRPs, intermediate-PGRPs, and long-PGRPs. PGN is an important molecule of the microbial cell wall, including Lys type (L-lysine) and Dap type (diaminoheptanedioic acid), with the PGN in Lys type mainly existing in the cell wall of Gram-positive microbes, while PGN in Dap type being the main cell wall element of Gram-negative bacteria (Royet and Dziarski 2007). In grass carp (Ctenopharyngodon idella), PGRPs can recognize pathogenic Edwardsiella tarda by combining bacterial PGN in Lys and Dap types (Li et al. 2014). In rainbow trout (Oncorhynchus mykiss), the PGRP negatively regulates the NODs-mediated antimicrobial immune response (Jang et al. 2016), which suggests that, in sensing microbes, different PRRs families also interact with each other.
The sensing of gut microorganisms by the fish intestinal immune system is summarized in Fig. 1. The immune cells sense gut microorganisms through PRRs (TLRs, NLRs, CLRs, and PGRPs), which can recognize MAMPs of microorganisms. Fish TLRs are able to recognize PAMPs, such as LTA, PGN, LPS, and flagellin components. Fish NLRs can recognize PAMPs or iE-DAP, CLRs can recognize microbial antigens, and PGRPs can specifically recognize PGN. The recognition of microorganisms by immune system would further regulate downstream signaling pathways, like mitogen-activated protein kinase (MAPK) or the NF-κB signaling pathway, and activate the downstream expression of cytokines and immune effector molecules.
The fish intestinal immune system senses gut microorganisms. The immune cells sense gut microorganisms through PRRs (TLRs, NLRs, CLRs, and PGRPs), which can recognize MAMPs of microorganisms. Fish TLRs are able to recognize PAMPs (LTA, PG, LPS, and flagellin). Fish NLRs can recognize PAMPs or iE-DAP, CLRs can recognize microbial antigens, and PGRPs can specifically recognize PG. The recognition of microorganisms by immune system would further induce downstream signaling pathways, like MAPK or the NF-κB signaling pathway, then activate the downstream expressions of cytokines and immune effector molecules
Host immunity shapes the intestinal microbiota
The immunity of fish consists of innate and adaptive immunity, which are interconnected to clear harmful substances and pathogens, and maintain host health (Fadel et al. 2018). Studies indicate that innate immunity can regulate microbial composition (Cao et al. 2021), i.e., improving the abundance of beneficial microbes and controlling harmful microbes in the intestine of fish (Thaiss et al. 2016). The immunity and intestinal microbes are intimately connected in fish, and in the first part of this review, we have summarized the zebrafish immunity in shaping the intestinal microbiota. Except for zebrafish, the relationships between immunity and gut microbes are also very close in other fish species. Immune responses were closely associated with intestinal microbes in sea cucumber (Apostichopus japonicus), the activities of alkaline phosphatase and lysozyme were strongly correlated with Shewanella, while the lectin expression levels were significantly negatively associated with Flavirhabdus (Song et al. 2019). The increased expression levels of intestinal IFN-induced helicase C domain containing protein 1, TNF-α, IL-1β, and major histocompatibility complex-1 were accompanied by the reduced relative abundances of Sphingomonas and Listonella and the enhanced relative abundances of Akkermansia, Faecalibacterium, and Bifidobacteium in turbot (Scophthalmus maximus) (Guo et al. 2022). In golden pompano (Trachinotus ovatus), the enhanced immunity (decreasing epithelial lymphocyte numbers and increasing intestinal goblet cells numbers) was accompanied by the improved diversity and richness of gut microbiota, with the relative abundance of harmful microbial taxa decreased, and the abundance of Firmicutes and Actinobacteria increased (Tan and Sun 2020).
The host immune system exerts key functions in the intestinal homeostasis of homoiothermic animals, and it can shape the microbial structure through the IgA (Petersen et al. 2019). Secretory immunoglobulin A (sIgA), an element of the adaptive immune system, is a vital protector of intestinal epithelial health, and an important factor of intestinal microbiological regulation (Dollé et al. 2016). The host secretes large amounts of IgA into the intestinal cavity, which either react extensively with microorganisms in low affinity or relatively react specifically with microorganism in high affinity, and in some cases binds to several common motifs of bacteria (Lynch and Hsiao 2019). Teleost fish has mucosal IgT, which can recognize mucosal symbiotic microorganisms and bind to their surfaces, playing a similar function as mammalian sIgA in the anti-infection process of parasitic and bacterial pathogens (Xu et al. 2016). Fish lacking IgT are highly sensitive to mucosal parasites and cannot produce compensatory IgM responses in rainbow trout. After the mucosal microbiota was disturbed, the amounts of conditional pathogenic microbes (such as Bacteroidetes) increased significantly, the surface microbes translocated, and eventually caused tissue lesions and inflammatory reactions. With the recovery level of secretory IgT, the proportion and type of IgT-coated microorganisms returned to normal level in mucosal tissue (Xu et al. 2020). This study reveals for the first time that like sIgA in mammalian, fish IgT plays an important role in maintaining intestinal mucosa immunity and microbial homeostasis.
Intestinal microbes tune host immunity
Although zebrafish is recognized as an ideal model for studying host immunity and microbial interaction, most of the current information about intestinal microbes regulating host immunity in fish are from the studies using host-derived intestinal microbes (Table 2). In common carp (Cyprinus carpio), commensal Enterococcus casseliflavus (EC-001) could significantly improve the immunity, with increased serum total protein (TP), Ig and C3 contents, and skin mucus protease activity (Akbari et al. 2021). In African catfish (Clarias gariepinus), host-derived Lactobicillus plantarum promoted the immune system, with improved levels of blood parameters, such as hemoglobin and pack cell volume, and white blood cells (Nwanna and Tope-Jegede 2017). In Nile tilapia (Oreochromis niloticus), host-derived Lactobacillus plantarum N11 and Bacillus velezensis H3.1 could promote mucosal and serum immunities, and enhance the serum lysozyme activity (Doan et al. 2018). Three host-associated Bacillus species, Bacillus velezensis TPS3N, Bacillus amyloliquefaciens TPS17, and Bacillus subtilis TPS4, were proved to significantly promote the intestinal and skin mucosal immunities of Nile tilapia (O. niloticus), with higher lysozyme, alkaline phosphatase activities, and nitric oxide and IgM contents (Kuebutornye et al. 2020). The host-derived Lactococcus lactis MM1 and Enterococcus faecium MM4 were proved to regulate the immune function and increase serum C3 levels in grouper E. coioides (Sun et al. 2012). Isolated from the intestinal tract of the healthy rainbow trout (O. mykiss), Bacillus subtilis AB1 can activate both humoral and cellular immune responses, increase the number of leucocytes, and enhance serum phagocytic activity and lysozyme activity (Newaj-Fyzul et al. 2007). Isolated from the intestine of Trachinotus ovatus, B. pumilus A97 could improve the host immune function, promote non-specific immune responses and intestinal TLR8 and TLR9 expressions in the kidney, and enhance the disease resistance (Liu et al. 2020a). Isolated from the intestine of grass carp (Ctenopharyngodon idellus), Citrobacter freundii GC01 triggers the natural mucosal immune system, induces the increasing expressions of immune system genes, such as c1s, c3, c4, c5, c7, c8a, and c8b, significantly reduces the α- and β-diversity of intestinal microbiota, and improves the relative abundance of Pasteurellales, Enterobacteriales, Citrobacter, and Neisseriales in grass carp (Xiong et al. 2020). L. lactis CLFP100 and Leuconostoc mesenteroides CLFP196 were isolated from the intestinal tract of brown trout (Balcázar et al. 2006), and they could enhance fish serum complement content and lysozyme activity (Balcázar et al. 2007) and activate head-kidney phagocytes (Balcázar et al. 2009). Our research group isolated B. clausii DE5 and B. pumilus SE5 from the gut of juvenile grouper (E. coioides) (Sun et al. 2013). The two probiotics can promote the immune function of grouper, with serum lysozyme activity, leucocyte phagocytic activity, and serum complement C3 and C4 levels increased significantly (Sun et al. 2010). The function of Bacillus pumilus SE5 is independent to its viability, as both heat-inactivated B. pumilus SE5 (Yan et al. 2016) and live B. pumilus SE5 (Yang et al. 2014) could improve the immunity of grouper (E. coioides). Our study group also demonstrated that commensal Psychrobacter sp. SE6 can regulate intestinal immunity and maintain the balance of intestinal microbiota in grouper (Sun et al. 2014), and commensal Lactococcus petauri LF3 and Bacillus siamensis LF4 can significantly increase the activity of serum lysozyme (Yang et al. 2021) and regulate the intestinal immunity and microbial structure in Japanese seabass (Lateolabrax japonicus) (Yang et al. 2022a, 2022b). It can be concluded that the host-derived beneficial microbes can regulate fish intestinal immune function, control the excessive proliferation of intestinal pathogens, and maintain the intestinal microbial homeostasis.
Cytokines are key signaling molecules for information transmission between cells and are important molecules regulating the immune cascades after the fish intestinal tract has been exposed to different microorganisms. Host-derived beneficial microbes can regulate the expression of gut pro-inflammatory cytokines, such as IL, IFN, and TNF, and anti-inflammatory cytokines, such as TGFs. Isolated from the gut of healthy Atlantic cod, live and heat-inactivated Pseudomonas sp. GP21 and Psychrobacter sp. GP12 could promote the expressions of IL-1β and IL-8 in the head kidney leukocytes of Atlantic cod (Lazado et al. 2010). Carnobacterium divergens B33 and C. maltaromaticum B26 are isolated from the intestinal tract of rainbow trout, and they can regulate the expressions of TNF-α, TGF-β, IL-1β, and IL-8 in the head kidney of rainbow trout (Kim and Austin 2006). Isolated from the gut of cobia (Rachycentron canadum), Pantoea agglomerans RCS2 could upregulate the expressions of IL-1β, TNF-α, and IL-8 in the gut of juvenile cobia (Amenyogbe et al. 2022). Isolated from the intestinal tract of Nile tilapia (O. niloticus), Rummeliibacillus stabekisii could significantly elevate expressions of the cytokine genes, like heat shock protein 70, TNF-α, IL-1β, and TGF-β in the head kidney of Nile tilapia (Tan et al. 2019b). Isolated from the intestinal tract of olive founder (Paralichthys olivaceus), Lactobacillus plantarum FGL0001 could significantly upregulate the expressions of TNF-α, IL-6, and IL-8 in the intestinal tract of olive founder (Beck et al. 2015). Isolated from the intestinal tract of Japanese seabass, L. petauri LF3 and B. siamensis LF4 could significantly reduce the expressions of TNF-α, IL-8, and IL-1β and increase the expression of TGF-β1 in the intestine of Japanese seabass (Yang et al. 2022b).
The intestinal microbes can sense fish PRRs and regulate the expressions of cytokines, then modulate the expressions of key molecules in downstream signaling pathways. Isolated from clinically healthy European seabass (Dicentrarchus labrax) larvae, probiotic candidate Vibrio lentus can modulate NF-κB pathway and upregulate the downstream molecules NF-κB inhibitor zeta and tyrosine-protein phosphatase non-receptor type 22 in sea bass (Schaeck et al. 2017). Isolated from the gut of rockfish (Sebastes schlegelii), Bacillus sp. KRF-7 was found to increase the expressions of NF-κB, IL-10, and B cell activating factors in both the kidney and spleen in rockfish (Jang et al. 2021). Our group demonstrated that B. pumilus SE5 originated LTA and PGN could enhance the intestinal imunity of grouper (E. coioides) by regulating the TLRs/MyD88 signaling (Yang et al. 2019). Based on the above studies, we postulate that fish intestinal microbes may modulate host immunity through TLRs/MyD88/NF-κB and MAPK signaling pathways, but more data are needed to clarify this.
Intestinal commensal bacteria could secrete many metabolites and nutrients, which can be involved in the development, maturation, function, and homeostasis of the immune system (Brestoff and Artis 2013). In rainbow trout (O. mykiss), administration of the shingolipids produced by Flectobacillus major can change the percentage of IgT+ to IgM+ B cells in the head kidney and regulate the mucosal homeostasis (Sepahi et al. 2016). Microorganisms within the Firmicutes phylum can generate short-chain fatty acids (SCFAs) that can offer energy for the intestinal mucosal cells (Koh et al. 2016). Adding SCFAs to the diets has beneficial functions by improving the immune systems in aquatic animals (Tran et al. 2020). SCFAs can increase intracellular bactericidal activity of head kidney macrophages via hypoxia inducible factor-1α in turbot (Scophthalmus maximus L.) (Zhang et al. 2020a). It is demonstrated that butyrate (0.2%) activated the expressions of IL-1β, IL-6, IL-8, IL-10, and TNF-α in European seabass (Dicentrarchus labrax L.) (Rimoldi et al. 2016). Besides secretory metabolites and nutrients, bacterial cell components such as bacterial DNA, LPS, PGN, and LTA have been confirmed as promising immunostimulants in fish (Giri et al. 2018). LPS could activate the innate immune system (O’Hagan et al. 2001), enhance the antibody level (Ackerman et al. 2000; Selvaraj et al. 2006), and induce the expressions of pro-inflammatory cytokines and acute-phase proteins in fish (Swain et al. 2008). Our group demonstrated that commensal Bacillus pumilus SE5-derived LTA and PGN could increase the expressions of MyD88, TLR1, TLR2, and TLR5, upregulate the expressions of antimicrobial effectors, and modulate the microbial composition in the intestine of grouper (E. coioides) (Yang et al. 2019).
Crosstalk between probiotics, intestinal immunity, and microbiota
Probiotics can improve the activities of the cytokines and innate immune cells on the surface of fish intestinal mucosa, control the proliferation of conditioned pathogens, and effectively prevent disease onset (Fazle Rohani et al. 2022). Also, probiotics can regulate the innate and adaptive immunity of fish, and then establish the correlation between intestinal and systemic immunity of fish by shaping microbial structure. Probiotics can regulate the fish microbiota in the three major mucosal tissues, namely, intestinal-associated lymphoid tissue, gill-associated lymphoid tissue, and skin-associated lymphoid tissue, thus exerting local immune effects (e.g., intestinal immunity) and systemic immune effects (e.g., cellular and humoral immunity) (Lazado and Caipang 2014).
By supplying significant signals for the development and maintenance of the immune system, a healthy microbiota has an important function in host immunophysiologic regulation (Salminen et al. 2005). The intestinal microbial composition in healthy fish is in a dynamic balance, and the pathogens are surveilled and their amounts are being controlled at an appropriate level. However, the intestinal microbial dysbiosis may disrupt immunomodulatory networks, which are closely related with gut inflammation and other immune-mediated diseases (Maynard et al. 2012). For example, Vibrio harveyi colonized in the liver led to a gut-liver immune response and substantially disrupted the intestinal microbiota balance of the pearl gentian grouper (Epinephelus lanceolatus ♂ × E. fuscoguttatus ♀), with the amounts of Photobacterium and Vibrio increased and the amounts of Lactobacillus, Blautia, Bradyrhizobium, and Faecalibaculum reduced, and causing the death of fish (Deng et al. 2020). In sea cucumbers (Apostichopus japonicus Selenka), benzo[a]pyrene exposure decreased the abundance of Firmicutes and increased the abundances of Bacteroidete and Proteobacteria, and caused the immune suppression, i.e., significantly downregulated the relative expressions of lysozyme, Ctl, and NF-κB1, and negatively impacted the gut health (Zhao et al. 2019). Therefore, the dysbiosis of intestinal microbiota could lead to the disorders of intestinal immune system and cause the diseases occurrence in fish (Pérez et al. 2010).
Probiotics are perhaps the most feasible therapeutic strategy to promote the healthy fish physiology by regulating gut microbiota and immunity (Langlois et al. 2021; Vargas-Albores et al. 2021). As one of the crucial regulators in fish immune responses, cytokines are also found to be positively regulated by probiotics (Kim and Austin 2006; He et al. 2013). Probiotic Clostridium butyricum could significantly improve the expressions of intestinal immune-related genes (MyD88, TLR2, and IL-10) and improve the diversity of the intestinal microbiota, with the abundance of Fusobacteria and Proteobacteria decreased and the abundance of Bacteroidetes increased in common carp (Cyprinus carpio L.) (Meng et al. 2021). Bacillus subtilis H2 could upregulate the expressions of IFN-α, colony stimulating factor, FOSB, IL-4, IL-11, IGHV3-11, MAPK12b, and IGHV3-21 in the intestinal tract of grass carp (Shi et al. 2020). Dietary Lactobacillus rhamnosus JCM1136 and L. lactis subsp. lactis JCM5805 could enhance the expressions of IL-1β, C-type lysozyme (lyzc), and IFN-γ, and shape the intestinal microbiota, with the abundances of Fusobacteria, Bacteroidetes, and Actinobacteria reduced and the abundance of Proteobacteria increased in Nile tilapia (Xia et al. 2018). Besides, our previous research discovered that two cell wall components of B. pumilus SE5, LTA, and PGN can enhance the expressions TLR1, TLR2, TLR5, and MyD88, activate the expression of immune effector molecules, and modulate the intestinal microbiota, i.e., increase the abundance of Lactobacillus and decrease the abundance of Vibrio significantly (Yang et al. 2019). Therefore, it is clear that probiotics/commensals can regulate fish intestinal immunity and intestinal microbiota and improve the overall health of host fish.
Summary and outlook
Intestinal microbes can improve the development and response of fish mucosal immune system, and the host immune system monitors and modulates the intestinal microbial balance. This complex interaction is very important to maintain intestinal health of fish. However, it is unclear how the intestinal immune system of fish accurately monitors and regulates the intestinal microbiota and maintains intestinal homeostasis. Existing studies have shown that both TLR and NLRs signaling may play important functions in this process, but more supporting data are needed. Commensals can regulate the immune system and intestinal microbiota of fish, improve the disease resistance, and then promote the feed utilization efficiency and growth performance. However, the molecular mechanisms of the interactions among commensals with host immunity and gut microbiota are still poorly understood. Future studies can use gnotobiotic zebrafish technology, in vitro intestinal epithelial cell culture technology, combined with RNA interference, multi-omics analysis, gene editing, and other biological techniques to further explore these issues. The elucidation of these issues will greatly promote the exploitation and application of intestinal functional microbes in fish.
Data availability
Not applicable.
References
Ackerman PA, Iwama GK, Thornton JC (2000) Physiological and immunological effects of adjuvanted Aeromonas salmonicida vaccines on juvenile rainbow trout. J Aquat Anim Health 12(2):157–164. https://doi.org/10.1577/1548-8667(200006)0122.0.CO;2
Akbari H, Shekrabi SPH, Soltani M, Mehrgan MS (2021) Effects of potential probiotic Enterococcus casseliflavus (EC-001) on growth performance, immunity, and resistance to Aeromonas hydrophila infection in common carp (Cyprinus carpio). Probiotics Antimicro 13:1316–1325. https://doi.org/10.1007/s12602-021-09771-x
Amenyogbe E, Yang EJ, Xie RT, Huang JS, Chen G (2022) Influences of indigenous isolates Pantoea agglomerans RCS2 on growth, proximate analysis, haematological parameters, digestive enzyme activities, serum biochemical parameters, antioxidants activities, intestinal morphology, disease resistance, and molecular immune response in juvenile’s cobia fish (Rachycentron canadum). Aquaculture 551:737942. https://doi.org/10.1016/j.aquaculture.2022.737942
Assefa A, Abunna F (2018) Maintenance of fish health in aquaculture: review of epidemiological approaches for prevention and control of infectious disease of fish. Vet Med Int 2018:5432497. https://doi.org/10.1155/2018/5432497
Balcázar JL, Blas ID, Ruiz-Zarzuela I, Vendrell D, Evora MD, Múzquiz JL (2006) Growth inhibition of Aeromonas species by lactic acid bacteria isolated from salmonids. MEHD 18(1):61–63. https://doi.org/10.1080/08910600600761331
Balcázar JL, Blas ID, Ruiz-Zarzuela I, Vendrell D, Calvo AC, Marquez I, Girones O, Muzquiz JL (2007) Changes in intestinal microbiota and humoral immune response following probiotic administration in brown trout (Salmo trutta). Br J Nutr 97(3):522–527. https://doi.org/10.1017/s0007114507432986
Balcázar JL, Vendrell D, Blas ID, Ruiz-Zarzuela I, Muzquiz JL (2009) Effect of Lactococcus lactis CLFP 100 and Leuconostoc mesenteroides CLFP 196 on Aeromonas salmonicida infection in brown trout (Salmo trutta). J Mol Microb Biotech 17(3):153–157. https://doi.org/10.1159/000226588
Beck BR, Kim D, Jeon JS, Lee SM, Kim HK, Kim OJ, Lee JI, Suh BS, Do HK, Lee KH, Holzapfel WH, Hwang JY, Kwon MG, Song SK (2015) The effects of combined dietary probiotics Lactococcus lactis BFE920 and Lactobacillus plantarum FGL0001 on innate immunity and disease resistance in olive founder (Paralichthys olivaceus). Fish Shellfsh Immun 42:177–183. https://doi.org/10.1016/j.fsi.2014.10.035
Bi DK, Gao YH, Chu Q, Cui JX, Xu TJ (2017) NOD1 is the innate immune receptor for iE-DAP and can activate NF-κB pathway in teleost fish. Dev Comp Immunol 76:238–246. https://doi.org/10.1016/j.dci.2017.06.012
Boltana S, Roher N, Goetz FW, MacKenzie SA (2011) PAMPs, PRRs and the genomics of gram negative bacterial recognition in fish. Dev Comp Immunol 35(12):1195–1203. https://doi.org/10.1016/j.dci.2011.02.010
Brestoff JR, Artis D (2013) Commensal bacteria at the interface of host metabolism and the immune system. Nat Immunol 14(7):676–684. https://doi.org/10.1038/ni.2640
Brugman S, Schneeberger K, Witte M, Klein MR (2014) T lymphocytes control microbial composition by regulating the abundance of Vibrio in the zebrafish gut. Gut Microbes 5(6):737. https://doi.org/10.4161/19490976.2014.972228
Cao SL, Guo JJ, Zhao WP, Yang WF, Zhang SL, Tao HZ, Li JN (2021) Impacts of oral Vibrio mimicus double-targeted DNA vaccine on the gut microbiota in grass carps (Ctenopharyngodon idella) and correlations with intestinal mucosa innate immunity. Aquaculture 533:736201. https://doi.org/10.1016/j.aquaculture.2020.736201
Chapot-Chartier MP, Vinogradov E, Sadovskaya I, Andre G, Mistou MY, Trieu-Cuot P, Furlan S, Bidnenko E, Courtin P, Péchoux C, Hols P, Dufre YF, Kulakauskas S (2010) Cell surface of Lactococcus lactis is covered by a protective polysaccharide pellicle. J Biol Chem 285(14):10464–10471. https://doi.org/10.1074/jbc.M109.082958
Deng YQ, Zhang YQ, Chen HX, Xu LW, Feng J (2020) Gut-liver immune response and gut microbiota profiling reveal the pathogenic mechanisms of Vibrio harveyi in Pearl Gentian Grouper (Epinephelus lanceolatus ♂× E. fuscoguttatus ♀). Front Immunol 11:607754. https://doi.org/10.3389/fimmu.2020.607754
Diwan AD, Harke SN, Panche AN (2023) Host-microbiome interaction in fish and shellfish: an overview. Fish Shellfish Immun Rep 4:100091. https://doi.org/10.1016/j.fsirep.2023.100091
Doan VH, Hoseinifar SH, Khanongnuch C, Kanpiengjai A, Unban K, Kim VV, Srichaiyo S (2018) Host-associated probiotics boosted mucosal and serum immunity, disease resistance and growth performance of Nile tilapia (Oreochromis niloticus). Aquaculture 491:94–100. https://doi.org/10.1016/j.aquaculture.2018.03.019
Dollé L, Tran HQ, Etienne-Mesmin L, Chassaing B (2016) Policing of gut microbiota by the adaptive immune system. BMC Med 14:1–4. https://doi.org/10.1186/s12916-016-0573-y
Earley AM, Graves CL, Shiau CE (2018) Critical role for a subset of intestinal macrophages in shaping gut microbiota in adult zebrafish. Cell Rep 25(2):424–436. https://doi.org/10.1016/j.celrep.2018.09.025
Fadel A, Plunkett A, Li W, Tessu Gyamfi VE, Nyaranga RR, Fadel F, Dakak S, Ranneh Y, Salmon Y, Ashworth JJ (2018) Modulation of innate and adaptive immune responses by arabinoxylans. J Food Biochem 42(2):e12473. https://doi.org/10.1111/jfbc.12473
Fazle Rohani MD, Majharul Islam SM, Kabir Hossain MD, Ferdous Z, Siddik MAB, Nuruzzaman M, Padeniya U, Brown C, Shahjahan MD (2022) Probiotics, prebiotics and synbiotics improved the functionality of aquafeed: upgrading growth, reproduction, immunity and disease resistance in fish. Fish Shellfish Immun 120:569–589. https://doi.org/10.1016/j.jsi.2021.12.037
Filardy AA, Ferreira JRM, Rezende RM, Kelsall BL, Oliveira RP (2023) The intestinal microenvironment shapes macrophage and dendritic cell identity and function. Immunol Lett 253:41–53. https://doi.org/10.1016/j.imlet.2023.01.003
Galindo-Villegas J, Garciá-Moreno D, Oliveira DS, Meseguer J, Mulero V (2012) Regulation of immunity and disease resistance by commensal microbes and chromatin modifications during zebrafish development. Proc Natl Acad Sci USA 109(39):15545–15546. https://doi.org/10.1073/pnas.1209920109
Ganz J (2018) Gut feelings: studying enteric nervous system development, function, and disease in the zebrafish model system. Dev Dyn 247(2):268–278. https://doi.org/10.1002/dvdy.24597
Gioacchini G, Giorgini E, Olivotto I, Maradonna F, Merrififield DL, Carnevali O (2014) The influence of probiotics on zebrafish Danio rerio innate immunity and hepatic stress. Zebrafish 11:98–106. https://doi.org/10.1089/zeb.2013.0932
Giri SS, Chi C, Jun JW, Park SC (2018) Use of bacterial subcellular components as immunostimulants in fish aquaculture. Rev Aquac 10(2):474–492. https://doi.org/10.1111/raq.12182
Guo GX, Li KX, Zhu QH, Zhao CY, Li C, He ZL, Hu SN, Ren YC (2022) Improvements of immune genes and intestinal microbiota composition of turbot (Scophthalmus maximus) with dietary oregano oil and probiotics. Aquaculture 547:737442. https://doi.org/10.1016/j.aquaculture.2021.737442
Gutierrez MW, van Tilburg BE, Changirwa D, McDonald B, Arrieta MC (2022) “Molding” immunity—modulation of mucosal and systemic immunity by the intestinal mycobiome in health and disease. Mucosal Immunol 15:573–583. https://doi.org/10.1038/s41385-022-00515-w
He SX, Zhang Y, Xu L, Yang YL, Marubashi T, Zhou ZG, Yao B (2013) Effects of dietary Bacillus subtilis C-3102 on the production, intestinal cytokine expression and autochthonous bacteria of hybrid tilapia Oreochromis niloticus♀×Oreochromis aureus♂. Aquaculture 412–413:125–130. https://doi.org/10.1016/j.aquaculture.2013.06.028
Hu ZG, Cao XB, Guo M, Li CH (2020) Identification and characterization of a novel short-type peptidoglycan recognition protein in Apostichopus japonicus. Fish Shellfish Immun 99:257–266. https://doi.org/10.1016/j.fsi.2020.02.013
Jang JH, Kim H, Jang MJ, Cho JH (2016) PGRP negatively regulates NOD-mediated cytokine production in rainbow trout liver cells. Sci Rep 6:39344. https://doi.org/10.1038/srep39344
Jang WJ, Lee SJ, Jeon MH, Kim TY, Lee JM, Hasan MT, Lee HT, Park JH, Lee BJ, Hur SW, Lee S, Kim KW, Lee EW (2021) Characterization of a Bacillus sp. KRF-7 isolated from the intestine of rockfish and effects of dietary supplementation with mannan oligosaccharide in rockfish aquaculture. Fish Shellfish Immun 119:189–192. https://doi.org/10.1016/j.fsi.2021.09.039
Jia PP, Junaid M, Wen PP, Yang YF, Li GW, Yang XG, Pei DS (2021) Role of germ-free animal models in understanding interactions of gut microbiota to host and environmental health: a special reference to zebrafish. Environ Pollut 279:116925. https://doi.org/10.1016/j.envpol.2021.116925
Kanther M, Sun XL, Mühlbauer M, Mackey LC, Flynn EJ, Bagnat M, Jobin C, Rawls JF (2011) Microbial colonization induces dynamic temporal and spatial patterns of NF-κB activation in the zebrafish digestive tract. Gastroenterology 141(1):197–207. https://doi.org/10.1053/j.gastro.2011.03.042
Kikuno R, Sato A, Mayer WE, Shintani S, Aoki T, Klein J (2004) Clustering of C-type lectin natural killer receptor-like loci in the bony fish Oreochromis niloticus. Scand J Immunol 59(2):133–142. https://doi.org/10.1111/j.0300-9475.2004.01372.x
Kim DH, Austin B (2006) Cytokine expression in leucocytes and gut cells of rainbow trout, Oncorhynchus mykiss Walbaum, induced by probiotics. Vet Immunol Immunop 114(3):297–304. https://doi.org/10.1016/j.vetimm.2006.08.015
Kinnebrew MA, Pamer EG (2012) Innate immune signaling in defense against intestinal microbes. Immunol Rev 245(1):113–131. https://doi.org/10.1111/j.1600-065x.2011.01081.x
Koch BEV, Yang SX, Lamers G, Stougaard J, Spaink HP (2018) Intestinal microbiome adjusts the innate immune setpoint during colonization through negative regulation of MyD88. Nat Commun 9(1):1–11. https://doi.org/10.1038/s41467-018-06658-4
Kogut MH, Lee A, Santin E (2020) Microbiome and pathogen interaction with the immune system. Poult Sci 99(4):1906–1913. https://doi.org/10.1016/j.psj.2019.12.011
Koh A, Vadder DF, Kovatcheva-Datchary P, Backhed F (2016) From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165:1332–1345. https://doi.org/10.1016/j.cell.2016.05.041
Kuebutornye FKA, Wang Z, Lu Y, Abarike ED, Sakyi ME, Li Y, Xie CX, Hlordzi V (2020) Effects of three host-associated Bacillus species on mucosal immunity and gut health of Nile tilapia, Oreochromis niloticus and its resistance against Aeromonas hydrophila infection. Fish Shellfish Immun 97:83–95. https://doi.org/10.1016/j.fsi.2019.12.046
Lam SH, Chua HL, Gong Z, Lam TL, Sin YM (2004) Development and maturation of the immune system in zebrafish, Danio rerio: a gene expression profiling, in situ hybridization and immunological study. Dev Comp Immunol 28(1):9–28. https://doi.org/10.1016/s0145-305x(03)00103-4
Langlois L, Akhtar N, Tam KC, Dixon B, Reid G (2021) Fishing for the right probiotic: host-microbe interactions at the interface of effective aquaculture strategies. FEMS Microbiol Rev 45(6):1–19. https://doi.org/10.1093/femsre/fuab030
Lazado CC, Caipang CMA (2014) Mucosal immunity and probiotics in fish. Fish Shellfish Immun 39:78–89. https://doi.org/10.1016/j.fsi.2014.04.015
Lazado CC, Caipang CMA, Gallage S, Brinchmann MF, Kiron V (2010) Expression profiles of genes associated with immune response and oxidative stress in Atlantic cod, Gadus morhua head kidney leukocytes modulated by live and heat-inactivated intestinal bacteria. Comp Biochem Phys B 155(3):249–255. https://doi.org/10.1016/j.cbpb.2009.11.006
Li JH, Yu ZL, Xue NN, Zou PF, Hu JY, Nie P, Chang MX (2014) Molecular cloning and functional characterization of peptidoglycan recognition protein 6 in grass carp Ctenopharyngodon idella. Dev Comp Immunol 42(2):244–255. https://doi.org/10.1016/j.dci.2013.09.014
Liang QT, Liu GX, Guo Z, Wang YT, Xu ZH, Ren YX, Zhang QZ, Cui M, Zhao XQ, Xu DL (2022) Application of potential probiotic strain Streptomyces sp. SH5 on anti-Aeromonas infection in zebrafish larvae. Fish Shellfish Immun 127:375–385. https://doi.org/10.1016/j.fsi.2022.06.049
Lin YS, Saputra F, Chen YC, Hu SY (2018) Dietary administration of Bacillus amyloliquefaciens R8 reduces hepatic oxidative stress and enhances nutrient metabolism and immunity against Aeromonas hydrophila and Streptococcus agalactiae in zebrafish (Danio rerio). Fish Shellfish Immun 86:410–419. https://doi.org/10.1016/j.fsi.2018.11.047
Liu SB, Wang SF, Cai Y, Li EC, Ren ZL, Wu Y, Guo WL, Sun Y, Zhou YC (2020a) Beneficial effects of a host gut-derived probiotic, Bacillus pumilus, on the growth, non-specific immune response and disease resistance of juvenile golden pompano. Trachinotus Ovatus Aquaculture 514:734446. https://doi.org/10.1016/j.aquaculture.2019.734446
Liu XD, Sun W, Zhang YB, Zhou YF, Xu JW, Gao XJ, Zhang SM, Zhang XJ (2020b) Impact of Aeromonas hydrophila and infectious spleen and kidney necrosis virus infections on susceptibility and host immune response in Chinese perch (Siniperca chuatsi). Fish Shellfish Immun 105:117–125. https://doi.org/10.1016/j.fsi.2020.07.012
Liu C, Zhao LP, Shen YQ (2021a) A systematic review of advances in intestinal microflora of fish. Fish Physiol Biochem 47(6):2041–2053. https://doi.org/10.1007/s10695-021-01027-3
Liu X, Wang H, Zhao H (2021b) Prevalence of antibiotic resistance genes in wastewater collected from ornamental fish market in northern China. Environ Pollut 271:116316. https://doi.org/10.1016/j.envpol.2020.116316
Lv YN, Xu Q, Mao Y, Xu YD, Zhang R, Zhong H, Zhou Y, Xiao J, Du MK, Song HY, Liang YH, Yan JP (2020) TRAF3 of blunt snout bream participates in host innate immune response to pathogenic bacteria via NF-κB signaling pathway. Fish Shellfish Immun 104:592–604. https://doi.org/10.1016/j.fsi.2020.06.022
Lynch JB, Hsiao EY (2019) Microbiomes as sources of emergent host phenotypes. Science 365(6460):1405–1409. https://doi.org/10.1126/science.aay0240
Maynard CL, Elson CO, Hatton RD, Weaver CT (2012) Reciprocal interactions of the intestinal microbiota and immune system. Nature 489(7415):231–241. https://doi.org/10.1038/nature11551
Meng XL, Wu SK, Hu WP, Zhu ZX, Yang GK, Zhang YM, Qin CB, Yang LP, Nie GX (2021) Clostridium butyricum improves immune responses and remodels the intestinal microbiota of common carp (Cyprinus carpio L.). Aquaculture 530:735753. https://doi.org/10.1016/j.aquaculture.2020.735753
Murdoch CC, Rawls JF (2019) Commensal microbiota regulate vertebrate innate immunity-insights from the zebrafish. Front Immunol 10:2100. https://doi.org/10.3389/fimmu.2019.02100
Nadal AL, Ikeda-Ohtsubo W, Sipkema D, Peggs D, McGurk C, Forlenza M, Wiegertjes GF, Brugman S (2020) Feed, microbiota, and gut immunity: using the zebrafish model to understand fish health. Front Immunol 11:114. https://doi.org/10.3389/fimmu.2020.00114
Newaj-Fyzul A, Adesiyun AA, Mutani A, Ramsubhag A, Brunt J, Austin B (2007) Bacillus subtilis AB1 controls Aeromonas infection in rainbow trout (Oncorhynchus mykiss, Walbaum). J Appl Microbiol 103(5):1699–1706. https://doi.org/10.1111/j.1365-2672.2007.03402.x
Nwanna LC, Tope-Jegede H (2017) Effects of dietary Lactobicillus plantarum on the growth, carcass quality, and immune response of African catfish (Clarias gariepinus) challenged with Salmonella typhi. J Appl Aquacult 29(1):62–80. https://doi.org/10.1080/10454438.2016.1209713
O’Hagan DT, MacKichan ML, Singh M (2001) Recent developments in adjuvants for vaccines against infectious diseases. Biomol Eng 18(3):69–85. https://doi.org/10.1016/s1389-0344(01)00101-0
Ou WH, Yu GJ, Zhang YJ, Mai KS (2021) Recent progress in the understanding of the gut microbiota of marine fishes. Mar Life Sci Tech 3(4):434–448. https://doi.org/10.1007/s42995-021-00094-y
Pérez T, Balcázar JL, Ruiz-Zarzuela I, Halaihel N, Vendrell D, Blas DI, Múzquiz JL (2010) Host–microbiota interactions within the fish intestinal ecosystem. Mucosal Immunol (1933-0219) 3(4):355–360. https://doi.org/10.1038/mi.2010.12
Petersen C, Bell R, Klag KA, Lee SH, Soto R, Ghazaryan A, Buhrke K, Ekiz HA, Ost KS, Boudina S, O’Connell RM, Cox JE, Villanueva CJ, Stephens WZ, Round JL (2019) T cell–mediated regulation of the microbiota protects against obesity. Science 365:eaat9351. https://doi.org/10.1126/science.aat9351
Petit J, Bailey EC, Wheeler RT, De Oliveira CAF, Forlenza M, Wiegertjes GF (2019) Studies into β-glucan recognition in fish suggests a key role for the C-type lectin pathway. Front Immunol 10:280. https://doi.org/10.3389/fimmu.2019.00280
Rawls JF, Samuel BS, Gordon JI (2004) Gnotobiotic zebrafish reveal evolutionarily conserved responses to the gut microbiota. Proc Natl Acad Sci USA 101(13):4596–4601. https://doi.org/10.1073/pnas.0400706101
Rawls JF, Mahowald MA, Goodman AL, Trent CM, Gordon JI (2007) In vivo imaging and genetic analysis link bacterial motility and symbiosis in the zebrafish gut. Proc Natl Acad Sci USA 104(18):7622–7627. https://doi.org/10.1073/pnas.0702386104
Ribeiro CMS, Hermsen T, Taverne-Thiele AJ, Savelkoul HFJ, Wiegertjes GF (2010) Evolution of recognition of ligands from gram-positive bacteria: similarities and differences in the TLR2-mediated response between mammalian vertebrates and teleost fish. J Immunol 184(5):2355–2368. https://doi.org/10.4049/jimmunol.0900990
Rimoldi S, Finzi G, Ceccotti C, Girardello R, Grimaldi A, Ascione C, Terova G (2016) Butyrate and taurine exert a mitigating effect on the inflamed distal intestine of European sea bass fed with a high percentage of soybean meal. Fish Aquat Sci 19:40. https://doi.org/10.1186/s41240-016-0041-9
Royet J, Dziarski R (2007) Peptidoglycan recognition proteins: pleiotropic sensors and effectors of antimicrobial defences. Nat Rev Microbiol 5(4):264–277. https://doi.org/10.1038/nrmicro1620
Ruff WE, Greiling TM, Kriegel MA (2020) Host-microbiota interactions in immune-mediated diseases. Nat Rev Microbiol 9:521. https://doi.org/10.1038/s41579-020-0367-2
Salminen SJ, Gueimonde M, Isolauri E (2005) Probiotic that modify disease risk. J Nutr 135(5):1294–1298. https://doi.org/10.1093/jn/135.5.1294
Samanta M, Swain B, Basu M, Panda P, Mohapatra GB, Sahoo BR, Maiti NK (2012) Molecular characterization of toll-like receptor 2 (TLR2), analysis of its inductive expression and associated down-stream signaling molecules following ligands exposure and bacterial infection in the Indian major carp, rohu (Labeo rohita). Fish Shellfish Immun 32(3):411–425. https://doi.org/10.1016/j.fsi.2011.11.029
Schaeck M, Reyes-López FE, Vallejos-Vidal E, Cleemput JV, Duchateau L, Broeck WVD, Tort L, Decostere A (2017) Cellular and transcriptomic response to treatment with the probiotic candidate Vibrio lentus in gnotobiotic sea bass (Dicentrarchus labrax) larvae. Fish Shellfish Immun 63:147–156. https://doi.org/10.1016/j.fsi.2017.01.028
Selvaraj V, Sampath K, Sekar V (2006) Adjuvant and immunostimulatory effects of β-glucan administration in combination with lipopolysaccharide enhances survival and some immune parameters in carp challenged with Aeromonas hydrophila. Vet Immunol Immunop 114(1):15–24. https://doi.org/10.1016/j.vetimm.2006.06.011
Sepahi A, Cordero H, Goldfine H, Esteban MÁ, Salinas I (2016) Symbiont-derived sphingolipids modulate mucosal homeostasis and B cells in teleost fish. Sci Rep 6:srep39054. https://doi.org/10.1038/srep39054
Shephard KL (1994) Functions for fish mucus. Rev Fish Biol Fisher 4(4):401–429
Shi F, Zi YJ, Lu ZJ, Li FL, Yang MX, Zhan FB, Li YN, Li J, Zhao LJ, Lin L, Qin ZD (2020) Bacillus subtilis H2 modulates immune response, fat metabolism and bacterial flora in the gut of grass carp (Ctenopharyngodon idellus). Fish Shellfish Immun 106:8–20. https://doi.org/10.1016/j.fsi.2020.06.061
Song XJ, Feng ZF, Zhang YP, Zhu W (2019) Regulation of dietary astragalus polysaccharide (APS) supplementation on the non-specific immune response and intestinal microbiota of sea cucumber Apostichopus japonicus. Fish Shellfish Immun 94:517–524. https://doi.org/10.1016/j.fsi.2019.09.049
Stagaman K, Burns AR, Guillemin K, Bohannan BJM (2017) The role of adaptive immunity as an ecological filter on the gut microbiota in zebrafish. ISME 11:1630–1639. https://doi.org/10.1038/ismej.2017.28
Stream A, Madigan CA (2022) Zebrafish: an underutilized tool for discovery in host-microbe interactions. Trends Immunol 43(6):426–437. https://doi.org/10.1016/j.it.2022.03.011
Sun YZ, Yang HL, Ma RL, Lin WY (2010) Probiotic applications of two dominant gut Bacillus strains with antagonistic activity improved the growth performance and immune responses of grouper Epinephelus coioides. Fish Shellfish Immun 29(5):803–809. https://doi.org/10.1016/j.fsi.2010.07.018
Sun YZ, Yang HL, Ma RL, Song K, Li JS (2012) Effect of Lactococcus lactis and Enterococcus faecium on growth performance, digestive enzymes and immune response of grouper Epinephelus coioides. Aquac Nutr 18(3):281–289. https://doi.org/10.1111/j.1365-2095.2011.00894.x
Sun YZ, Yang HL, Huang KP, Ye JD, Zhang CX (2013) Application of autochthonous Bacillus bioencapsulated in copepod to grouper Epinephelus coioides larvae. Aquaculture 392–395:44–50. https://doi.org/10.1016/j.aquaculture.2013.01.037
Sun YZ, Xia HQ, Yang HL, Wang YL, Zou WC (2014) TLR2 signaling may play a key role in the probiotic modulation of intestinal microbiota in grouper Epinephelus coioides. Aquaculture 430:50–56. https://doi.org/10.1016/j.aquaculture.2014.03.042
Sundha H, Finne-Fridell F, Ellisc T, Tarangerd GL, Niklassona L, Pettersene EF, Wergelande HI, Sundell K (2019) Reduced water quality associated with higher stocking density disturbs the intestinal barrier functions of Atlantic salmon (Salmo salar L.). Aquaculture 137:734356. https://doi.org/10.1016/j.aquaculture.2019.734356
Swain P, Nayak SK, Nanda PK, Dash S (2008) Biological effects of bacterial lipopolysaccharide (endotoxin) in fish: a review. Fish Shellfish Immun 25(3):191–201. https://doi.org/10.1016/j.fsi.2008.04.009
Swain B, Basu M, Samanta M (2013) NOD1 and NOD2 receptors in mrigal (Cirrhinus mrigala): inductive expression and downstream signalling in ligand stimulation and bacterial infections. J Biosci 38(3):533–548. https://doi.org/10.1007/s12038-013-9330-y
Tan XH, Sun ZZ (2020) Dietary dandelion extract improved growth performance, immunity, intestinal morphology and microbiota composition of golden pompano Trachinotus ovatus. Aquacult Rep 18:100491. https://doi.org/10.1016/j.aqrep.2020.100491
Tan F, Limbu SM, Qian Y, Qiao F, Du ZY, Zhang ML (2019a) The responses of germ-free zebrafish (Danio rerio) to varying bacterial concentrations, colonization time points, and exposure duration. Front Microbiol 10:2156. https://doi.org/10.3389/fmicb.2019.02156
Tan HY, Chen SW, Hu SY (2019b) Improvements in the growth performance, immunity, disease resistance, and gut microbiota by the probiotic Rummeliibacillus stabekisii in Nile tilapia (Oreochromis niloticus). Fish Shellfish Immun 92:265–275. https://doi.org/10.1016/j.fsi.2019.06.027
Teame T, Zhang Z, Ran C, Zhang HL, Yang YL, Ding QW, Xie MX, Gao CC, Ye YG, Duan M, Zhou ZG (2019) The use of zebrafish (Danio rerio) as biomedical models. Anim Front 9(3):68–77. https://doi.org/10.1093/af/vfz020
Thaiss CA, Zmora N, Levy M, Elinav E (2016) The microbiome and innate immunity. Nature 7610:65. https://doi.org/10.1038/nature18847
Tran NT, Li ZZ, Wang SQ, Zheng HP, Aweya JJ, Wen XB, Li SK (2020) Progress and perspectives of short-chain fatty acids in aquaculture. Rev Aquac 12:283–298. https://doi.org/10.1111/raq.12317
Vaart MVD, Soest JJV, Spaink HP, Meijer AH (2013) Functional analysis of a zebrafish myd88 mutant identifies key transcriptional components of the innate immune system. Dis Model Mech 6(3):841–854. https://doi.org/10.1242/dmm.010843
Vargas-Albores F, Martínez-Córdova LR, Hernández-Mendoza A, Cicala F, Lago-Lestón A, Martínez-Porchas M (2021) Therapeutic modulation of fish gut microbiota, a feasible strategy for aquaculture? Aquaculture 544:737050. https://doi.org/10.1016/j.aquaculture.2021.737050
Wang L, Zhang J, Kong XH, Zhao XL, Pei C, Li L (2017) A C-type lectin, Nattectin-like protein (CaNTC) in Qihe crucian carp Carassius auratus: binding ability with LPS, PGN and various bacteria, and agglutinating activity against bacteria. Fish Shellfish Immun 67:382–392. https://doi.org/10.1016/j.fsi.2017.06.012
Wang J, Jaramillo-Torres A, Li YX, Brevik ØJ, Jakobsen JV, Kortner TM, Krogdahl Å (2022) Gut health and microbiota in out-of-season Atlantic Salmon (Salmo salar L.) smolts before and after seawater transfer under commercial arctic conditions: modulation by functional feed ingredients. Front Mar Sci 9:860081. https://doi.org/10.3389/fmars.2022.860081
Wang LC, Chen LH, Chiu YC, Liou CY, Chen HC, Lu CY, Chen JL (2023) Teleost skin microbiome: an intimate interplay between the environment and the host immunity. Fish Shellfish Immun 139:108869. https://doi.org/10.1016/j.fsi.2023.108869
Xia Y, Lu MX, Chen G, Cao JM, Gao FY, Wang M, Liu ZG, Zhang DF, Zhu HP, Yi MM (2018) Effects of dietary Lactobacillus rhamnosus JCM1136 and Lactococcus lactis subsp. lactis JCM5805 on the growth, intestinal microbiota, morphology, immune response and disease resistance of juvenile Nile tilapia, Oreochromis niloticus. Fish Shellfish Immun 76:368–379. https://doi.org/10.1016/j.fsi.2018.03.020
Xiong JB, Nie L, Chen J (2019) Current understanding on the roles of gut microbiota in fish disease and immunity. Zool Res 40(2):70–76. https://doi.org/10.24272/j.issn.2095-8137.2018.069
Xiong F, Qin L, Hao YT, Zhao D, Li WX, Zou H, Li M, Wu SG, Wang GT (2020) Gut microbiota modulation and immunity response induced by Citrobacter freundii strain GC01 in grass carp (Ctenopharyngodon idellus). Aquaculture 521:735015. https://doi.org/10.1016/j.aquaculture.2020.735015
Xu Z, Takizawa F, Parra D, Gómez D, Jørgensen LVG, Lapatra SE, Sunyer JO (2016) Mucosal immunoglobulins at respiratory surfaces mark an ancient association that predates the emergence of tetrapods. Nat Commun 7(1):10728. https://doi.org/10.1038/ncomms10728
Xu Z, Takizawa F, Casadei E, Shibasaki Y, Ding Y, Sauters TJC, Yu Y, Salinas I, Sunyer JO (2020) Specialization of mucosal immunoglobulins in pathogen control and microbiota homeostasis occurred early in vertebrate evolution. Sci Immunol 5:3254. https://doi.org/10.1126/sciimmunol.aay3254
Xue J, Ajuwon KM, Fang R (2020) Mechanistic insight into the gut microbiome and its interaction with host immunity and inflammation. Anim Nutr 6(4):421–428. https://doi.org/10.1016/j.aninu.2020.05.007
Yan YY, Xia HQ, Yang HL, Hoseinifar SH, Sun YZ (2016) Effects of dietary live or heat-inactivated autochthonous Bacillus pumilus SE5 on growth performance, immune responses and immune gene expression in grouper Epinephelus coioides. Aquac Nutr 22(3):698–707. https://doi.org/10.1111/anu.12297
Yang HL, Xia HQ, Ye YD, Zou WC, Sun YZ (2014) Probiotic Bacillus pumilus SE5 shapes the intestinal microbiota and mucosal immunity in grouper Epinephelus coioides. Dis Aquat Organ 111(2):119–127. https://doi.org/10.3354/dao02772
Yang HT, Zou SS, Zhai LJ, Wang Y, Zhang FM, An LG, Yang GW (2017) Pathogen invasion changes the intestinal microbiota composition and induces innate immune responses in the zebrafish intestine. Fish Shellfish Immun 17:35–42. https://doi.org/10.1016/j.fsi.2017.09.075
Yang HL, Sun YZ, Hu X, Ye JD, Lu KL, Hu LH, Zhang JJ (2019) Bacillus pumilus SE5 originated PG and LTA tuned the intestinal TLRs/MyD88 signaling and microbiota in grouper (Epinephelus coioides). Fish Shellfish Immun 88:266–271. https://doi.org/10.1016/j.fsi.2019.03.005
Yang HL, Liu ZY, Jian JT, Ye JD, Sun YZ (2021) Host-associated Bacillus siamensis and Lactococcus petauri improved growth performance, innate immunity, antioxidant activity and ammonia tolerance in juvenile Japanese seabass (Lateolabrax japonicus). Aquacult Nutr 27:2739–2748. https://doi.org/10.1111/anu.13399
Yang HL, Liu ZY, Jian JT, Liu ZX, Ye JD, Sun YZ (2022a) Host-derived probiotics shape the intestinal microbial composition, but not putative function in juvenile Japanese seabass (Lateolabrax japonicus). Front Mar Sci 9:878633. https://doi.org/10.3389/fmars.2022.878633
Yang HL, Liu ZY, Jin YM, Liu ZX, Zhang BY, Yuan ZH, Ye JD, Sun YZ (2022b) Preventive and reparative functions of host-associated probiotics against soybean meal induced growth, immune suppression and gut injury in Japanese seabass (Lateolabrax japonicus). Fish Shellfish Immun 128:651–663. https://doi.org/10.1016/j.fsi.2022.08.034
Yu YY, Ding LG, Huang ZY, Xu HY, Xu Z (2021) Commensal bacteria-immunity crosstalk shapes mucosal homeostasis in teleost fish. Rev Aquacult 13(4):2322–2343. https://doi.org/10.1111/raq.12570
Zang LX, Ma Y, Huang WH, Ling YH, Sun LM, Wang XD, Zeng AB, Dahlgren RA, Wang CH, Wang HL (2019) Dietary Lactobacillus plantarum ST-III alleviates the toxic effects of triclosan on zebrafish (Danio rerio) via gut microbiota modulation. Fish Shellfish Immun 84:1157–1169. https://doi.org/10.1016/j.fsi.2018.11.007
Zhang L, Gao ZY, Yu L, Zhang B, Wang J, Zhou J (2018) Nucleotide-binding and oligomerization domain (NOD)-like receptors in teleost fish: current knowledge and future perspectives. J Fish Dis 9:1317. https://doi.org/10.1111/jfd.12841
Zhang JJ, Zhang H, Liu M, Lan YW, Sun HY, Mai KS, Wan M (2020a) Short-chain fatty acids promote intracellular bactericidal activity in head kidney macrophages from turbot (Scophthalmus maximus L.) via hypoxia inducible factor-1α. Front Immunol 11:615536. https://doi.org/10.3389/fimmu.2020.615536
Zhang Ml, Shan CJ, Tan F, Limbu SM, Chen LQ, Du ZY (2020b) Gnotobiotic models: powerful tools for deeply understanding intestinal microbiota-host interactions in aquaculture. Aquaculture 517:734800. https://doi.org/10.1016/j.aquaculture.2019.734800
Zhao Y, Liu H, Wang Q, Li BJ, Zhang HX, Pi YR (2019) The effects of benzo[a]pyrene on the composition of gut microbiota and the gut health of the juvenile sea cucumber Apostichopus japonicus Selenka. Fish Shellfish Immun 93:369–379. https://doi.org/10.1016/j.fsi.2019.07.073
Zou PF, Chang MX, Li Y, Xue NN, Li JH, Chen SN, Nie P (2016) NOD2 in zebrafish functions in antibacterial and also antiviral responses via NF-κB, and also MDA5, RIG-I and MAVS. Fish Shellfish Immun 55:173–185. https://doi.org/10.1016/j.fsi.2016.05.031
Funding
This work was supported by the National Natural Science Foundation of China (32072990; 32303022), Xiamen Marine and Fisheries Development Fund (19CZP018HJ04), Science and Technology Major/Special Project of Fujian Province (2021NZ029022), and Natural Science Foundation of Fujian Province (2021J05157).
Author information
Authors and Affiliations
Contributions
Conceptualization, Y.Z.S., B.Y.Z., and H.L.Y.; writing—original draft preparation, B.Y.Z., H.L.Y., G.H.C., Q.J.N.; writing—review and editing, G.H.C., H.L.Y., and Y.Z.S.; supervision and funding acquisition, Y.Z.S. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, B., Yang, H., Cai, G. et al. The interactions between the host immunity and intestinal microorganisms in fish. Appl Microbiol Biotechnol 108, 30 (2024). https://doi.org/10.1007/s00253-023-12934-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-023-12934-1