Abstract
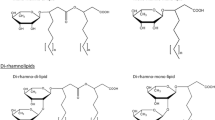
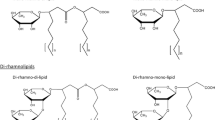
We previously discovered that Pseudomonas chlororaphis NRRL B-30761 produces monorhamnolipids (R1Ls) with predominantly 3-hydroxydodecenoyl-3-hydroxydecanoate (C12:1-C10) or 3-hydroxydodecanoyl-3-hydroxydecanoate (C12-C10) as the lipid moiety under static growth conditions only. We have now cloned, sequenced, and analyzed in silico the gene locus of NRRL B-30761 containing the putative coding sequences of rhamnosyltransferase chain A (rhlA Pch , 894 bps), rhamnosyltransferase chain B (rhlB Pch , 1272 bps), and N-acyl-homoserine lactone-dependent transcriptional regulatory protein (rhlR Pch , 726 bps). The putative gene products RhlAPch (297 amino acid residues or a.a.), RhlBPch (423 a.a.), and RhlRPch (241 a.a.) only have between 60 and 65 % a.a. identities to their respective closest matched homologs in P. aeruginosa. Polymerase chain reaction (PCR)-based assay did not detect the presence of rhamnosyltransferase C gene (rhlC) in P. chlororaphis, suggesting a genetic basis for the lack of dirhamnose-lipid (R2L) synthesis in this organism. We thus genetically constructed an R2L-synthesizing P. chlororaphis by expressing a rhamnosyltransferase C (rhlC) gene of P. aeruginosa using an expression vector (pBS29-P2-gfp) containing a Pseudomonas syringae promoter. The R2L/R1L ratio is 2.4 in the rhamnolipid (RL) sample isolated from the genetically engineered (GE) P. chlororaphis [pBS29-P2-rhlC], in contrast to undetectable R2L in the GE P. chlororaphis [pBS29-P2-gfp] control cells based on LC-MS analysis. The critical micelle concentrations of the R2L and R1L samples from GE P. chlororaphis [pBS29-P2-rhlC] and the control [pBS29-P2-gfp] cells were ca. 0.1 mM, and their minimum surface tensions were ca. 26 mN/m with no significant difference.




Similar content being viewed by others
References
Abdel-Mawgoud AM, Lépine F, Déziel E (2010) Rhamnolipids: diversity of structures, microbial origins and roles. Appl Microbiol Biotechnol 86:1323–1336
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (1987) Current protocols in molecular biology. Wiley, NY
Cabrera-Valladares N, Richardson AP, Olvera C, Treviño LG, Déziel E, Lépine F, Soberón-Chávez G (2006) Monorhamnolipids and 3-(3-hydroxyalkanoyloxy)alkanoic acids (HAAs) production using Escherichia coli as a heterologous host. Appl Microbiol Biotechnol 73:187–194
Campos-García J, Caro AD, Nájera R, Miller-Maier RM, Al-Tahhan RA, Soberón-Chávez G (1998) The Pseudomonas aeruginosa rhlG gene encodes an NADPH-dependent β- ketoacyl reductase which is specifically involved in rhamnolipid synthesis. J Bacteriol 180:4442–4451
Chen ML, Penfold J, Thomas RK, Smyth TJP, Perfumo A, Marchant R, Banat IM, Stevenson P, Parry A, Tucker I, Grillo I (2010a) Solution self-assembly and adsorption at the air-water interface of the monorhamnose and dirhamnose rhamnolipids and their mixtures. Langmuir 26:18281–18292
Chen ML, Penfold J, Thomas RK, Smyth TJP, Perfumo A, Marchant R, Banat IM, Stevenson P, Parry A, Tucker I, Grillo I (2010b) Mixing behavior of the biosurfactant, rhamnolipid, with a conventional anionic surfactant, sodium dodecyl benzene sulfonate. Langmuir 26:17958–17968
Choi MH, Xu J, Gutierrez M, Yoo T, Cho Y-H, Yoon SC (2011) Metabolic relationship between polyhydroxyalkanoic acid and rhamnolipid synthesis in Pseudomonas aeruginosa: comparative 13C NMR analysis of the products in wild-type and mutants. J Biotechnol 151:30–42
Déziel E, Lépine F, Milot S, Villemur R (2003) rhlA is required for the production of a novel biosurfactant promoting swarming motility in Pseudomonas aeruginosa: 3-(3-hydroxyalkanoyloxy)alkanoic acids (HAAs), the precursors of rhamnolipids. Microbiology 149:2005–2013
Faivre V, Rosilio V (2010) Interest of glycolipids in drug delivery: from physicochemical properties to drug targeting. Expert Opin Drug Deliv 7:1031–1048
Giani C, Wullbrandt D, Rothert R, Meiwes J (1997) Pseudomonas aeruginosa and its use in a process for the biotechnological preparation of L-rhamnose. U.S. Patent No. 5,658,793
Gunther NW IV, Nuñez A, Fett W, Solaiman DKY (2005) Production of rhamnolipids by Pseudomonas chlororaphis, a nonpathogenic bacterium. Appl Environ Microbiol 71:2288–2293
Gunther NW IV, Nuñez A, Fortis L, Solaiman DKY (2006) Proteomic based investigation of rhamnolipid production by Pseudomonas chlororaphis strain NRRL B-30761. J Ind Microbiol Biotechnol 33:914–920
Gunther NW, Solaiman DKY, Fett WF (2007) Processes for the production of rhamnolipids. U.S. Patent No. 7,202,063
Hoffmann N, Amara AA, Beermann BB, Qi Q, Hinz H-J, Rehm BHA (2002) Biochemical characterization of the Pseudomonas putida 3-hydroxyacyl ACP:CoA transacylase, which diverts intermediates of fatty acid de novo biosynthesis. J Biol Chem 277:42926–42936
Hošková M, Schreiberová O, Ježdík R, Chudoba J, Masák J, Sigler K, Řezanka T (2013) Characterization of rhamnolipids produced by non-pathogenic Acinetobacter and Enterobacter bacteria. Bioresour Technol 130:510–516
Jarvis FG, Johnson MJ (1949) A glyco-lipide produced by Pseudomonas aeruginosa. J Am Chem Soc 71:4128–4130
Kitamoto D, Yanagishita H, Shinbo T, Nakane T, Kamisawa C, Nakahara T (1993) Surface active properties and antimicrobial activities of mannosylerythritol lipids as biosurfactants produced by Candida antarctica. J Biotechnol 29:91–96
Lourith N, Kanlayavattanakul M (2009) Natural surfactants used in cosmetics: glycolipids. Int J Cosmet Sci 31:255–261
Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang D, Zhang N, Zheng C, Bryant SH (2011) CDD: a conserved domain database for the functional annotation of proteins. Nucleic Acids Res 39:D225–D229
Miller DJ, Zhang Y-M, Rock CO, White SW (2006) Structure of RhlG, an essential β-ketoacyl reductase in the rhamnolipid biosynthetic pathway of Pseudomonas aeruginosa. J Biol Chem 281:18025–18032
Müller MM, Hörmann B, Kugel M, Syldatk C, Hausmann R (2011) Evaluation of rhamnolipid production capacity of Pseudomonas aeruginosa PAO1 in comparison to the rhamnolipid over-producer strains DSM 7108 and DSM 2874. Appl Microbiol Biotechnol 89:585–592
Müller MM, Kügler JH, Henkel M, Gerlitzki M, Hörmann B, Pöhnlein M, Syldatk C, Hausmann R (2012) Rhamnolipids - next generation surfactants? J Biotechnol 162:366–380
Neilson JW, Zhang L, Veres-Schalnat TA, Chandler KB, Neilson CH, Crispin JD, Pemberton JE, Maier RM (2010) Cadmium effects on transcriptional expression of rhlB/rhlC genes and congener distribution of monorhamnolipid and dirhamnolipid in Pseudomonas aeruginosa IGB83. Appl Microbiol Biotechnol 88:953–963
Nguyen TT, Sabatini DA (2011) Characterization and emulsification properties of rhamnolipid and sophorolipid biosurfactants and their applications. Int J Mol Sci 12:1232–1244
Nitschke M, Costa SGVAO, Contiero J (2011) Rhamnolipids and PHAs: recent reports on Pseudomonas-derived molecules of increasing industrial interest. Process Biochem 46:621–630
Nuñez A, Ashby RD, Foglia TA, Solaiman DKY (2004) LC/MS analysis and lipase modification of the sophorolipids produced by Rhodotorula bogoriensis. Biotechnol Lett 26:1087–1093
Ochsner UA, Reiser J, Fiechter A, Witholt B (1995) Production of Pseudomonas aeruginosa rhamnolipid biosurfactants in heterologous hosts. Appl Environ Microbiol 61:3503–3506
Otto RT, Daniel H-J, Pekin G, Müller-Decker K, Fürstenberger G, Reuss M, Syldatk C (1999) Production of sophorolipids from whey. Appl Microbiol Biotechnol 52:495–501
Özdemir G, Peker S, Helvaci SS (2004) Effect of pH on the surface and interfacial behavior of rhamnolipids R1 and R2. Colloids Surf A 234:135–143
Parra JL, Guinea J, Manresa MA, Robert M, Mercadé ME, Comelles F, Bosch MP (1989) Chemical characterization and physicochemical behavior of biosurfactants. JAOCS 66:141–145
Pham TH, Webb JS, Rehm BHA (2004) The role of polyhydroxyalkanoate biosynthesis by Pseudomonas aeruginosa in rhamnolipid and alginate production as well as stress tolerance and biofilm formation. Microbiology 150:3405–3413
Piljac T, Piljac G (2007) Use of rhamnolipids in wound healing, treating burn shock, atherosclerosis, organ transplants, depression, schizophrenia and cosmetics. U.S. Patent No. 7,262,171 B1
Pinzon NM, Zhang Q, Koganti S, Ju L-K (2009) Advances in bioprocess development of rhamnolipid and sophorolipid production. In: Hayes DG, Kitamoto D, Solaiman DKY, Ashby RD (eds) Biobased surfactants and detergents: synthesis, properties, and applications, chapter 4. AOCS Press, Urbana, pp 77–105
Prithiviraj B, Bais HP, Weir T, Suresh B, Najarro EH, Dayakar BV, Schweizer HP, Vivanco JM (2005) Down regulation of virulence factors of Pseudomonas aeruginosa by salicylic acid attenuates its virulence on Arabidopsis thaliana and Caenorhabditis elegans. Infect Immun 73:5319–5328
Rahim R, Ochsner UA, Olvera C, Graninger M, Messner P, Lam JS, Soberón-Chávez G (2001) Cloning and functional characterization of the Pseudomonas aeruginosa rhlC gene that encodes rhamnosyltransferase 2, an enzyme responsible for di-rhamnolipid biosynthesis. Mol Microbiol 40:708–718
Rehm BHA, Krüger N, Steinbüchel A (1998) A new metabolic link between fatty acid de novo synthesis and polyhydroxyalkanoic acid synthesis. The phaG gene from Pseudomonas putida KT2440 encodes a 3-hydroxyacyl-acyl carrier protein-coenzyme A transferase. J Biol Chem 273:24044–24051
Rehm BHA, Mitsky TA, Steinbüchel A (2001) Role of fatty acid de novo biosynthesis in polyhydroxyalkanoic acid (PHA) and rhamnolipid synthesis by pseudomonads: establishment of the transacylase (PhaG)-mediated pathway for PHA biosynthesis in Escherichia coli. Appl Environ Microbiol 67:3102–3109
Řezanka T, Siristova L, Sigler K (2011) Rhamnolipid-producing thermophilic bacteria of species Thermus and Meiothermus. Extremophiles 15:697–709
Solaiman DKY (1998) Genetic transformation of Pseudomonas oleovorans by electroporation. Biotechnol Tech 12:829–832
Solaiman DKY (2000) PCR cloning of Pseudomonas resinovorans polyhydroxyalkanoate biosynthesis genes and expression in Escherichia coli. Biotechnol Lett 22:789–794
Solaiman DKY, Swingle BM (2010) Isolation of novel Pseudomonas syringae promoters and functional characterization in polyhydroxyalkanoate-producing pseudomonads. New Biotechnol 27:1–9
Solaiman DKY, Ashby RD, Nuñez A, Foglia TA (2004) Production of sophorolipids by Candida bombicola grown on soy molasses as substrate. Biotechnol Lett 26:1241–1245
Solaiman DKY, Ashby RD, Licciardello G, Catara V (2008) Genetic organization of pha gene locus affects phaC expression, poly(hydroxyalkanoate) composition and granule morphology in Pseudomonas corrugata. J Ind Microbiol Biotechnol 35:111–120
Solaiman DKY, Ashby RD, Crocker N, Lai B-H, Zerkowski JA (2014) Rhamnolipid and poly(hydroxyalkanoate) biosynthesis in 3-hydroxyacyl-ACP:CoA transacylase (phaG)-knockouts of Pseudomonas chlororaphis. Biocatalysis Agric Biotechnol 3:159–166
Sotirova AV, Spasova DI, Galabova DN, Karpenko E, Shulga A (2008) Rhamnolipid-biosurfactant permeabilizing effects on Gram-positive and Gram-negative bacterial strains. Curr Microbiol 56:639–644
Stipcevic T, Piljac A, Piljac G (2006) Enhanced healing of full-thickness burn wounds using di-rhamnolipid. Burns 32:24–34
Takemoto JY, Bensaci M, De Lucca AJ, Cleveland TE, Gandhi NR, Palmer Skebba V (2010) Inhibition of fungi from diseased grape by syringomycin E-rhamnolipid mixture. Am J Enol Vitic 61:120–124
Tavares LFD, Silva PM, Junqueira M, Mariano DCO, Nogueira FCS, Domont GB, Freire DMG, Neves BC (2013) Characterization of rhamnolipids produced by wild-type and engineered Burkholderia kururiensis. J Appl Microbiol Biotechnol 97:1909–1921
Toribio J, Escalante AE, Soberón-Chávez G (2010) Rhamnolipids: production in bacteria other than Pseudomonas aeruginosa. Eur J Lipid Sci Technol 112:1082–1087
Transparency Market Research (2012) Biosurfactants market - global scenario, raw material and consumption trends, industry analysis, size, share and forecasts, 2011 – 2018
Van Delden C, Iglewski BH (1998) Cell-to-cell signaling and Pseudomonas aeruginosa infections. Emerg Infect Dis 4:551–560
Van Gennip M, Christensen LD, Alhede M, Phipps R, Jensen PO, Christophersen L, Pamp SJ, Moser C, Mikkelsen PJ, Koh AY, Tolker-Nielsen T, Pier GB, HØiby N, Givskov M, Bjarnsholt T (2009) Inactivation of the rhlA gene in Pseudomonas aeruginosa prevents rhamnolipid production, disabling the protection against polymorphonuclear leukocytes. A.P.M.I.S. 117:537-546
Vatsa P, Sanchez L, Clement C, Baillieul F, Dorey S (2010) Rhamnolipid biosurfactants as new players in animal and plant defense against microbes. Int J Mol Sci 11:5095–5108
Wang Q, Fang X, Bai B, Liang X, Shuler PJ, Goddard WA III, Tang Y (2007) Engineering bacteria for production of rhamnolipid as an agent for enhanced oil recovery. Biotechnol Bioeng 98:842–853
Wittgens A, Tiso T, Arndt TT, Wenk P, Hemmerich J, Müller C, Wichmann R, Küpper B, Zwick M, Wilhelm S, Hausmann R, Syldatk C, Rosenau F, Blank LM (2011) Growth independent rhamnolipid production from glucose using the non-pathogenic Pseudomonas putida KT2440. Microb Cell Factories 10: art. no. 80
Yoo DS, Lee BS, Kim EK (2005) Characteristics of microbial biosurfactant as an antifungal agent against plant pathogenic fungus. J Microbiol Biotechnol 15:1164–1169
Zhu K, Rock CO (2008) RhlA converts β-hydroxyacyl-acyl carrier protein intermediates in fatty acid synthesis to the β-hydroxydecanoyl-β-hydroxydecanoate component of rhamnolipids in Pseudomonas aeruginosa. J Bacteriol 190:3147–3154
Acknowledgments
The authors thank Bun-Hong Lai and Nicole Crocker for providing technical assistance. Nucleic acid sequence determination was performed by David S. Needleman, PhD. and Sue Lawlor of Eastern Regional Research Center Core Technologies’ Integrated Biomolecular Resources group. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture. USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Solaiman, D.K.Y., Ashby, R.D., Gunther, N.W. et al. Dirhamnose-lipid production by recombinant nonpathogenic bacterium Pseudomonas chlororaphis . Appl Microbiol Biotechnol 99, 4333–4342 (2015). https://doi.org/10.1007/s00253-015-6433-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6433-4