Abstract
Purpose
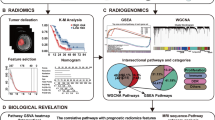
PTEN mutation status is a pivotal biomarker for glioblastoma. This study aimed to establish a radiomic signature to predict PTEN mutation status in patients with glioblastoma, and to investigate the genetic background behind this radiomic signature.
Methods
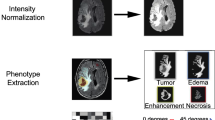
In this study, a total of 862 radiomic features were extracted from each patient. The training (n = 69) and validation (n = 40) sets were retrospectively collected from the Cancer Genome Atlas and the Chinese Glioma Genome Atlas, respectively. The minimum redundancy maximum relevance (mRMR) algorithm was used to select the best predictive features of PTEN status. A machine learning model was then built with the selected features using a support vector machine classifier. The predictive performance of each selected feature and the complete model were evaluated via the area under the curve from receiver operating characteristic analysis in both the training and validation sets. The genetic background underlying the radiomic signature was determined using radiogenomic analysis.
Results
Six features were selected using the mRMR algorithm, including two features derived from contrast-enhanced images and four features derived from T2-weighted images. The predictive performance of the machine learning model for the training and validation sets were 0.925 and 0.787, respectively, which were better than the individual features. Radiogenomics analysis revealed that the PTEN-associated biological processes could be described using the radiomic signature.
Conclusion
These results show that radiomic features derived from preoperative MRI can predict PTEN mutation status in glioblastoma patients, thus providing a novel noninvasive imaging biomarker.




Similar content being viewed by others
Abbreviations
- PTEN:
-
Phosphatase and tensin homolog
- mRMR:
-
Minimum redundancy maximum relevance
- CE:
-
Contrast enhancement
- T2:
-
T2-weighted
- MR:
-
Magnetic resonance
- TCGA:
-
The Cancer Genome Atlas
- AUC:
-
Area under the curve
- SVM:
-
Support vector machine
- ROC:
-
Receiver operating characteristic
- DAVID:
-
Database for Annotation, Visualization and Integrated Discovery
References
Korfiatis P, Kline TL, Coufalova L, Lachance DH, Parney IF, Carter RE, Buckner JC, Erickson BJ (2016) MRI texture features as biomarkers to predict MGMT methylation status in glioblastomas. Med Phys 43(6):2835–2844. https://doi.org/10.1118/1.4948668
Chai RC, Liu YQ, Zhang KN, Wu F, Zhao Z, Wang KY, Jiang T, Wang YZ (2019) A novel analytical model of MGMT methylation pyrosequencing offers improved predictive performance in patients with gliomas. Mod Pathol 32(1):4–15. https://doi.org/10.1038/s41379-018-0143-2
Jiang T, Mao Y, Ma W, Mao Q, You Y, Yang X, Jiang C, Kang C, Li X, Chen L, Qiu X, Wang W, Li W, Yao Y, Li S, Li S, Wu A, Sai K, Bai H, Li G, Chen B, Yao K, Wei X, Liu X, Zhang Z, Dai Y, Lv S, Wang L, Lin Z, Dong J, Xu G, Ma X, Cai J, Zhang W, Wang H, Chen L, Zhang C, Yang P, Yan W, Liu Z, Hu H, Chen J, Liu Y, Yang Y, Wang Z, Wang Z, Wang Y, You G, Han L, Bao Z, Liu Y, Wang Y, Fan X, Liu S, Liu X, Wang Y, Wang Q, Chinese Glioma Cooperative G (2016) CGCG clinical practice guidelines for the management of adult diffuse gliomas. Cancer Lett 375(2):263–273. https://doi.org/10.1016/j.canlet.2016.01.024
Katsetos CD, Draberova E, Smejkalova B, Reddy G, Bertrand L, de Chadarevian JP, Legido A, Nissanov J, Baas PW, Draber P (2007) Class III beta-tubulin and gamma-tubulin are co-expressed and form complexes in human glioblastoma cells. Neurochem Res 32(8):1387–1398. https://doi.org/10.1007/s11064-007-9321-1
Koul D (2008) PTEN signaling pathways in glioblastoma. Cancer Biol Ther 7(9):1321–1325
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114(2):97–109. https://doi.org/10.1007/s00401-007-0243-4
Myung JK, Cho HJ, Park C-K, Kim S-K, Phi JH, Park S-H (2012) IDH1 mutation of gliomas with long-term survival analysis. Oncol Rep 28(5):1639–1644. https://doi.org/10.3892/or.2012.1994
Yang Y, Shao N, Luo G, Li L, Zheng L, Nilsson-Ehle P, Xu N (2010) Mutations of PTEN gene in gliomas correlate to tumor differentiation and short-term survival rate. Anticancer Res 30(3):981–985
Benitez JA, Ma J, D'Antonio M, Boyer A, Camargo MF, Zanca C, Kelly S, Khodadadi-Jamayran A, Jameson NM, Andersen M, Miletic H, Saberi S, Frazer KA, Cavenee WK, Furnari FB (2017) PTEN regulates glioblastoma oncogenesis through chromatin-associated complexes of DAXX and histone H3.3. Nat Commun 8:15223. https://doi.org/10.1038/ncomms15223
Xu PF, Yang JA, Liu JH, Yang X, Liao JM, Yuan FE, Liu BH, Chen QX (2019) PI3Kbeta inhibitor AZD6482 exerts antiproliferative activity and induces apoptosis in human glioblastoma cells. Oncol Rep 41(1):125–132. https://doi.org/10.3892/or.2018.6845
Wu DM, Hong XW, Wen X, Han XR, Wang S, Wang YJ, Shen M, Fan SH, Zhuang J, Zhang ZF, Shan Q, Li MQ, Hu B, Sun CH, Lu J, Zheng YL (2019) MCL1 gene silencing promotes senescence and apoptosis of glioma cells via inhibition of the PI3K/Akt signaling pathway. IUBMB Life 71(1):81–92. https://doi.org/10.1002/iub.1944
Gu J-J, Fan K-C, Zhang J-H, Chen H-J, Wang S-S (2017) Suppression of microRNA-130b inhibits glioma cell proliferation and invasion, and induces apoptosis by PTEN/AKT signaling. Int J Mol Med. https://doi.org/10.3892/ijmm.2017.3233
Karsy M, Neil JA, Guan J, Mahan MA, Colman H, Jensen RL (2015) A practical review of prognostic correlations of molecular biomarkers in glioblastoma. Neurosurg Focus 38(3):E4. https://doi.org/10.3171/2015.1.FOCUS14755
Nakamura JL, Karlsson A, Arvold ND, Gottschalk AR, Pieper RO, Stokoe D, Haas-Kogan DA (2005) PKB/Akt mediates radiosensitization by the signaling inhibitor LY294002 in human malignant gliomas. J Neuro-Oncol 71(3):215–222. https://doi.org/10.1007/s11060-004-1718-y
Abounader R (2009) Interactions between PTEN and receptor tyrosine kinase pathways and their implications for glioma therapy. Expert Rev Anticancer Ther 9(2):235–245. https://doi.org/10.1586/14737140.9.2.235
Luo S, Lei K, Xiang D, Ye K (2018) NQO1 is regulated by PTEN in glioblastoma, mediating cell proliferation and oxidative stress. Oxidative Med Cell Longev 2018:9146528. https://doi.org/10.1155/2018/9146528
Wang Y, Fan X, Zhang C, Zhang T, Peng X, Qian T, Ma J, Wang L, Li S, Jiang T (2014) Identifying radiographic specificity for phosphatase and tensin homolog and epidermal growth factor receptor changes: a quantitative analysis of glioblastomas. Neuroradiology 56(12):1113–1120. https://doi.org/10.1007/s00234-014-1427-y
Ryoo I, Choi SH, Kim JH, Sohn CH, Kim SC, Shin HS, Yeom JA, Jung SC, Lee AL, Yun TJ, Park CK, Park SH (2013) Cerebral blood volume calculated by dynamic susceptibility contrast-enhanced perfusion MR imaging: preliminary correlation study with glioblastoma genetic profiles. PLoS One 8(8):e71704. https://doi.org/10.1371/journal.pone.0071704
Li Y, Ji F, Jiang Y, Zhao T, Xu C (2018) Correlation analysis of expressions of PTEN and p53 with the value obtained by magnetic resonance spectroscopy and apparent diffusion coefficient in the tumor and the tumor-adjacent area in magnetic resonance imaging for glioblastoma. J BUON 23(2):391–397
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278(2):563–577. https://doi.org/10.1148/radiol.2015151169
Zhang B, Chang K, Ramkissoon S, Tanguturi S, Bi WL, Reardon DA, Ligon KL, Alexander BM, Wen PY, Huang RY (2016) Multimodal MRI features predict isocitrate dehydrogenase genotype in high-grade gliomas. Neuro-oncology:now121. https://doi.org/10.1093/neuonc/now121
Li Y, Liu X, Xu K, Qian Z, Wang K, Fan X, Li S, Wang Y, Jiang T (2017) MRI features can predict EGFR expression in lower grade gliomas: a voxel-based radiomic analysis. Eur Radiol 28:356–362. https://doi.org/10.1007/s00330-017-4964-z
Kickingereder P, Bonekamp D, Nowosielski M, Kratz A, Sill M, Burth S, Wick A, Eidel O, Schlemmer HP, Radbruch A, Debus J, Herold-Mende C, Unterberg A, Jones D, Pfister S, Wick W, von Deimling A, Bendszus M, Capper D (2016) Radiogenomics of glioblastoma: machine learning-based classification of molecular characteristics by using multiparametric and multiregional MR imaging features. Radiology 281(3):907–918. https://doi.org/10.1148/radiol.2016161382
Zinn PO, Singh SK, Kotrotsou A, Abrol S, Thomas G, Mosley J, Elakkad A, Hassan I, Kumar A, Colen RR (2017) Distinct radiomic phenotypes define glioblastoma TP53-PTEN-EGFR mutational landscape. Neurosurgery 64(CN_suppl_1):203–210. https://doi.org/10.1093/neuros/nyx316
Hu LS, Ning S, Eschbacher JM, Baxter LC, Gaw N, Ranjbar S, Plasencia J, Dueck AC, Peng S, Smith KA, Nakaji P, Karis JP, Quarles CC, Wu T, Loftus JC, Jenkins RB, Sicotte H, Kollmeyer TM, O'Neill BP, Elmquist W, Hoxworth JM, Frakes D, Sarkaria J, Swanson KR, Tran NL, Li J, Mitchell JR (2016) Radiogenomics to characterize regional genetic heterogeneity in glioblastoma. Neuro-oncology. 19:128–137. https://doi.org/10.1093/neuonc/now135
McCann SM, Jiang Y, Fan X, Wang J, Antic T, Prior F, VanderWeele D, Oto A (2016) Quantitative multiparametric MRI features and PTEN expression of peripheral zone prostate cancer: a pilot study. Am J Roentgenol 206(3):559–565. https://doi.org/10.2214/ajr.15.14967
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Aerts HJ, Velazquez ER, Leijenaar RT, Parmar C, Grossmann P, Carvalho S, Bussink J, Monshouwer R, Haibe-Kains B, Rietveld D, Hoebers F, Rietbergen MM, Leemans CR, Dekker A, Quackenbush J, Gillies RJ, Lambin P (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun 5:4006. https://doi.org/10.1038/ncomms5006
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ, Berman SH, Beroukhim R, Bernard B, Wu CJ, Genovese G, Shmulevich I, Barnholtz-Sloan J, Zou L, Vegesna R, Shukla SA, Ciriello G, Yung WK, Zhang W, Sougnez C, Mikkelsen T, Aldape K, Bigner DD, Van Meir EG, Prados M, Sloan A, Black KL, Eschbacher J, Finocchiaro G, Friedman W, Andrews DW, Guha A, Iacocca M, O'Neill BP, Foltz G, Myers J, Weisenberger DJ, Penny R, Kucherlapati R, Perou CM, Hayes DN, Gibbs R, Marra M, Mills GB, Lander E, Spellman P, Wilson R, Sander C, Weinstein J, Meyerson M, Gabriel S, Laird PW, Haussler D, Getz G, Chin L, Network TR (2013) The somatic genomic landscape of glioblastoma. Cell 155(2):462–477. https://doi.org/10.1016/j.cell.2013.09.034
De Jay N, Papillon-Cavanagh S, Olsen C, El-Hachem N, Bontempi G, Haibe-Kains B (2013) mRMRe: an R package for parallelized mRMR ensemble feature selection. Bioinformatics 29(18):2365–2368. https://doi.org/10.1093/bioinformatics/btt383
Prasanna P, Patel J, Partovi S, Madabhushi A, Tiwari P (2016) Radiomic features from the peritumoral brain parenchyma on treatment-naïve multi-parametric MR imaging predict long versus short-term survival in glioblastoma multiforme: preliminary findings. Eur Radiol 27:4188–4197. https://doi.org/10.1007/s00330-016-4637-3
Orru G, Pettersson-Yeo W, Marquand AF, Sartori G, Mechelli A (2012) Using support vector machine to identify imaging biomarkers of neurological and psychiatric disease: a critical review. Neurosci Biobehav Rev 36(4):1140–1152. https://doi.org/10.1016/j.neubiorev.2012.01.004
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57. https://doi.org/10.1038/nprot.2008.211
Huang d W, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37(1):1–13. https://doi.org/10.1093/nar/gkn923
Aerts HJ, Grossmann P, Tan Y, Oxnard GG, Rizvi N, Schwartz LH, Zhao B (2016) Defining a radiomic response phenotype: a pilot study using targeted therapy in NSCLC. Sci Rep 6:33860. https://doi.org/10.1038/srep33860
Lehrer M, Bhadra A, Ravikumar V, Chen JY, Wintermark M, Hwang SN, Holder CA, Huang EP, Fevrier-Sullivan B, Freymann JB, Rao A, Group TGPR (2017) Multiple-response regression analysis links magnetic resonance imaging features to de-regulated protein expression and pathway activity in lower grade glioma. Oncoscience 4(5–6):57–66. https://doi.org/10.18632/oncoscience.353
Kini V, Chavez A, Mehta D (2010) A new role for PTEN in regulating transient receptor potential canonical channel 6-mediated Ca2+ entry, endothelial permeability, and angiogenesis. J Biol Chem 285(43):33082–33091. https://doi.org/10.1074/jbc.M110.142034
Ng F, Ganeshan B, Kozarski R, Miles KA, Goh V (2013) Assessment of primary colorectal cancer heterogeneity by using whole-tumor texture analysis: contrast-enhanced CT texture as a biomarker of 5-year survival. Radiology 266(1):177–184. https://doi.org/10.1148/radiol.12120254
Yuan F, Zhang Y-H, Kong X-Y, Cai Y-D (2017) Identification of candidate genes related to inflammatory bowel disease using minimum redundancy maximum relevance, incremental feature selection, and the shortest-path approach. Biomed Res Int 2017:1–15. https://doi.org/10.1155/2017/5741948
Xu Y, Ding Y-X, Ding J, Wu L-Y, Xue Y (2016) Mal-Lys: prediction of lysine malonylation sites in proteins integrated sequence-based features with mRMR feature selection. Sci Rep 6(1). https://doi.org/10.1038/srep38318
Mortazavi A, Moattar MH (2016) Robust feature selection from microarray data based on cooperative game theory and qualitative mutual information. Adv Bioinforma 2016:1–16. https://doi.org/10.1155/2016/1058305
Huang MW, Chen CW, Lin WC, Ke SW, Tsai CF (2017) SVM and SVM ensembles in breast Cancer prediction. PLoS One 12(1):e0161501. https://doi.org/10.1371/journal.pone.0161501
Funding
This study was funded by the National Natural Science Foundation of China (No. 81601452), the Beijing Natural Science Foundation (No. 7174295), the National Key Research and Development Plan (No. 2016YFC0902500) and the National Key Research and Development Program of China (2018YFC0115604).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in the studies involving human participants were in accordance with the ethical standards of the institutional research committee (Clinical Research Adoption Committee) and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Table 1
(XLSX 12 kb)
Supplementary Table 2
(DOCX 23 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Liang, Y., Sun, Z. et al. Radiogenomic analysis of PTEN mutation in glioblastoma using preoperative multi-parametric magnetic resonance imaging. Neuroradiology 61, 1229–1237 (2019). https://doi.org/10.1007/s00234-019-02244-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00234-019-02244-7