Abstract
MS-based metabolite profiling of adherent mammalian cells comprises several challenging steps such as metabolism quenching, cell detachment, cell disruption, metabolome extraction, and metabolite measurement. In LC-MS, the final metabolome coverage is strongly determined by the separation technique and the MS conditions used. Human liver-derived cell line HepG2 was chosen as adherent mammalian cell model to evaluate the performance of several commonly used procedures in both sample processing and LC-MS analysis. In a first phase, metabolite extraction and sample analysis were optimized in a combined manner. To this end, the extraction abilities of five different solvents (or combinations) were assessed by comparing the number and the levels of the metabolites comprised in each extract. Three different chromatographic methods were selected for metabolites separation. A HILIC-based method which was set to specifically separate polar metabolites and two RP-based methods focused on lipidome and wide-ranging metabolite detection, respectively. With regard to metabolite measurement, a Q-ToF instrument operating in both ESI (+) and ESI (−) was used for unbiased extract analysis. Once metabolite extraction and analysis conditions were set up, the influence of cell harvesting on metabolome coverage was also evaluated. Therefore, different protocols for cell detachment (trypsinization or scraping) and metabolism quenching were compared. This study confirmed the inconvenience of trypsinization as a harvesting technique, and the importance of using complementary extraction solvents to extend metabolome coverage, minimizing interferences and maximizing detection, thanks to the use of dedicated analytical conditions through the combination of HILIC and RP separations. The proposed workflow allowed the detection of over 300 identified metabolites from highly polar compounds to a wide range of lipids.

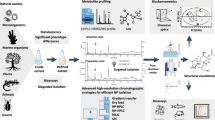
A novel analytical workflow for the LC-MS-based metabolomic analysis of adherent cultured hepatic cells was developed. Three key steps were evaluated: a) cell harvesting, which includes cell detachment and metabolism quenching; b) metabolite extraction; and c) LC-MS analysis, assessing the use of RP and HILIC chromatographies. The final protocol allowed us to extend the metabolome coverage and enabled the detection of a wide range of metabolites from highly polar compounds to a wide range of lipids






Similar content being viewed by others
References
Bai J, Wang MX, Chowbay B, Ching CB, Chen WN (2011) Metabolic profiling of HepG2 cells incubated with S(−) and R(+) enantiomers of anti-coagulating drug warfarin. Metabolomics 7(3):353–362
Croixmarie V, Umbdenstock T, Cloarec O, Moreau A, Pascussi JM, Boursier-Neyret C et al (2009) Integrated comparison of drug-related and drug-induced ultra performance liquid chromatography/mass spectrometry metabonomic profiles using human hepatocyte cultures. Anal Chem 81(15):6061–6069
Brown MV, Compton SA, Milburn MV, Lawton KA, Cheatham B (2013) Metabolomic signatures in lipid-loaded HepaRGs reveal pathways involved in steatotic progression. Obesity (Silver Spring) 21(12):E561–E570
Paglia G, Hrafnsdottir S, Magnusdottir M, Fleming RM, Thorlacius S, Palsson BO et al (2012) Monitoring metabolites consumption and secretion in cultured cells using ultra-performance liquid chromatography quadrupole-time of flight mass spectrometry (UPLC-Q-ToF-MS). Anal Bioanal Chem 402(3):1183–1198
Panopoulos AD, Yanes O, Ruiz S, Kida YS, Diep D, Tautenhahn R et al (2012) The metabolome of induced pluripotent stem cells reveals metabolic changes occurring in somatic cell reprogramming. Cell Res 22(1):168–177
Frezza C, Zheng L, Tennant DA, Papkovsky DB, Hedley BA, Kalna G et al (2011) Metabolic profiling of hypoxic cells revealed a catabolic signature required for cell survival. PLoS One 6(9):e24411
Yizhak K, Gaude E, Le Devedec S, Waldman YY, Stein GY, van de Water B et al (2014) Phenotype-based cell-specific metabolic modeling reveals metabolic liabilities of cancer. eLife 3:e03641
Ibanez C, Valdes A, Garcia-Canas V, Simo C, Celebier M, Rocamora-Reverte L et al (2012) Global Foodomics strategy to investigate the health benefits of dietary constituents. J Chromatogr A 1248:139–153
Leon Z, Garcia-Canaveras JC, Donato MT, Lahoz A (2013) Mammalian cell metabolomics: experimental design and sample preparation. Electrophoresis 34(19):2762–2775
Gomez-Lechon MJ, Castell JV, Donato MT (2008) An update on metabolism studies using human hepatocytes in primary culture. Expert Opin Drug metab Toxicol 4(7):837–854
Vuckovic D (2012) Current trends and challenges in sample preparation for global metabolomics using liquid chromatography-mass spectrometry. Anal Bioanal Chem 403(6):1523–1548
Dettmer K, Nurnberger N, Kaspar H, Gruber MA, Almstetter MF, Oefner PJ (2011) Metabolite extraction from adherently growing mammalian cells for metabolomics studies: optimization of harvesting and extraction protocols. Anal Bioanal Chem 399(3):1127–1139
Fei F, Bowdish DM, McCarry BE (2014) Comprehensive and simultaneous coverage of lipid and polar metabolites for endogenous cellular metabolomics using HILIC-TOF-MS. Anal Bioanal Chem 406(15):3723–3733
Lorenz MA, Burant CF, Kennedy RT (2011) Reducing time and increasing sensitivity in sample preparation for adherent mammalian cell metabolomics. Anal Chem 83(9):3406–3414
Bi H, Krausz KW, Manna SK, Li F, Johnson CH, Gonzalez FJ (2013) Optimization of harvesting, extraction, and analytical protocols for UPLC-ESI-MS-based metabolomic analysis of adherent mammalian cancer cells. Anal Bioanal Chem 405(15):5279–5289
Ritter JB, Genzel Y, Reichl U (2008) Simultaneous extraction of several metabolites of energy metabolism and related substances in mammalian cells: optimization using experimental design. Anal Biochem 373(2):349–369
Ivanisevic J, Zhu ZJ, Plate L, Tautenhahn R, Chen S, O’Brien PJ et al (2013) Toward ‘omic scale metabolite profiling: a dual separation-mass spectrometry approach for coverage of lipid and central carbon metabolism. Anal Chem 85(14):6876–6884
Yuan M, Breitkopf SB, Yang X, Asara JM (2012) A positive/negative ion-switching, targeted mass spectrometry-based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat Protoc 7(5):872–881
Martano G, Delmotte N, Kiefer P, Christen P, Kentner D, Bumann D et al (2015) Fast sampling method for mammalian cell metabolic analyses using liquid chromatography-mass spectrometry. Nat Protoc 10(1):1–11
Sellick CA, Hansen R, Stephens GM, Goodacre R, Dickson AJ (2011) Metabolite extraction from suspension-cultured mammalian cells for global metabolite profiling. Nat Protoc 6(8):1241–1249
Lahoz A, Gombau L, Donato MT, Castell JV, Gomez-Lechon MJ (2006) In vitro ADME medium/high-throughput screening in drug preclinical development. Mini Rev Med Chem 6(9):1053–1062
Lenz EM, Wilson ID (2007) Analytical strategies in metabonomics. J Proteome Res 6(2):443–458
Garcia-Cañaveras JC, Donato MT, Castell JV, Lahoz A (2011) A comprehensive untargeted metabonomic analysis of human steatotic liver tissue by RP and HILIC chromatography coupled to mass spectrometry reveals important metabolic alterations. J Proteome Res 10(10):4825–4834
Rojo D, Barbas C, Ruperez FJ (2012) LC-MS metabolomics of polar compounds. Bioanalysis 4(10):1235–1243
Donato MT, Lahoz A, Castell JV, Gomez-Lechon MJ (2008) Cell lines: a tool for in vitro drug metabolism studies. Curr Drug Metab 9(1):1–11
Gomez-Lechon MJ, Donato MT, Martinez-Romero A, Jimenez N, Castell JV, O’Connor JE (2007) A human hepatocellular in vitro model to investigate steatosis. Chem Biol Interact 165(2):106–116
García-Cañaveras JC, Jiménez N, Gómez-Lechón MJ, Castell JV, Donato MT, Lahoz A (2015) LC-MS untargeted metabolomic analysis of drug-induced hepatotoxicity in HepG2 cells. Electrophoresis 36(18):2294–2302
Gomez-Lechon MJ, Ponsoda X, O’Connor E, Donato T, Jover R, Castell JV (2003) Diclofenac induces apoptosis in hepatocytes. Toxicol in Vitro 17(5–6):675–680
Cao B, Aa J, Wang G, Wu X, Liu L, Li M et al (2011) GC–TOFMS analysis of metabolites in adherent MDCK cells and a novel strategy for identifying intracellular metabolic markers for use as cell amount indicators in data normalization. Anal Bioanal Chem 400(9):2983–2993
Wu H, Southam AD, Hines A, Viant MR (2008) High-throughput tissue extraction protocol for NMR- and MS-based metabolomics. Anal Biochem 372(2):204–212
Want EJ, Masson P, Michopoulos F, Wilson ID, Theodoridis G, Plumb RS et al (2013) Global metabolic profiling of animal and human tissues via UPLC-MS. Nat Protoc 8(1):17–32
Nygren H, Seppanen-Laakso T, Castillo S, Hyotylainen T, Oresic M (2011) Liquid chromatography-mass spectrometry (LC-MS)-based lipidomics for studies of body fluids and tissues. Methods Mol Biol 708:247–257
Cortes M, Pareja E, Garcia-Canaveras JC, Donato MT, Montero S, Mir J et al (2014) Metabolomics discloses donor liver biomarkers associated with early allograft dysfunction. J Hepatol 61(3):564–574
Pluskal T, Castillo S, Villar-Briones A, Oresic M (2010) MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinf 11:395
Quintás G, Portillo N, García-Cañaveras JC, Castell JV, Ferrer A, Lahoz A (2012) Chemometric approaches to improve PLSDA model outcome for predicting human non-alcoholic fatty liver disease using UPLC-MS as a metabolic profiling tool. Metabolomics 8(1):86–98
Wishart DS, Jewison T, Guo AC, Wilson M, Knox C, Liu Y et al (2013) HMDB 3.0—the human metabolome database in 2013. Nucleic Acids Res 41(Database issue):D801–D807
Fahy E, Sud M, Cotter D, Subramaniam S (2007) LIPID MAPS online tools for lipid research. Nucleic Acids Res 35(Web Server issue):W606–W612
Smith CA, O’Maille G, Want EJ, Qin C, Trauger SA, Brandon TR et al (2005) METLIN: a metabolite mass spectral database. Ther Drug Monit 27(6):747–751
Horai H, Arita M, Kanaya S, Nihei Y, Ikeda T, Suwa K et al (2010) MassBank: a public repository for sharing mass spectral data for life sciences. J Mass Spectrom 45(7):703–714
Sumner LW, Amberg A, Barrett D, Beale MH, Beger R, Daykin CA et al (2007) Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 3(3):211–221
R Core Team (2014) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Álvarez-Sánchez B, Priego-Capote F, Castro M (2010) Metabolomics analysis II. Preparation of biological samples prior to detection. TrAC Trends Anal Chem 29(2):120–127
Villas-Boas SG, Mas S, Akesson M, Smedsgaard J, Nielsen J (2005) Mass spectrometry in metabolome analysis. Mass Spectrom Rev 24(5):613–646
Seppanen-Laakso T, Oresic M (2009) How to study lipidomes. J Mol Endocrinol 42(3):185–190
Dunn WB, Broadhurst D, Begley P, Zelena E, Francis-McIntyre S, Anderson N et al (2011) Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nat Protoc 6(7):1060–1083
Chen J, Zhou L, Zhang X, Lu X, Cao R, Xu C et al (2012) Urinary hydrophilic and hydrophobic metabolic profiling based on liquid chromatography-mass spectrometry methods: differential metabolite discovery specific to ovarian cancer. Electrophoresis 33(22):3361–3369
Teng Q, Huang W, Collette TW, Ekman DR, Tan C (2009) A direct cell quenching method for cell-culture based metabolomics. Metabolomics 5(2):199–208
Atkinson DE (1968) The energy charge of the adenylate pool as a regulatory parameter. Interaction with feedback modifiers. Biochemistry 7(11):4030–4034
Saric J, Want EJ, Duthaler U, Lewis M, Keiser J, Shockcor JP et al (2012) Systematic evaluation of extraction methods for multiplatform-based metabotyping: application to the Fasciola hepatica metabolome. Anal Chem 84(16):6963–6972
Vorkas PA, Isaac G, Anwar MA, Davies AH, Want EJ, Nicholson JK et al (2015) Untargeted UPLC-MS profiling pipeline to expand tissue metabolome coverage: application to cardiovascular disease. Anal Chem 87(8):4184–4193
Yamazaki M, Miyake M, Sato H, Masutomi N, Tsutsui N, Adam K-P et al (2013) Perturbation of bile acid homeostasis is an early pathogenesis event of drug induced liver injury in rats. Toxicol Appl Pharmacol 268(1):79–89
Sarafian MH, Gaudin M, Lewis MR, Martin FP, Holmes E, Nicholson JK et al (2014) Objective set of criteria for optimization of sample preparation procedures for ultra-high throughput untargeted blood plasma lipid profiling by ultra performance liquid chromatography-mass spectrometry. Anal Chem 86(12):5766–5774
Acknowledgments
This work has been supported by the Instituto de Salud Carlos III of the Spanish Ministry of Science and Innovation (FIS PI14/00026 and FIS PI13/0986). A.L. is grateful for a Miguel Server II contract (CPII14/00004) from the above Ministry/Instituto de Salud Carlos III. J.C. G.-C. is grateful for a pre-doctoral contract from the Vali + d program of the Conselleria d’Educació (Regional Valencian Ministry of Education). S.L. is grateful for a contract (PTA2012-7224-I) from the Spanish Ministry of Economy and Competitiveness.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
All the animals received human care and all experimental protocols were approved by the institutional animal ethics committee and performed in accordance with national and institutional regulations.
Conflict of interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 2.28 mb)
Rights and permissions
About this article
Cite this article
García-Cañaveras, J.C., López, S., Castell, J.V. et al. Extending metabolome coverage for untargeted metabolite profiling of adherent cultured hepatic cells. Anal Bioanal Chem 408, 1217–1230 (2016). https://doi.org/10.1007/s00216-015-9227-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-015-9227-8