Abstract
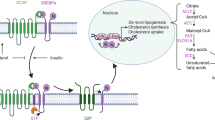
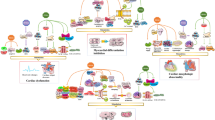
The mechanisms by which pollutants participate in the development of diverse pathologies are not completely understood. The pollutant 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) activates the AhR (aryl hydrocarbon receptor) signaling pathway. We previously showed that TCDD (25 nM, 30 h) decreased the expression of several alcohol metabolism enzymes (cytochrome P450 2E1, alcohol dehydrogenases ADH1, 4 and 6) in differentiated human hepatic cells (HepaRG). Here, we show that, as rapidly as 8 h after treatment (25 nM TCDD) ADH expression decreased 40 % (p < 0.05). ADH1 and 4 protein levels decreased 40 and 27 %, respectively (p < 0.05), after 72 h (25 nM TCDD). The protein half-lives were not modified by TCDD which suggests transcriptional regulation of expression. The AhR antagonist CH-223191 or AhR siRNA reduced the inhibitory effect of 25 nM TCDD on ADH1A, 4 and 6 expression 50–100 % (p < 0.05). The genomic pathway (via the AhR/ARNT complex) and not the non-genomic pathway involving c-SRC mediated these effects. Other AhR ligands (3-methylcholanthrene and PCB 126) decreased ADH1B, 4 and 6 mRNAs by more than 78 and 55 %, respectively (p < 0.01). TCDD also regulated the expression of ADH4 in the HepG2 human hepatic cell line, in primary human hepatocytes and in C57BL/6J mouse liver. In conclusion, activation of the AhR/ARNT signaling pathway by AhR ligands represents a novel mechanism for regulating the expression of ADHs. These effects may be implicated in the toxicity of AhR ligands as well as in the alteration of ethanol or retinol metabolism and may be associated further with higher risk of liver diseases or/and alcohol abuse disorders.






Similar content being viewed by others
References
Alavanja MCR, Dosemeci M, Samanic C et al (2004) Pesticides and lung cancer risk in the agricultural health study cohort. Am J Epidemiol 160:876–885. doi:10.1093/aje/kwh290
Ambolet-Camoit A, Ottolenghi C, Leblanc A et al (2015) Two persistent organic pollutants which act through different xenosensors (alpha-endosulfan and 2,3,7,8 tetrachlorodibenzo-p-dioxin) interact in a mixture and downregulate multiple genes involved in human hepatocyte lipid and glucose metabolism. Biochimie 116:79–91. doi:10.1016/j.biochi.2015.07.003
Aninat C, Piton A, Glaise D et al (2006) Expression of cytochromes P450, conjugating enzymes and nuclear receptors in human hepatoma HepaRG cells. Drug Metab Dispos Biol Fate Chem 34:75–83. doi:10.1124/dmd.105.006759
Barouki R, Coumoul X, Fernandez-Salguero PM (2007) The aryl hydrocarbon receptor, more than a xenobiotic-interacting protein. FEBS Lett 581:3608–3615. doi:10.1016/j.febslet.2007.03.046
Bauman JW, Goldsworthy TL, Dunn CS, Fox TR (1995) Inhibitory effects of 2,3,7,8-tetrachlorodibenzo-p-dioxin on rat hepatocyte proliferation induced by 2/3 partial hepatectomy. Cell Prolif 28:437–451
Bhopale KK, Wu H, Boor PJ et al (2006) Metabolic basis of ethanol-induced hepatic and pancreatic injury in hepatic alcohol dehydrogenase deficient deer mice. Alcohol 39:179–188. doi:10.1016/j.alcohol.2006.09.005
Braeuning A, Thomas M, Hofmann U et al (2015) Comparative analysis and functional characterization of HC-AFW1 hepatocarcinoma cells: CYP expression and induction by nuclear receptor agonists. Drug Metab Dispos Biol Fate Chem. doi:10.1124/dmd.115.064667
Bui L-C, Tomkiewicz C, Chevallier A et al (2009) Nedd9/Hef1/Cas-L mediates the effects of environmental pollutants on cell migration and plasticity. Oncogene 28:3642–3651. doi:10.1038/onc.2009.224
Carlson EA, McCulloch C, Koganti A et al (2009) Divergent transcriptomic responses to aryl hydrocarbon receptor agonists between rat and human primary hepatocytes. Toxicol Sci Off J Soc Toxicol 112:257–272. doi:10.1093/toxsci/kfp200
Chang JS, Straif K, Guha N (2012) The role of alcohol dehydrogenase genes in head and neck cancers: a systematic review and meta-analysis of ADH1B and ADH1C. Mutagenesis 27:275–286. doi:10.1093/mutage/ger073
Clissold PM, Hainey S, Bishop JO (1984) Messenger RNAs coding for mouse major urinary proteins are differentially induced by testosterone. Biochem Genet 22:379–387
Dannenberg LO, Chen H-J, Edenberg HJ (2005) GATA-2 and HNF-3beta regulate the human alcohol dehydrogenase 1A (ADH1A) gene. DNA Cell Biol 24:543–552. doi:10.1089/dna.2005.24.543
Denison MS, Soshilov AA, He G et al (2011) Exactly the same but different: promiscuity and diversity in the molecular mechanisms of action of the aryl hydrocarbon (dioxin) receptor. Toxicol Sci Off J Soc Toxicol 124:1–22. doi:10.1093/toxsci/kfr218
Dong Y, Poellinger L, Okret S et al (1988) Regulation of gene expression of class I alcohol dehydrogenase by glucocorticoids. Proc Natl Acad Sci USA 85:767–771
Duester G, Farrés J, Felder MR et al (1999) Recommended nomenclature for the vertebrate alcohol dehydrogenase gene family. Biochem Pharmacol 58:389–395
Edenberg HJ (2000) Regulation of the mammalian alcohol dehydrogenase genes. Prog Nucleic Acid Res Mol Biol 64:295–341
Fader KA, Nault R, Ammendolia DA et al (2015) 2,3,7,8-Tetrachlorodibenzo-p-Dioxin (TCDD) Alters Lipid Metabolism and Depletes Immune Cell Populations in the Jejunum of C57BL/6 Mice. Toxicol Sci Off J Soc Toxicol. doi:10.1093/toxsci/kfv206
Fletcher N, Wahlström D, Lundberg R et al (2005) 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD) alters the mRNA expression of critical genes associated with cholesterol metabolism, bile acid biosynthesis, and bile transport in rat liver: a microarray study. Toxicol Appl Pharmacol 207:1–24. doi:10.1016/j.taap.2004.12.003
Forgacs AL, Dere E, Angrish MM, Zacharewski TR (2013) Comparative analysis of temporal and dose-dependent TCDD-elicited gene expression in human, mouse, and rat primary hepatocytes. Toxicol Sci Off J Soc Toxicol 133:54–66. doi:10.1093/toxsci/kft028
Guyot E, Chevallier A, Barouki R, Coumoul X (2013) The AhR twist: ligand-dependent AhR signaling and pharmaco-toxicological implications. Drug Discov Today 18:479–486. doi:10.1016/j.drudis.2012.11.014
He L, Simmen FA, Ronis MJJ, Badger TM (2004) Post-transcriptional regulation of sterol regulatory element-binding protein-1 by ethanol induces class I alcohol dehydrogenase in rat liver. J Biol Chem 279:28113–28121. doi:10.1074/jbc.M400906200
Hellgren M, Strömberg P, Gallego O et al (2007) Alcohol dehydrogenase 2 is a major hepatic enzyme for human retinol metabolism. Cell Mol Life Sci CMLS 64:498–505. doi:10.1007/s00018-007-6449-8
Hernández-Tobías A, Julián-Sánchez A, Piña E, Riveros-Rosas H (2011) Natural alcohol exposure: is ethanol the main substrate for alcohol dehydrogenases in animals? Chem Biol Interact 191:14–25. doi:10.1016/j.cbi.2011.02.008
Hines RN, McCarver DG (2002) The ontogeny of human drug-metabolizing enzymes: phase I oxidative enzymes. J Pharmacol Exp Ther 300:355–360. doi:10.1124/jpet.300.2.355
Höög J-O, Ostberg LJ (2011) Mammalian alcohol dehydrogenases–a comparative investigation at gene and protein levels. Chem Biol Interact 191:2–7. doi:10.1016/j.cbi.2011.01.028
Huang W, Booth DM, Cane MC et al (2014) Fatty acid ethyl ester synthase inhibition ameliorates ethanol-induced Ca2+-dependent mitochondrial dysfunction and acute pancreatitis. Gut 63:1313–1324. doi:10.1136/gutjnl-2012-304058
Ishii Y, Kato H, Hatsumura M et al (2001) Effects of a highly toxic coplanar polychlorinated biphenyl, 3,3′,4,4′,5-pentachlorobiphenyl on intermediary metabolism: reduced triose phosphate content in rat liver cytosol. Fukuoka Igaku Zasshi Hukuoka Acta Medica 92:190–200
Kaphalia L, Calhoun WJ (2013) Alcoholic lung injury: metabolic, biochemical and immunological aspects. Toxicol Lett 222:171–179. doi:10.1016/j.toxlet.2013.07.016
Kim S-H, Henry EC, Kim D-K et al (2006) Novel compound 2-methyl-2H-pyrazole-3-carboxylic acid (2-methyl-4-o-tolylazo-phenyl)-amide (CH-223191) prevents 2,3,7,8-TCDD-induced toxicity by antagonizing the aryl hydrocarbon receptor. Mol Pharmacol 69:1871–1878. doi:10.1124/mol.105.021832
Kumar S, Sandell LL, Trainor PA et al (2012) Alcohol and aldehyde dehydrogenases: retinoid metabolic effects in mouse knockout models. Biochim Biophys Acta 1821:198–205. doi:10.1016/j.bbalip.2011.04.004
Larsen GL, Bergman A, Klasson-Wehler E (1990) A methylsulphonyl metabolite of a polychlorinated biphenyl can serve as a ligand for alpha 2mu-globulin in rat and major-urinary-protein in mice. Xenobiotica Fate Foreign Compd Biol Syst 20:1343–1352. doi:10.3109/00498259009046632
Lee D-H, Steffes MW, Sjödin A et al (2010) Low dose of some persistent organic pollutants predicts type 2 diabetes: a nested case-control study. Environ Health Perspect 118:1235–1242. doi:10.1289/ehp.0901480
Liu Z, Ma Y, Yang J, Qin H (2011) Upregulated and downregulated proteins in hepatocellular carcinoma: a systematic review of proteomic profiling studies. Omics J Integr Biol 15:61–71. doi:10.1089/omi.2010.0061
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods San Diego Calif 25:402–408. doi:10.1006/meth.2001.1262
Lo R, Matthews J (2012) High-resolution genome-wide mapping of AHR and ARNT binding sites by ChIP-Seq. Toxicol Sci Off J Soc Toxicol 130:349–361. doi:10.1093/toxsci/kfs253
Magne L, Blanc E, Marchand A et al (2007) Stabilization of IGFBP-1 mRNA by ethanol in hepatoma cells involves the JNK pathway. J Hepatol 47:691–698. doi:10.1016/j.jhep.2007.05.018
McCarver DG, Hines RN (2002) The ontogeny of human drug-metabolizing enzymes: phase II conjugation enzymes and regulatory mechanisms. J Pharmacol Exp Ther 300:361–366. doi:10.1124/jpet.300.2.361
Morel Y, de Waziers I, Barouki R (2000) A repressive cross-regulation between catalytic and promoter activities of the CYP1A1 and CYP2E1 genes: role of H(2)O(2). Mol Pharmacol 57:1158–1164
Mutka SC, Green LH, Verderber EL et al (2012) ADH IB expression, but not ADH III, is decreased in human lung cancer. PLoS One 7:e52995. doi:10.1371/journal.pone.0052995
Okada H, Honda M, Campbell JS et al (2012) Acyclic retinoid targets platelet-derived growth factor signaling in the prevention of hepatic fibrosis and hepatocellular carcinoma development. Cancer Res 72:4459–4471. doi:10.1158/0008-5472.CAN-12-0028
Parola M, Robino G (2001) Oxidative stress-related molecules and liver fibrosis. J Hepatol 35:297–306
Pierre S, Chevallier A, Teixeira-Clerc F et al (2014) Aryl hydrocarbon receptor-dependent induction of liver fibrosis by dioxin. Toxicol Sci Off J Soc Toxicol 137:114–124. doi:10.1093/toxsci/kft236
Pochareddy S, Edenberg HJ (2010) Identification of a FOXA-dependent enhancer of human alcohol dehydrogenase 4 (ADH4). Gene 460:1–7. doi:10.1016/j.gene.2010.03.013
Podechard N, Le Ferrec E, Rebillard A et al (2009) NPC1 repression contributes to lipid accumulation in human macrophages exposed to environmental aryl hydrocarbons. Cardiovasc Res 82:361–370. doi:10.1093/cvr/cvp007
Podevin P, Carpentier A, Pène V et al (2010) Production of infectious hepatitis C virus in primary cultures of human adult hepatocytes. Gastroenterology 139:1355–1364. doi:10.1053/j.gastro.2010.06.058
Procházková J, Kabátková M, Šmerdová L et al (2013) Aryl hydrocarbon receptor negatively regulates expression of the plakoglobin gene (jup). Toxicol Sci Off J Soc Toxicol 134:258–270. doi:10.1093/toxsci/kft110
Ramirez MC, Bourguignon NS, Bonaventura MM et al (2012) Neonatal xenoestrogen exposure alters growth hormone-dependent liver proteins and genes in adult female rats. Toxicol Lett 213:325–331. doi:10.1016/j.toxlet.2012.07.015
Rouach H, Fataccioli V, Gentil M et al (1997) Effect of chronic ethanol feeding on lipid peroxidation and protein oxidation in relation to liver pathology. Hepatol 25:351–355. doi:10.1002/hep.510250216
Sebastiani G, Gkouvatsos K, Maffettone C et al (2011) Accelerated CCl4-induced liver fibrosis in Hjv-/-mice, associated with an oxidative burst and precocious profibrogenic gene expression. PLoS One 6:e25138. doi:10.1371/journal.pone.0025138
Sellin S, Holmquist B, Mannervik B, Vallee BL (1991) Oxidation and reduction of 4-hydroxyalkenals catalyzed by isozymes of human alcohol dehydrogenase. Biochemistry 30:2514–2518
Stewart MJ, Shean ML, Paeper BW, Duester G (1991) The role of CCAAT/enhancer-binding protein in the differential transcriptional regulation of a family of human liver alcohol dehydrogenase genes. J Biol Chem 266:11594–11603
Tomkiewicz C, Herry L, Bui L-C et al (2013) The aryl hydrocarbon receptor regulates focal adhesion sites through a non-genomic FAK/Src pathway. Oncogene 32:1811–1820. doi:10.1038/onc.2012.197
Triano EA, Slusher LB, Atkins TA et al (2003) Class I alcohol dehydrogenase is highly expressed in normal human mammary epithelium but not in invasive breast cancer: implications for breast carcinogenesis. Cancer Res 63:3092–3100
Wang F, Samudio I, Safe S (2001) Transcriptional activation of cathepsin D gene expression by 17beta-estradiol: mechanism of aryl hydrocarbon receptor-mediated inhibition. Mol Cell Endocrinol 172:91–103
Wei R-R, Zhang M-Y, Rao H-L et al (2012) Identification of ADH4 as a novel and potential prognostic marker in hepatocellular carcinoma. Med Oncol Northwood Lond Engl 29:2737–2743. doi:10.1007/s12032-011-0126-3
White SS, Birnbaum LS (2009) An overview of the effects of dioxins and dioxin-like compounds on vertebrates, as documented in human and ecological epidemiology. J Environ Sci Health Part C Environ Carcinog Ecotoxicol Rev 27:197–211. doi:10.1080/10590500903310047
Wu H, Cai P, Clemens DL et al (2006) Metabolic basis of ethanol-induced cytotoxicity in recombinant HepG2 cells: role of nonoxidative metabolism. Toxicol Appl Pharmacol 216:238–247. doi:10.1016/j.taap.2006.05.003
Zelner I, Matlow JN, Natekar A, Koren G (2013) Synthesis of fatty acid ethyl esters in mammalian tissues after ethanol exposure: a systematic review of the literature. Drug Metab Rev 45:277–299. doi:10.3109/03602532.2013.795584
Zheng YW, Bey M, Liu H, Felder MR (1993) Molecular basis of the alcohol dehydrogenase-negative deer mouse. Evidence for deletion of the gene for class I enzyme and identification of a possible new enzyme class. J Biol Chem 268:24933–24939
Acknowledgments
This work was supported by the IREB association (Institut de Recherche scientifique sur les Boissons, 2015/03), the FRA (Fondation pour la Recherche en Alcoologie, 2016/02), the ANR (Agence Nationale de la Recherche), and the INCa (Institut National du Cancer, Caroline Duval, postdoctoral fellowship, CESA 201101), and Université Paris Descartes, MESR (Ministère de l’Enseignement Supérieur et de la Recherche) for the doctoral fellowships (Eléonore Attignon, Alix Leblanc). We thank Pr X. Coumoul for critical reading and Dr L. Aggerbeck for editing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Eléonore A. Attignon and Alix F. Leblanc have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
204_2016_1700_MOESM2_ESM.eps
Supplementary material 2 a: The viability of HepaRG cells exposed for 8 days to different concentrations of TCDD was evaluated by the WST-1 test. b: Quantitative real-time PCR analysis of ADH1B and ADH4 mRNA expression after 8 days of exposure to several concentrations of TCDD. * p < 0.05, ** p < 0.01, *** p < 0.001 (EPS 13747 kb)
204_2016_1700_MOESM3_ESM.eps
Supplementary material 3 The half-lives of ADH mRNAs are unchanged after exposure of cells to TCDD. The half-lives of ADH mRNAs, as measured by quantitative real-time PCR, in the presence of DRB, an inhibitor of RNA polymerase II, are shown for HepaRG cells exposed or not (Control) to TCDD. Cells were exposed or not to 25 nM TCDD for 20 h then DRB was added and cells were harvested at 0, 4, 8, 12 and 24 h. The means are shown and the error bars represent the SD of three independent experiments (EPS 5041 kb)
204_2016_1700_MOESM4_ESM.eps
Supplementary material 4 siRNAs reduce the expression of AhR, ARNT and c-SRC mRNAs. Differentiated HepaRG cells were transfected with siRNA directed against AhR, ARNT or SRC or scrambled (siC). Three days after, the cells were exposed or not to 25 nM TCDD for 24 h. The levels of AhR (a) ARNT (b) and c-SRC (c) and CYP1A1 (d) mRNAs were measured by qRT-PCR. Relative mRNA levels were calculated using siC value as the reference. The means are shown and the error bars represent the SD of four independent experiments. ** p < 0.01 *** p < 0.001 as compared to untreated and untransfected cells (EPS 17465 kb)
204_2016_1700_MOESM6_ESM.eps
Supplementary material 6 Values (means and SD) used for Fig. 1b: qRT-PCR measurement of ADH (ADH1A, 1B, 4, 5 and 6) mRNA expression as a function of the concentration of TCDD (0–25 nM for 24 h) (n = 3-4), in differentiated HepaRG cells (EPS 3338 kb)
Rights and permissions
About this article
Cite this article
Attignon, E.A., Leblanc, A.F., Le-Grand, B. et al. Novel roles for AhR and ARNT in the regulation of alcohol dehydrogenases in human hepatic cells. Arch Toxicol 91, 313–324 (2017). https://doi.org/10.1007/s00204-016-1700-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-016-1700-4