Abstract
Key message
Karyotypes evolve through numerical and structural chromosome rearrangements. We show that Phaseolus leptostachyus, a wild bean, underwent a rapid genome reshuffling associated with the reduction from 11 to 10 chromosome pairs, but without whole genome duplication, the highest chromosome evolution rate known for plants.
Abstract
Plant karyotypes evolve through structural rearrangements often associated with polyploidy or dysploidy. The genus Phaseolus comprises ~ 90 species, five of them domesticated due to their nutritional relevance. Most of the species have 2n = 22 karyotypes and are highly syntenic, except for three dysploid karyotypes of species from the Leptostachyus group (2n = 20) that have accumulated several rearrangements. Here, we investigated the degrees of structural rearrangements among Leptostachyus and other Phaseolus groups by estimating their chromosomal evolution rates (CER). For this, we combined our oligo-FISH barcode system for beans and chromosome-specific painting probes for chromosomes 2 and 3, with rDNA and a centromeric probe to establish chromosome orthologies and identify structural rearrangements across nine Phaseolus species. We also integrated the detected rearrangements with a phylogenomic approach to estimate the CERs for each Phaseolus lineage. Our data allowed us to identify translocations, inversions, duplications and deletions, mostly in species belonging to the Leptostachyus group. Phaseolus leptostachyus showed the highest CER (12.31 rearrangements/My), a tenfold increase in contrast to the 2n = 22 species analysed. This is the highest rate known yet for plants, making it a model species for investigating the mechanisms behind rapid genome reshuffling in early species diversification.






Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in the manuscript or as supplementary materials.
References
Almeida C, Pedrosa-Harand A (2011) Contrasting rDNA evolution in lima bean (Phaseolus lunatus L.) and common bean (P. vulgaris L., Fabaceae). Cytogenet Genome Res 132:212–217. https://doi.org/10.1159/000321677
Barker MS, Baute GJ, Liu S-L (2012) Duplications and turnover in plant genomes. In: Wendel JF, Greilhuber J, Dolezel J, Leitch IJ (eds) Plant genome diversity: plant genomes, their residents, and their evolutionary dynamics, vol 1. Springer, Vienna, pp 155–169
Benathar TCM, Nagamachi CY, Rodrigues LRR et al (2019) Karyotype, evolution and phylogenetic reconstruction in Micronycterinae bats with implications for the ancestral karyotype of Phyllostomidae. BMC Evol Biol 19:1–13. https://doi.org/10.1186/s12862-019-1421-4
Bonifácio EM, Fonsêca A, Almeida C et al (2012) Comparative cytogenetic mapping between the lima bean (Phaseolus lunatus L.) and the common bean (P. vulgaris L.). Theor Appl Genet 124:1513–1520. https://doi.org/10.1007/s00122-012-1806-x
Braz GT, He L, Zhao H et al (2018) Comparative oligo-FISH mapping: an efficient and powerful methodology to reveal karyotypic and chromosomal evolution. Genetics 208:513–523. https://doi.org/10.1534/genetics.117.300344
Braz GT, Yu F, Zhao H et al (2021) Preferential meiotic chromosome pairing among homologous chromosomes with cryptic sequence variation in tetraploid maize. New Phytol 229:3294–3302. https://doi.org/10.1111/nph.17098
Braz GT, Yu F, de Vale Martins L, Jiang J (2020) Fluorescent in situ Hybridization using oligonucleotide-based probes. In: In situ hybridization protocols, Methods in Molecular Biology. Humana, New York, pp 71–83 DOI: https://doi.org/10.1007/978-1-0716-0623-0_4
Broughton WJ, Hernández G et al (2003) Beans (Phaseolus spp.)—model food legumes. Plant Soil 252:55–128. https://doi.org/10.1023/A:1024146710611
Danecek P, Bonfield JK, Liddle J et al (2021) Twelve years of SAMtools and BCFtools. GigaScience 10:1–4. https://doi.org/10.1093/gigascience/giab008
De Carvalho CR, Saraiva LS (1993) An air drying technique for maize chromosomes without enzymatic maceration. Biotech Histochem 68:142–145. https://doi.org/10.3109/10520299309104684
De Storme N, Mason A (2014) Plant speciation through chromosome instability and ploidy change: cellular mechanisms, molecular factors and evolutionary relevance. Curr Plant Biol 1:10–33. https://doi.org/10.1016/j.cpb.2014.09.002
De OliveiraBustamante F, de Nascimento TH, Montenegro C et al (2021) Oligo-FISH barcode in beans: a new chromosome identification system. Theor Appl Genetics. 134:3675-3686. https://doi.org/10.1007/s00122-021-03921-z
Delgado Salinas A, Bibler R, Lavin M (2006) Phylogeny of the genus Phaseolus (Leguminosae): a recent diversification in an ancient landscape. Syst Bot 31:779–791. https://doi.org/10.1600/036364406779695960
Do Vale Martins L, Yu F, Zhao H et al (2019) Meiotic crossovers characterized by haplotype-specific chromosome painting in maize. Nat Commun 10:4604. https://doi.org/10.1038/s41467-019-12646-z
Do Vale Martins L, De Oliveira BF, da Silva Oliveira AR et al (2021) BAC- and oligo-FISH mapping reveals chromosome evolution among Vigna angularis, V. unguiculata, and Phaseolus vulgaris. Chromosoma 130:133–147. https://doi.org/10.1007/s00412-021-00758-9
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol 29:1969–1973. https://doi.org/10.1093/molbev/mss075
Ferraz ME, Fonsêca A, Pedrosa-Harand A (2020) Multiple and independent rearrangements revealed by comparative cytogenetic mapping in the dysploid Leptostachyus group (Phaseolus L., Leguminosae). Chromosom Res 28:395–405. https://doi.org/10.1007/s10577-020-09644-z
Fonsêca A, Pedrosa-Harand A (2013) Karyotype stability in the genus Phaseolus evidenced by the comparative mapping of the wild species Phaseolus microcarpus. Genome 56:335–343. https://doi.org/10.1139/gen-2013-0025
Fonsêca A, Pedrosa-Harand A (2017) Cytogenetics and comparative analysis of Phaseolus Species. In: Fonsêca A (ed) The common bean Genome. Springer, Cham, pp 57–68
Fonsêca A, Ferreira J, dos Santos TRB et al (2010) Cytogenetic map of common bean (Phaseolus vulgaris L.). Chromosom Res 18:487–502. https://doi.org/10.1007/s10577-010-9129-8
Fonsêca A, Ferraz ME, Pedrosa-Harand A (2016) Speeding up chromosome evolution in Phaseolus: multiple rearrangements associated with a one-step descending dysploidy. Chromosoma 125:413–421. https://doi.org/10.1007/s00412-015-0548-3
Fransz P, Linc G, Lee C et al (2016) Molecular, genetic and evolutionary analysis of a paracentric inversion in Arabidopsis thaliana. Plant J 88:159–178. https://doi.org/10.1111/tpj.13262
Garcia T, Duitama J, Zullo SS et al (2021) Comprehensive genomic resources related to domestication and crop improvement traits in Lima bean. Nat Commun 12:1–17. https://doi.org/10.1038/s41467-021-20921-1
Gernhard T (2008) The conditioned reconstructed process. J Theor Biol 253:769–778. https://doi.org/10.1016/j.jtbi.2008.04.005
Guo X, Castillo-Ramírez S, González V et al (2007) Rapid evolutionary change of common bean (Phaseolus vulgaris L.) plastome, and the genomic diversification of legume chloroplasts. BMC Genomics 8:228. https://doi.org/10.1186/1471-2164-8-228
Han Y, Zhang T, Thammapichai P et al (2015) Chromosome-specific painting in Cucumis species using bulked oligonucleotides. Genetics 200:771–779. https://doi.org/10.1534/genetics.115.177642
He L, Zhao H, He J et al (2020) Extraordinarily conserved chromosomal synteny of Citrus species revealed by chromosome-specific painting. Plant J 103:2225–2235. https://doi.org/10.1111/tpj.14894
Iwata A, Tek AL, Richard MMS et al (2013) Identification and characterization of functional centromeres of the common bean. Plant J 76:47–60. https://doi.org/10.1111/tpj.12269
Jiang J (2019) Fluorescence in situ hybridization in plants: recent developments and future applications. Chromosome Res 27:153–165. https://doi.org/10.1007/s10577-019-09607-z
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Lewis G, Schrire B, Mackinder B, Lock M (2005) Legumes of the world, 1 ed. Royal botanic garden. Kew, London
Li H, Wang W, Lin L et al (2013) Diversification of the phaseoloid legumes: effects of climate change, range expansion and habit shift. Front Plant Sci 4:1–9. https://doi.org/10.3389/fpls.2013.00386
Liu X, Sun S, Wu Y et al (2020) Dual-color oligo-FISH can reveal chromosomal variations and evolution in Oryza species. Plant J 101:112–121. https://doi.org/10.1111/tpj.14522
Llaca V, Delgado Salinas A, Gepts P (1994) Chloroplast DNA as an evolutionary marker in the Phaseolus vulgaris complex. Theor Appl Genet 88:646–652. https://doi.org/10.1007/BF01253966
Lonardi S, Muñoz-Amatriaín M, Liang Q et al (2019) The genome of cowpea (Vigna unguiculata [L.] Walp.). Plant J 98:767–782. https://doi.org/10.1111/tpj.14349
Lysak MA, Berr A, Pecinka A et al (2006) Mechanisms of chromosome number reduction in Arabidopsis thaliana and related Brassicaceae species. PNAS 103:5224–5229. https://doi.org/10.1073/pnas.0510791103
Lysak MA, Mandáková T, Schranz ME (2016) Comparative paleogenomics of crucifers: ancestral genomic blocks revisited. Curr Opin Plant Biol 30:108–115. https://doi.org/10.1016/j.pbi.2016.02.001
Mandáková T, Lysak MA (2018) Post-polyploid diploidization and diversification through dysploid changes. Curr Opin Plant Biol 42:55–65. https://doi.org/10.1016/j.pbi.2018.03.001
Mandáková T, Hloušková P, Koch MA, Lysak MA (2020) Genome evolution in Arabideae was marked by frequent centromere repositioning. Plant Cell 32:650–665. https://doi.org/10.1105/tpc.19.00557
McClintock B (1979) The significance of responses of the genome to challenge. Science 226:792–801. https://doi.org/10.1126/science.15739260
Mercado-Ruaro P, Delgado-Salinas A (1996) Karyological studies in several Mexican species of Phaseolus L. and Vigna Savi (Phaseolinae, Fabaceae). In: Advances in legume systematics 8, Legumes of economic importance. Royal Botanic Gardens, Kew, pp 83–87
Mercado-Ruaro P, Delgado-Salinas A (1998) Karyotypic studies on species of Phaseolus (Fabaceae: Phaseolinae). Am J Bot 85:1–9. https://doi.org/10.2307/2446547
Mérot C, Oomen RA, Tigano A, Wellenreuther M (2020) A roadmap for understanding the evolutionary significance of structural genomic variation. Trends Ecol Evol 35:561–572. https://doi.org/10.1016/j.tree.2020.03.002
Mina-Vargas AM, McKeown PC, Flanagan NS et al (2016) Origin of year-long bean (Phaseolus dumosus Macfady, Fabaceae) from reticulated hybridization events between multiple Phaseolus species. Ann Bot 118:957–969. https://doi.org/10.1093/aob/mcw138
Moghaddam SM, Oladzad A, Koh C et al (2021) The tepary bean genome provides insight into evolution and domestication under heat stress. Nat Commun 12:1–14. https://doi.org/10.1038/s41467-021-22858-x
Moscone EA, Klein F, Lambrou M et al (1999) Quantitative karyotyping and dual-color FISH mapping of 5S and 18S–25S rDNA probes in the cultivated Phaseolus species (Leguminosae). Genome 42:1224–1233. https://doi.org/10.1139/g99-070
Mudd AB, Bredeson JV, Baum R et al (2020) Analysis of muntjac deer genome and chromatin architecture reveals rapid karyotype evolution. Commun Biol 3:480. https://doi.org/10.1038/s42003-020-1096-9
Nelson TC, Monnahan PJ, McIntosh MK et al (2019) Extreme copy number variation at a tRNA ligase gene affecting phenology and fitness in yellow monkeyflowers. Mol Ecol 28:1460–1475. https://doi.org/10.1111/mec.14904
Pedrosa-Harand A, De Almeida CCS, Mosiolek M et al (2006) Extensive ribosomal DNA amplification during Andean common bean (Phaseolus vulgaris L.) evolution. Theor Appl Genet 112:924–933. https://doi.org/10.1007/s00122-005-0196-8
Piperidis N, D’Hont A (2020) Sugarcane genome architecture decrypted with chromosome-specific oligo probes. Plant J 103:2039–2051. https://doi.org/10.1111/tpj.14881
Porubsky D, Sanders AD, Höps W et al (2020) Recurrent inversion toggling and great ape genome evolution. Nat Genet 52:849–858. https://doi.org/10.1038/s41588-020-0646-x
POWO (2023) Plants of the world online. Facilitated by the royal botanic gardens, Kew. http://www.plantsoftheworldonline.org/. Accessed 23 July 2023
Rambaut A, Drummond AJ, Xie D et al (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Rambaut A, Drummond AJ (2013) TreeAnnotator v1.10.4. Available as Part of the BEAST package. In: http://beast.bio.ed.ac.uk
Rambaut A (2018) FigTree v 1.4.4. http://tree.bio.ed.ac.uk/software/figtree/. Accessed 23 July 2023
Rendón-Anaya M, Herrero-Estrella A (2017) Requirement of whole-genome sequencing. In: Pérez de la Vega M, Santalla M, Marsolais F (eds) The common bean genome compendium of plant genomes. Springer, Cham. https://doi.org/10.1007/978-3-319-63526-2_5
Rendón-Anaya M, Montero-Vargas JM, Saburido-Álvarez S et al (2017) Genomic history of the origin and domestication of common bean unveils its closest sister species. Genome Biol 18:1–17. https://doi.org/10.1186/s13059-017-1190-6
Ribeiro T, dos Santos KGB, Richard MMS et al (2017) Evolutionary dynamics of satellite DNA repeats from Phaseolus beans. Protoplasma 254:791–801. https://doi.org/10.1007/s00709-016-0993-8
Ronquist F, Teslenko M, van der Mark P et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Schmutz J, McClean PE, Mamidi S et al (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713. https://doi.org/10.1038/ng.3008
Sotero-Caio CG, Volleth M, Hoffmann FG et al (2015) Integration of molecular cytogenetics, dated molecular phylogeny, and model-based predictions to understand the extreme chromosome reorganization in the Neotropical genus Tonatia (Chiroptera: Phyllostomidae). BMC Evol Biol 15:1–15. https://doi.org/10.1186/s12862-015-0494-y
Thompson MJ, Jiggins CD (2014) Supergenes and their role in evolution. Heredity 113:1–8. https://doi.org/10.1038/hdy.2014.20
Tillich M, Lehwark P, Pellizzer T et al (2017) GeSeq–versatile and accurate annotation of organelle genomes. Nucleic Acids Res 45:W6–W11. https://doi.org/10.1093/nar/gkx391
Van De Peer Y, Mizrachi E, Marchal K (2017) The evolutionary significance of polyploidy. Nat Rev Genet 18:411–424. https://doi.org/10.1038/nrg.2017.26
Vlasova A, Capella-Gutiérrez S, Rendón-Anaya M et al (2016) Genome and transcriptome analysis of the Mesoamerican common bean and the role of gene duplications in establishing tissue and temporal specialization of genes. Genome Biol 17:1–18. https://doi.org/10.1186/s13059-016-0883-6
Vogel JP, Garvin DF, Mockler TC et al (2010) Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature 463:763–768. https://doi.org/10.1038/nature08747
Waminal NE, Pellerin RJ, Kang S-H, Kim HH (2021) Chromosomal mapping of tandem repeats revealed massive chromosomal rearrangements and insights into Senna tora dysploidy. Front Plant Sci 12:1–13. https://doi.org/10.3389/fpls.2021.629898
Wellenreuther M, Bernatchez L (2018) Eco-Evolutionary genomics of chromosomal inversions. Trends Ecol Evol 33:427–440. https://doi.org/10.1016/j.tree.2018.04.002
Winterfeld G, Becher H, Voshell S et al (2018) Karyotype evolution in Phalaris (Poaceae): the role of reductional dysploidy, polyploidy and chromosome alteration in a wide-spread and diverse genus. PLoS One 13:e0192869. https://doi.org/10.1371/journal.pone.0192869
Xin H, Zhang T, Wu Y et al (2019) An extraordinarily stable karyotype of the woody Populus species revealed by chromosome painting. Plant J 101:253–264. https://doi.org/10.1111/tpj.14536
Acknowledgements
We would like to kindly thank the CIAT, EMBRAPA Cenargen and IPK for providing the seeds, as well as Fernanda de Oliveira Bustamante, Lívia do Vale Martins, Guilherme Tomaz Braz, Jiming Jiang, and Ana Christina Brasileiro-Vidal for the development of oligo-FISH probes.
Funding
This study was supported by CNPq (National Council for Scientific and Technological Development, acronym in Portuguese, Grant no. 310804/2017–5) and CAPES (Coordination of Improvement of Higher Education Personnel, acronym in Portuguese, Financial Code 001).
Author information
Authors and Affiliations
Contributions
T.N. conducted experiments, analyzsd the data, constructed the figures, and wrote the paper. A.P.H. conceived this research, provided the resources, discussed the results, and helped writing the paper. Both authors read and approved the final version of this text.
Corresponding author
Ethics declarations
Conflict of interests
The authors declare no conflicts of interest.
Additional information
Communicated by Annaliese S. Mason.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
122_2023_4462_MOESM1_ESM.pdf
Supplementary figure S1 Distribution of structural rearrangements among Phaseolus species over evolutionary time. (PDF 882 kb)
122_2023_4462_MOESM4_ESM.pdf
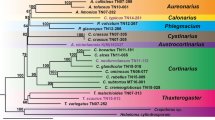
Supplementary figure S4 Phylogenetic tree based on concatenated ITS loci and chloroplast genomes; the negative numbers on the nodes correspond to the ages of each clade in millions of years ago. (PDF 80 kb)
122_2023_4462_MOESM5_ESM.pdf
Supplementary Table S1 Accession codes for the sequencing data deposited in the GenBank used for phylogenetic analyses. (PDF 4 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nascimento, T., Pedrosa-Harand, A. High rates of structural rearrangements have shaped the chromosome evolution in dysploid Phaseolus beans. Theor Appl Genet 136, 215 (2023). https://doi.org/10.1007/s00122-023-04462-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04462-3