Abstract
Key message
Structural variations are common in plant genomes, affecting meiotic recombination and distorted segregation in wheat. And presence/absence variations can significantly affect drought tolerance in wheat.
Abstract
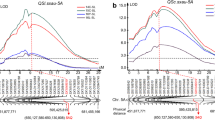
Drought is a major abiotic stress limiting wheat production. Common wheat has a complex genome with three sub-genomes, which host large numbers of structural variations (SVs). SVs play critical roles in understanding the genetic contributions of plant domestication and phenotypic plasticity, but little is known about their genomic characteristics and their effects on drought tolerance. In the present study, high-resolution karyotypes of 180 doubled haploids (DHs) were developed. Signal polymorphisms between the parents involved with 8 presence-absence variations (PAVs) of tandem repeats (TR) distributed on the 7 (2A, 4A, 5A, 7A, 3B, 7B, and 2D) of 21 chromosomes. Among them, PAV on chromosome 2D showed distorted segregation, others transmit normal conforming to a 1:1 segregation ration in the population; and a PAVs recombination occurred on chromosome 2A. Association analysis of PAV and phenotypic traits under different water regimes, we found PAVs on chromosomes 4A, 5A, and 7B showed negative effect on grain length (GL) and grain width (GW); PAV.7A had opposite effect on grain thickness (GT) and spike length (SL), with the effect on traits differing under different water regimes. PAVs on linkage group 2A, 4A, 7A, 2D, and 7B associated with the drought tolerance coefficients (DTCs), and significant negative effect on drought resistance values (D values) were detected in PAV.7B. Additionally, quantitative trait loci (QTL) associated with phenotypic traits using the 90 K SNP array showed QTL for DTCs and grain-related traits in chromosomes 4A, and 5A, 3B were co-localized in differential regions of PAVs. These PAVs can cause the differentiation of the target region of SNP and could be used for genetic improvement of agronomic traits under drought stress via marker-assisted selection (MAS) breeding.





Similar content being viewed by others
Data availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Alonge M, Wang X, Benoit M, Soyk S, Pereira L, Zhang L, Suresh H, Ramakrishnan S, Maumna F, Ciren D, Levy Y, Harel TH, Shalev-Schlosser G, Amsellem Z, Razifard H, Caicedo AL, Tieman DM, Klee H, Kirsche M, Aganezov S, Ranallo-Benavidez TR, Lemmon ZH, Kim J, Robitaille G, Kramer M, Goodwin S, McCombie WR, Hutton S, Eck JV, Gillis J, Eshed Y, Sedlazeck FJ, Knaap E, Schatz MC, Lippman ZB (2020) Major impacts of widespread structural variation on gene expression and crop improvement in tomato. Cell 182:145–61.e23. https://doi.org/10.1016/j.cell.2020.05.021
Badaeva ED, Dedkova OS, Gay G, Pukhalskyi VA, Zelenin AV, Bernard S, Bernard M (2007) Chromosomal rearrangements in wheat: their types and distribution. Genome 50:907–926. https://doi.org/10.1139/g07-072
Borde V (2007) The multiple roles of the Mre11 complex for meiotic recombination. Chromosome Res 15:551–563. https://doi.org/10.1007/s10577-007-1147-9
Cao S, Xu D, Hanif M, Xia X, He Z (2020) Genetic architecture underpinning yield component traits in wheat. Theor Appl Genet 133:1811–1823. https://doi.org/10.1007/s00122-020-03562-8
Chen J, Tang Y, Yao L, Wu H, Tu X, Zhuang L, Qi ZJ (2019) Cytological and molecular characterization of Thinopyrum bessarabicum chromosomes and structural rearrangements introgressed in wheat. Mol Breed 39:146. https://doi.org/10.1007/s11032-019-1054-8
Collins RL, Brand H, Karczewski KJ, Zhao X, Alföldi J, Francioli LC, Khera AV, Lowther C, Gauthier LD, Wang H, Watts NA, Solomonson M, O’Donnell-Luria A, Baumann A, Munshi R, Walker M, Whelan CW, Huang Y, Brookings T, Sharpe T, Stone MR, Valkanas E, Fu J, Tiao G, Laricchia KM, Ruano-Rubio V, Stevens C, Gupta N, Cusick C, Margolin L, Taylor KD, Lin HJ, Rich SS, Post WS, Chen YI, Rotter JI, Nusbaum C, Philippakis A, Lander E, Gabriel S, Neale BM, Kathiresan S, Daly MJ, Banks E, MacArthur DG, Talkowski ME (2020) A structural variation reference for medical and population genetics. Nature 581:444–451. https://doi.org/10.1038/s41586-020-2287-8
Contento A, Heslop-Harrison JS, Schwarzacher T (2005) Diversity of a major repetitive DNA sequence in diploid and polyploid Triticeae. Cytogenet Genome Res 109:34–42. https://doi.org/10.1159/000082379
Crespo Herrera LA, Garkava Gustavsson L, Åhman I (2017) A systematic review of rye (Secale cereale L.) as a source of resistance to pathogens and pests in wheat (Triticum aestivum L.). Hereditas 154:14. https://doi.org/10.1186/s41065-017-0033-5
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST, McVean G, Durbin R (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Doolittle WF, Sapienza C (1980) Selfish genes, the phenotype paradigm and genome evolution. Nature 284:601–603. https://doi.org/10.1038/284601a0
Dvorak J, Wang L, Zhu T, Jorgensen CM, Luo MC, Deal KR, Gu YQ, Gill BS, Distelfeld A, Devos KM, Qi P, McGuire PE (2018) Reassessment of the evolution of wheat chromosomes 4A, 5A, and 7B. Theor Appl Genet 131:2451–2462. https://doi.org/10.1007/s00122-018-3165-8
Erayman M, Sandhu D, Sidhu D, Dilbirligi M, Baenziger PS, Gill KS (2004) Demarcating the gene-rich regions of the wheat genome. Nucleic Acids Res 32:3546–3565. https://doi.org/10.1093/nar/gkh639
Feuk L, Carson AR, Scherer SW (2006) Structural variation in the human genome. Nat Rev Genet 7:85–97. https://doi.org/10.1038/nrg1767
Flagel L, Brandvain Y, Schrider DR (2019) The unreasonable effectiveness of convolutional neural networks in population genetic inference. Mol Biol Evol 36:220–238. https://doi.org/10.1093/molbev/msy224
Fransz P, Linc G, Lee CR, Aflitos SA, Lasky JR, Toomajian C, Ali H, Peters J, van Dam P, Ji X, Kuzak M, Gerats T, Schubert I, Schneeberger K, Colot V, Martienssen R, Koornneef M, Nordborg M, Juenger TE, de Jong H, Schranz ME (2016) Molecular, genetic and evolutionary analysis of a paracentric inversion in Arabidopsis thaliana. Plant J 88:159–178. https://doi.org/10.1111/tpj.13262
Frova C, Krajewski P, di Fonzo N, Villa M, Sari-Gorla M (1999) Genetic analysis of drought tolerance in maize by molecular markers I. yield components. Theor Appl Genet 99:280–288. https://doi.org/10.1007/s001220051233
Gabur I, Chawla HS, Snowdon RJ, Parkin IAP (2019) Connecting genome structural variation with complex traits in crop plants. Theor Appl Genet 132:733–750. https://doi.org/10.1007/s00122-018-3233-0
Gardiner LJ, Wingen LU, Bailey P, Joynson R, Brabbs T, Wright J, Higgins JD, Hall N, Griffiths S, Clavijo BJ, Hall A (2019) Analysis of the recombination landscape of hexaploid bread wheat reveals genes controlling recombination and gene conversion frequency. Genome Biol 20:69. https://doi.org/10.1186/s13059-019-1675-6
Gemayel R, Cho J, Boeynaems S, Verstrepen KJ (2012) Beyond junk-variable tandem repeats as facilitators of rapid evolution of regulatory and coding sequences. Genes (basel) 3:461–480. https://doi.org/10.3390/genes3030461
Guo Z, Yang W, Chang Y, Ma X, Tu H, Xiong F, Jiang N, Feng H, Huang C, Yang P, Zhao H, Chen G, Liu H, Luo L, Hu H, Liu Q, Xiong L (2018) Genome-wide association studies of image traits reveal genetic architecture of drought resistance in rice. Mol Plant 11:789–805. https://doi.org/10.1016/j.molp.2018.03.018
Harushima Y, Kurata N, Yano M, Nagamura Y, Sasaki T, Minobe Y, Nakagahra M (1996) Detection of segregation distortions in an indica-japonica rice cross using a high- resolution molecular map. Theor Appl Genet 92:145–150. https://doi.org/10.1007/BF00223368
He L, Dooner HK (2009) Haplotype structure strongly affects recombination in a maize genetic interval polymorphic for Helitron and retrotransposon insertions. Proc Natl Acad Sci U S A 106:8410–8416. https://doi.org/10.1073/pnas.0902972106
Huang X, Zhu M, Zhuang L, Zhang S, Wang J, Chen X, Wang D, Chen J, Bao Y, Guo J, Zhang J, Feng Y, Chu C, Du P, Qi Z, Wang H, Chen P (2018) Structural chromosome rearrangements and polymorphisms identified in Chinese wheat cultivars by high-resolution multiplex oligonucleotide FISH. Theor Appl Genet 131:1967–1986. https://doi.org/10.1007/s00122-018-3126-2
IWGSC (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science. https://doi.org/10.1126/science.aar7191
Jia ML, Li YN, Wang ZY, Tao S, Sun GL, Kong XC, Wang K, Ye XG, Liu SS, Geng SF, Mao L, Li AL (2021) TaIAA21 represses TaARF25-mediated expression of TaERFs required for grain size and weight development in wheat. Plant J 108:1754–1767. https://doi.org/10.1111/tpj.15541
Jordan KW, Wang S, He F, Chao S, Lun Y, Paux E, Sourdille P, Sherman J, Akhunova A, Blake NK, Pumphrey MO, Glover K, Dubcovsky J, Talbert L, Akhunov ED (2018) The genetic architecture of genome-wide recombination rate variation in allopolyploid wheat revealed by nested association mapping. Plant J 95:1039–1054. https://doi.org/10.1111/tpj.14009
Keidar-Friedman D, Bariah I, Domb K, Kashkush K (2020) The evolutionary dynamics of a novel miniature transposable element in the wheat genome. Front Plant Sci 11:1173. https://doi.org/10.3389/fpls.2020.01173
Klepikova AV, Kasianov AS, Gerasimov ES, Logacheva MD, Penin AA (2016) A high resolution map of the Arabidopsis thaliana developmental transcriptome based on RNA-seq profiling. Plant J 88:1058–1070. https://doi.org/10.1111/tpj.13312
Kianian SF, Quiros CF (1992) Generation of a Brassica oleracea composite RFLP map: linkage arrangements among various populations and evolutionary implications. Theor Appl Genet 84:544–554. https://doi.org/10.1007/BF00224150
Knox AK, Dhillon T, Cheng H, Tondelli A, Pecchioni N, Stockinger EJ (2010) CBF gene copy number variation at Frost Resistance-2 is associated with levels of freezing tolerance in temperate-climate cereals. Theor Appl Genet 121:21–35. https://doi.org/10.1007/s00122-010-1288-7
Kou Y, Liao Y, Toivainen T, Lv Y, Tian X, Emerson JJ, Gaut BS, Zhou Y (2020) Evolutionary genomics of structural variation in Asian rice (Oryza sativa) domestication. Mol Biol Evol 37:3507–3524. https://doi.org/10.1093/molbev/msaa185
Lang T, Li G, Wang H, Yu Z, Chen Q, Yang E, Fu S, Tang Z, Yang Z (2019) Physical location of tandem repeats in the wheat genome and application for chromosome identification. Planta 249:663–675. https://doi.org/10.1007/s00425-018-3033-4
Lein A (1943) Die genetische grundlage der kreuzbarkeit zwischen weizen und roggen. Z Indukt Abstamm Vererbungsl 81:28–61. https://doi.org/10.1007/BF01847441
Li A, Hao C, Wang Z, Geng S, Jia M, Wang F, Han X, Kong X, Yin L, Tao S, Deng Z, Liao R, Sun G, Wang K, Ye X, Jiao C, Lu H, Zhou Y, Liu D, Fu X, Zhang X, Mao L (2022) Wheat breeding history reveals synergistic selection of pleiotropic genomic sites for plant architecture and grain yield. Mol Plant 15:504–519. https://doi.org/10.1016/j.molp.2022.01.004
Lin Y, Chen G, Hu H, Yang X, Zhang Z, Jiang X, Wu F, Shi H, Wang Q, Zhou K, Li C, Ma J, Zheng Y, Wei Y, Liu Y (2020) Phenotypic and genetic variation in phosphorus-deficiency-tolerance traits in Chinese wheat landraces. BMC Plant Biol 20:330. https://doi.org/10.1186/s12870-020-02492-3
Liu C, Gong W, Han R, Guo J, Li G, Li H, Song J, Liu A, Cao X, Zhai S, Cheng D, Li G, Zhao Z, Yang Z, Liu J, Reader SM (2019) Characterization, identification and evaluation of a set of wheat-Aegilops comosa chromosome lines. Sci Rep 9:4773. https://doi.org/10.1038/s41598-019-41219-9
Lu F, Romay MC, Glaubitz JC, Bradbury PJ, Elshire RJ, Wang T, Li Y, Li Y, Semagn K, Zhang X, Hernandez AG, Mikel MA, Soifer I, Barad O, Buckler ES (2015) High-resolution genetic mapping of maize pan-genome sequence anchors. Nat Commun 6:6914. https://doi.org/10.1038/ncomms7914
Luo DP, Xu H, Liu ZL, Guo JX, Li HY, Chen LT, Fang C, Zhang QY, Bai M, Yao N, Wu H, Ji CH, Zheng HQ, Chen YL, Ye S, Li XY, Zhao XC, Li RQ, Liu YG (2013) A detrimental mitochondrial-nuclear interaction causes cytoplasmic male sterility in rice. Nat Genet 45:573–577. https://doi.org/10.1038/ng.2570
Ma J, Li HB, Zhang CY, Yang XM, Liu YX, Yan GJ, Liu CJ (2010) Identification and validation of a major QTL conferring crown rot resistance in hexaploid wheat. Theor Appl Genet 120:1119–1128. https://doi.org/10.1007/s00122-009-1239-3
Ma S, Wang M, Wu J, Guo W, Chen Y, Li G, Wang Y, Shi W, Xia G, Fu D, Kang Z, Ni F (2021) WheatOmics: a platform combining multiple omics data to accelerate functional genomics studies in wheat. Mol Plant 14:1965–1968. https://doi.org/10.1016/j.molp.2021.10.006
Mahmoud M, Gobet N, Cruz-Dávalos DI, Mounier N, Dessimoz C, Sedlazeck FJ (2019) Structural variant calling: the long and the short of it. Genome Biol 20:246. https://doi.org/10.1186/s13059-019-1828-7
Mahmoud M, Gracz-Bernaciak J, Żywicki M, Karłowski W, Twardowski T, Tyczewska A (2020) Identification of structural variants in two novel genomes of Maize inbred lines possibly related to glyphosate tolerance. Plants (basel). https://doi.org/10.3390/plants9040523
Maron LG, Guimarães CT, Kirst M, Albert PS, Birchler JA, Bradbury PJ, Buckler ES, Coluccio AE, Danilova TV, Kudrna D, Magalhaes JV, Piñeros MA, Schatz MC, Wing RA, Kochian LV (2013) Aluminum tolerance in maize is associated with higher MATE1 gene copy number. Proc Natl Acad Sci USA 110:5241–5246. https://doi.org/10.1073/pnas.1220766110
McHale LK, Haun WJ, Xu WW, Bhaskar PB, Anderson JE, Hyten DL, Gerhardt DJ, Jeddeloh JA, Stupar RM (2012) Structural variants in the soybean genome localize to clusters of biotic stress-response genes. Plant Physiol 159:1295–1308. https://doi.org/10.1104/pp.112.194605
Messmer MM, Keller M, Zanetti S, Keller B (1999) Genetic linkage map of a wheat×spelt cross. Theor Appl Genet 98:1163–1170. https://doi.org/10.1007/s001220051181
Milner MJ, Swarbreck SM, Craze M, Bowden S, Griffiths H, Bentley AR, Wallington EJ (2022) Over-expression of TaDWF4 increases wheat productivity under low and sufficient nitrogen through enhanced carbon assimilation. Commun Biol 5:193. https://doi.org/10.1038/s42003-022-03139-9
Mizuno N, Kinoshita M, Kinoshita S, Nishida H, Fujita M, Kato K et al (2016) Loss-of-function mutations in three homoeologous PHYTOCLOCK 1 Genes in common wheat are associated with the extra-early flowering phenotype. PLoS ONE 11:e0165618. https://doi.org/10.1371/journal.pone.0165618
Mizuta Y, Harushima Y, Kurata N (2010) Rice pollen hybrid incompatibility caused by reciprocal gene loss of duplicated genes. Proc Natl Acad Sci USA 107:20417–20422. https://doi.org/10.1073/pnas.1003124107
Nelson JC, Sorrells ME, Van Deynze AE, Lu YH, Atkinson M, Bernard M, Leroy P, Faris JD, Anderson JA (1995) Molecular mapping of wheat: major genes and rearrangements in homoeologous groups 4, 5, and 7. Genetics 141:721–731. https://doi.org/10.1093/genetics/141.2.721
Pauline T, Julien B, Chris B, Radoslaw S, Ute B, Priyanka K, Peter L, Penny T, Delphine F (2021) The wheat seven in absentia gene is associated with increases in biomass and yield in hot climates. J Exp Bot 72:3774–3791. https://doi.org/10.1093/jxb/erab044
Saintenac C, Faure S, Remay A, Choulet F, Ravel C, Paux E, Balfourier F, Feuillet C, Sourdille P (2011) Variation in crossover rates across a 3-Mb contig of bread wheat (Triticum aestivum) reveals the presence of a meiotic recombination hotspot. Chromosoma 120:185–198. https://doi.org/10.1007/s00412-010-0302-9
Sandler L, Novitski E (1957) Meiotic drive as an evolutionary force. Am Nat 91:105–110. https://doi.org/10.1086/281969
Sandler L, Hiraizumi Y, Sandler I (1959) Meiotic drive in natural populations of Drosophila melanogaster. I. the cytogenetic basis of segregation-distortion. Genetics 44:233–250. https://doi.org/10.1093/genetics/44.2.233
Sari-Gorla M, Krajewski P, Di Fonzo N, Villa M, Frova C (1999) Genetic analysis of drought tolerance in maize by molecular markers. II. Plant height and flowering. Theor Appl Genet 99:289–295. https://doi.org/10.1007/s001220051234
Saxena RK, Edwards D, Varshney RK (2014) Structural variations in plant genomes. Brief Funct Genomics 13:296–307. https://doi.org/10.1093/bfgp/elu016
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M (2008) Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet 40:1023–1028. https://doi.org/10.1038/ng.169
Schiessl S-V, Katche E, Ihien E, Chawla HS, Mason AS (2019) The role of genomic structural variation in the genetic improvement of polyploid crops. Crop J 7:127–140. https://doi.org/10.1016/j.cj.2018.07.006
Song JM, Guan Z, Hu J, Guo C, Yang Z, Wang S, Liu D, Wang B, Lu S, Zhou R, Xie WZ, Cheng Y, Zhang Y, Liu K, Yang QY, Chen LL, Guo L (2020) Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat Plants 6:34–45. https://doi.org/10.1038/s41477-019-0577-7
Sorrells M, Gustafson J, Somers D, Chao S, Benscher D, Guedira-Brown G, Huttner E, Kilian A, McGuire P, Ross K, Tanaka J, Wenzl P, Williams K, Qualset C (2011) Reconstruction of the Synthetic W7984 × Opata M85 wheat reference population. Genome 54:875–882. https://doi.org/10.1139/g11-054
Sutton T, Baumann U, Hayes J, Collins NC, Shi BJ, Schnurbusch T, Hay A, Mayo G, Pallotta M, Tester M, Langridge P (2007) Boron-toxicity tolerance in barley arising from efflux transporter amplification. Science 318:1446–1449. https://doi.org/10.1126/science.1146853
Stephens SG (1949) The cytogenetics of speciation in Gossypium. I. Selective elimination of the donor parent genotype in interspecific backcrosses. Genetics 34:627–637. https://doi.org/10.1093/genetics/34.5.627
Tang ZX, Wang XF, Zhang MZ, Zhang YH, Deng DX, Xu CW (2013) The maternal cytoplasmic environment may be involved in the viability selection of gametes and zygotes. Heredity 110:331–337. https://doi.org/10.1038/hdy.2012.89
Tao Y, Zhao X, Mace E, Henry R, Jordan D (2019) Exploring and exploiting pan-genomics for crop improvement. Mol Plant 12:156–169. https://doi.org/10.1016/j.molp.2018.12.016
Tian T, Liu Y, Yan H, You Q, Yi X, Du Z, Xu W, Su Z (2017) AgriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res 45:w122–w129. https://doi.org/10.1093/nar/gkx382
Wang H, Yin H, Jiao C, Fang X, Wang G, Li G, Ni F, Li P, Su P, Ge W, Lyu Z, Xu S, Yang Y, Hao Y, Cheng X, Zhao J, Liu C, Xu F, Ma X, Sun S, Zhao Y, Bao Y, Liu C, Zhang J, Pavlicek T, Li A, Yang Z, Nevo E, Kong L (2020) Sympatric speciation of wild emmer wheat driven by ecology and chromosomal rearrangements. Proc Natl Acad Sci U S A 117:5955–5963. https://doi.org/10.1073/pnas.1920415117
Wang Y, Xiong G, Hu J, Jiang L, Yu H, Xu J, Fang Y, Zeng L, Xu E, Xu J, Ye W, Meng X, Liu R, Chen H, Jing Y, Wang Y, Zhu X, Li J, Qian Q (2015) Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat Genet 47:944–948. https://doi.org/10.1038/ng.3346
Wu CI, Ting CT (2004) Genes and speciation. Nat Rev Genet 5:114–122. https://doi.org/10.1038/nrg1269
Wang Y, Qiao L, Yang CK, Li XH, Zhao JJ, Wu BB, Zheng XW, Zheng J, Li PB (2022) Identification of genetic loci for flag-leaf-related traits in wheat (Triticum aestivum L.) and their effects on grain yield. Front Plant Sci 13:990287. https://doi.org/10.3389/fpls.2022.990287
Xu X, Liu X, Ge S, Jensen JD, Hu F, Li X, Dong Y, Gutenkunst RN, Fang L, Huang L, Li J, He W, Zhang G, Zheng X, Zhang F, Li Y, Yu C, Kristiansen K, Zhang X, Wang J, Wright M, McCouch S, Nielsen R, Wang J, Wang W (2012) Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat Biotechnol 30:105–111. https://doi.org/10.1038/nbt.2050
Xu Y, Zhu L, Xiao J, Huang N, McCouch SR (1997) Chromosomal regions associated with segregation distortion of molecular markers in F2, backcross, doubled haploid, and recombinant inbred populations in rice (Oryza sativa L.). Mol Gen Genet 253:535–545. https://doi.org/10.1007/s004380050355
Yamamori M, Nakamura T, Endo TR, Nagamine T (1994) Waxy protein deficiency and chromosomal location of coding genes in common wheat. Theor Appl Genet 89:179–184. https://doi.org/10.1007/bf00225138
Yamagata Y, Yamamoto E, Aya K, Win KT, Doi K, Sobrizal IT, Kanamori H, Wu JZ, Matsumoto T, Matsuoka M, Ashikari M, Yoshimura A (2010) Mitochondrial gene in the nuclear genome induces reproductive barrier in rice. Proc Natl Acad Sci USA 107:1494–1499. https://doi.org/10.1073/pnas.0908283107
Yang JY, Zhao XB, Cheng K, Du HY, Ouyang YD, Chen JJ, Qiu SQ, Huang JY, Jiang YH, Jiang LW, Ding JH, Wang J, Xu CG, Li XH, Zhang QF (2012) A killer-protector system regulates both hybrid sterility and segregation distortion in rice. Science 337:1336–1340. https://doi.org/10.1126/science.1223702
Yang Y, Amo A, Wei D, Chai Y, Zheng J, Qiao P, Cui C, Lu S, Chen L, Hu YG (2021) Large-scale integration of meta-QTL and genome-wide association study discovers the genomic regions and candidate genes for yield and yield-related traits in bread wheat. Theor Appl Genet 134:3083–3109. https://doi.org/10.1007/s00122-021-03881-4
Yang Z, Ge X, Yang Z, Qin W, Sun G, Wang Z, Li Z, Liu J, Wu J, Wang Y, Lu L, Wang P, Mo H, Zhang X, Li F (2019) Extensive intraspecific gene order and gene structural variations in upland cotton cultivars. Nat Commun 10:2989. https://doi.org/10.1038/s41467-019-10820-x
Yu P, Wang C, Xu Q, Feng Y, Yuan X, Yu H, Wang Y, Tang S, Wei X (2011) Detection of copy number variations in rice using array-based comparative genomic hybridization. BMC Genomics 12:372. https://doi.org/10.1186/1471-2164-12-372
Zhang J, Hao C, Ren Q, Chang X, Liu G, Jing R (2011) Association mapping of dynamic developmental plant height in common wheat. Planta 234:891–902. https://doi.org/10.1007/s00425-011-1434-8
Zhao LB, Xie D, Huang L, Zhang SJ, Luo JT, Jiang B, Ning SZ, Zhang LQ, Yuan ZW, Wang JR, Zheng YI, Liu DC, Hao M (2021) Integrating the physical and genetic map of bread wheat facilitates the detection of chromosomal rearrangements. J Integr Agric 20:2333–2342. https://doi.org/10.1016/S2095-3119(20)63289-0
Zhang X, Chen X, Liang P, Tang H (2018) Cataloging plant genome structural variations. Curr Issues Mol Biol 27:181–194. https://doi.org/10.21775/cimb.027.181
Zhang X, Zhu Y, Kremling KAG, Romay MC, Bukowski R, Sun Q, Gao S, Buckler ES, Lu F (2022) Genome-wide analysis of deletions in maize population reveals abundant genetic diversity and functional impact. Theor Appl Genet 135:273–290. https://doi.org/10.1007/s00122-021-03965-1
Zheng LY, Guo XS, He B, Sun LJ, Peng Y, Dong SS, Liu TF, Jiang S, Ramachandran S, Liu CM, Jing HC (2011) Genome-wide patterns of genetic variation in sweet and grain sorghum (Sorghum bicolor). Genome Biol 12:R114. https://doi.org/10.1186/gb-2011-12-11-r114
Zhou YF, Minio A, Massonnet M, Solares E, Lv YD, Beridze T, Cantu D, Gaut BS (2019) The population genetics of structural variants in grapevine domestication. Nat Plants 5:965–979. https://doi.org/10.1038/s41477-019-0507-8
Zhu XL, Rong W, Wang K, Guo W, Zhou M, Wu JZ, Ye XG, Wei XN, Zhang ZY (2022) Overexpression of TaSTT3b-2B improves resistance to sharp eyespot and increases grain weight in wheat. Plant Biotechnol J 20:777–793. https://doi.org/10.1111/pbi.13760
Zou Y, Wan LR, Luo J, Tang ZX, Fu SL (2021) FISH landmarks reflecting meiotic recombination and structural alterations of chromosomes in wheat (Triticum aestivum L.). BMC Plant Biol 21:167. https://doi.org/10.1186/s12870-021-02947-1
Acknowledgements
This project was supported by State Key Laboratory of Sustainable Dryland Agriculture (No. YJHZKF2107), National Natural Science Foundation of China (31871571), and Research Program of Shanxi Agricultural University (YBSJJ2006) Bioinformatics analyses were supported by the Bioinformatics Center of Nanjing Agricultural University, China. We thank the editor and anonymous reviewers for critical comments and suggestions for improving the manuscript.
Funding
This project was supported by State Key Laboratory of Sustainable Dryland Agriculture (No. YJHZKF2107), National Natural Science Foundation of China (31871571), and Research Program of Shanxi Agricultural University (YBSJJ2006).
Author information
Authors and Affiliations
Contributions
WDY and JZ planned and designed the experiments. JJZ, BBW, XWZ, and MJG performed the experiments. JJZ, ZJQ, LQ, and MCF analyzed the data. JJZ and XHL wrote the article. All authors read and commented on the final version for publication.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Peter Langridge.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, J., Li, X., Qiao, L. et al. Identification of structural variations related to drought tolerance in wheat (Triticum aestivum L.). Theor Appl Genet 136, 37 (2023). https://doi.org/10.1007/s00122-023-04283-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04283-4